Abstract
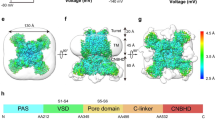
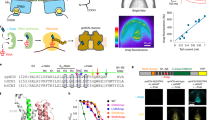
Hv1 voltage-gated proton channels mediate rapid and selective transmembrane H+ flux and are gated by both voltage and pH gradients. Selective H+ transfer in membrane proteins is commonly achieved by Grotthuss proton 'hopping' in chains of ionizable amino acid side chains and intraprotein water molecules. To identify whether ionizable residues are required for proton permeation in Hv1, we neutralized candidate residues and measured expressed voltage-gated H+ currents. Unexpectedly, charge neutralization was insufficient to abrogate either the Hv1 conductance or coupling of pH gradient and voltage-dependent activation. Molecular dynamics simulations revealed water molecules in the central crevice of Hv1 model structures but not in homologous voltage-sensor domain (VSD) structures. Our results indicate that Hv1 most likely forms an internal water wire for selective proton transfer and that interactions between water molecules and S4 arginines may underlie coupling between voltage- and pH-gradient sensing.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$189.00 per year
only $15.75 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
References
Bezanilla, F. How membrane proteins sense voltage. Nat. Rev. Mol. Cell Biol. 9, 323–332 (2008).
Long, S.B., Campbell, E.B. & Mackinnon, R. Voltage sensor of Kv1.2: structural basis of electromechanical coupling. Science 309, 903–908 (2005).
Swartz, K.J. Sensing voltage across lipid membranes. Nature 456, 891 (2008).
Ramsey, I.S., Moran, M.M., Chong, J.H.A. & Clapham, D.E. A voltage-gated proton-selective channel lacking the pore domain. Nature 440, 1213–1216 (2006).
Sasaki, M., Takagi, M. & Okamura, Y. A voltage sensor-domain protein is a voltage-gated proton channel. Science 312, 589–592 (2006).
DeCoursey, T.E. Voltage-gated proton channels. Cell. Mol. Life Sci. 65, 2554–2573 (2008).
Musset, B. et al. Detailed comparison of expressed and native voltage-gated proton channel currents. J. Physiol. (Lond.) 586, 2477 (2008).
Tombola, F., Ulbrich, M.H. & Isacoff, E.Y. The voltage-gated proton channel Hv1 has two pores, each controlled by one voltage sensor. Neuron 58, 546–556 (2008).
Koch, H.P. et al. Multimeric nature of voltage-gated proton channels. Proc. Natl. Acad. Sci. USA 105, 9111–9116 (2008).
Swanson, J.M. et al. Proton solvation and transport in aqueous and biomolecular systems: insights from computer simulations. J. Phys. Chem. B 111, 4300–4314 (2007).
Buch-Pedersen, M.J., Pedersen, B.P., Veierskov, B., Nissen, P. & Palmgren, M.G. Protons and how they are transported by proton pumps. Pflugers Arch. 457, 573 (2009).
Cukierman, S. The transfer of protons in water wires inside proteins. Front. Biosci. 8, s1118–s1139 (2003).
Cherny, V.V., Markin, V.S. & DeCoursey, T.E. The voltage-activated hydrogen ion conductance in rat alveolar epithelial cells is determined by the pH gradient. J. Gen. Physiol. 105, 861–896 (1995).
Starace, D.M. & Bezanilla, F. A proton pore in a potassium channel voltage sensor reveals a focused electric field. Nature 427, 548 (2004).
Tombola, F., Ulbrich, M., Kohout, S. & Isacoff, E. The opening of the two pores of the Hv1 voltage-gated proton channel is tuned by cooperativity. Nat. Struct. Mol. Biol. 17, 44–50 (2010).
Sakata, S. et al. Functionality of the voltage-gated proton channel truncated in S4. Proc. Natl. Acad. Sci. USA 107, 2313–2318 (2010).
Jiang, Y. et al. X-ray structure of a voltage-dependent K+ channel. Nature 423, 33–41 (2003).
Long, S.B., Tao, X., Campbell, E.B. & MacKinnon, R. Atomic structure of a voltage-dependent K+ channel in a lipid membrane-like environment. Nature 450, 376–382 (2007).
Scott, K.A. et al. Coarse-grained MD simulations of membrane protein-bilayer self-assembly. Structure 16, 621–630 (2008).
Wolf, S., Bockmann, M., Howeler, U., Schlitter, J. & Gerwert, K. Simulations of a G protein-coupled receptor homology model predict dynamic features and a ligand binding site. FEBS Lett. 582, 3335–3342 (2008).
Musset, B. et al. Zinc inhibition of monomeric and dimeric proton channels suggests cooperative gating. J. Physiol. 588, 1435–1449 (2010).
Gonzalez, C., Koch, H., Drum, B. & Larsson, H.P. Strong cooperativity between subunits in voltage-gated proton channels. Nat. Struct. Mol. Biol. 17, 51–56 (2010).
Freites, J.A., Tobias, D.J. & White, S.H. A voltage-sensor water pore. Biophys. J. 91, L90–L92 (2006).
Wang, D. & Voth, G.A. Proton transport pathway in the ClC Cl−/H+ antiporter. Biophys. J. 97, 121–131 (2009).
Chen, H., Wu, Y. & Voth, G. Origins of proton transport behavior from selectivity domain mutations of the aquaporin-1 channel. Biophys. J. 90, L73 (2006).
Garczarek, F. & Gerwert, K. Functional waters in intraprotein proton transfer monitored by FTIR difference spectroscopy. Nature 439, 109–112 (2006).
Tiwari-Woodruff, S.K., Lin, M.A., Schulteis, C.T. & Papazian, D.M. Voltage-dependent structural interactions in the Shaker K+ channel. J. Gen. Physiol. 115, 123–138 (2000).
Decoursey, T.E. Voltage-gated proton channels and other proton transfer pathways. Physiol. Rev. 83, 475–579 (2003).
Alabi, A.A., Bahamonde, M.I., Jung, H.J., Kim, J.I. & Swartz, K.J. Portability of paddle motif function and pharmacology in voltage sensors. Nature 450, 370–375 (2007).
Blanchet, J., Pilote, S. & Chahine, M. Acidic residues on the voltage-sensor domain determine the activation of the NaChBac sodium channel. Biophys. J. 92, 3513–3523 (2007).
Tao, X. et al. A gating charge transfer center in voltage sensors. Science 328, 67–73 (2010).
Markovitch, O. et al. Special pair dance and partner selection: elementary steps in proton transport in liquid water. J. Phys. Chem. B 112, 9456–9466 (2008).
DeCoursey, T.E. & Cherny, V.V. Deuterium isotope effects on permeation and gating of proton channels in rat alveolar epithelium. J. Gen. Physiol. 109, 415–434 (1997).
Kuno, M. et al. Temperature dependence of proton permeation through a voltage-gated proton channel. J. Gen. Physiol. 134, 191–205 (2009).
Chen, H. et al. Charge delocalization in proton channels, I: the aquaporin channels and proton blockage. Biophys. J. 92, 46–60 (2007).
DeCoursey, T.E. & Cherny, V.V. Voltage-activated proton currents in membrane patches of rat alveolar epithelial cells. J. Physiol. (Lond.) 489, 299–307 (1995).
Sansom, M.S., Kerr, I.D., Smith, G.R. & Son, H.S. The influenza A virus M2 channel: a molecular modeling and simulation study. Virology 233, 163–173 (1997).
Katoh, K., Misawa, K., Kuma, K. & Miyata, T. MAFFT: a novel method for rapid multiple sequence alignment based on fast Fourier transform. Nucleic Acids Res. 30, 3059–3066 (2002).
Henikoff, S. & Henikoff, J.G. Amino acid substitution matrices from protein blocks. Proc. Natl. Acad. Sci. USA 89, 10915–10919 (1992).
Shi, J., Blundell, T.L. & Mizuguchi, K. FUGUE: sequence-structure homology recognition using environment-specific substitution tables and structure-dependent gap penalties. J. Mol. Biol. 310, 243–257 (2001).
Bond, P.J., Holyoake, J., Ivetac, A., Khalid, S. & Sansom, M.S.P. Coarse-grained molecular dynamics simulations of membrane proteins and peptides. J. Struct. Biol. 157, 593–605 (2007).
Marrink, S.J., de Vries, A.H. & Mark, A.E. Coarse grained model for semiquantitative lipid simulations. J. Phys. Chem. B 108, 750–760 (2004).
Shelley, J.C., Shelley, M.Y., Reeder, R.C., Bandyopadhyay, S. & Klein, M.L. A coarse grain model for phospholipid simulations. J. Phys. Chem. B 105, 4464–4470 (2001).
Bond, P.J. & Sansom, M.S.P. Insertion and assembly of membrane proteins via simulation. J. Am. Chem. Soc. 128, 2697–2704 (2006).
Bond, P.J., Wee, C.L. & Sansom, M.S. Coarse-grained molecular dynamics simulations of the energetics of helix insertion into a lipid bilayer. Biochemistry 47, 11321–11331 (2008).
Berendsen, H.J.C., van der Spoel, D. & van Drunen, R. GROMACS: A message-passing parallel molecular dynamics implementation. Comput. Phys. Commun. 91, 43–56 (1995).
Clayton, G.M. et al. Structure of the transmembrane regions of a bacterial cyclic nucleotide-regulated channel. Proc. Natl. Acad. Sci. USA 105, 1511–1515 (2008).
Atilgan, A.R. et al. Anisotropy of fluctuation dynamics of proteins with an elastic network model. Biophys. J. 80, 505–515 (2001).
Keskin, O., Jernigan, R.L. & Bahar, I. Proteins with similar architecture exhibit similar large-scale dynamic behavior. Biophys. J. 78, 2093–2106 (2000).
Sands, Z.A. & Sansom, M.S. How does a voltage sensor interact with a lipid bilayer? Simulations of a potassium channel domain. Structure 15, 235–244 (2007).
Berger, O., Edholm, O. & Jahnig, F. Molecular dynamics simulations of a fluid bilayer of dipalmitoylphosphatidycholine at full hydration, constant pressure and constant temperature. Biophys. J. 72, 2002–2013 (1997).
Van Der Spoel, D. et al. GROMACS: fast, flexible, and free. J. Comput. Chem. 26, 1701–1718 (2005).
van Gunsteren, W.F. et al. in Biomolecular Simulation: The GROMOS96 Manual and User Guide (Biomos & Hochschulverlag AG an der ETH Zurich, Groningen & Zurich, 1996).
Darden, T., York, D. & Pedersen, L. Particle mesh Ewald - an N.log(N) method for Ewald sums in large systems. J. Chem. Phys. 98, 10089–10092 (1993).
Nose, S. & Klein, M.L. Constant pressure molecular-dynamics for molecular-systems. Mol. Phys. 50, 1055–1076 (1983).
Parrinello, M. & Rahman, A. Polymorphic transitions in single-crystals - a new molecular-dynamics method. J. Appl. Phys. 52, 7182–7190 (1981).
Hess, B., Bekker, H., Berendsen, H.J.C. & Fraaije, J.G.E.M. LINCS: A linear constraint solver for molecular simulations. J. Comput. Chem. 18, 1463–1472 (1997).
Humphrey, W., Dalke, A. & Schulten, K. VMD—visual molecular dynamics. J. Mol. Graph. 14, 33–38 (1996).
Acknowledgements
We are grateful to J.W. Klingelhoefer for writing MATLAB script to calculate the water-count profiles, M.M. Moran and J.A. Chong for their invaluable support and critical insight and E. Ruchti for superb technical assistance. The Mental Retardation/Developmental Disabilities Research Center Molecular Genetics Core Facility at Children's Hospital is supported by US National Institutes of Health Grant P30-HD18655. Work in the Sansom laboratory is supported by grants from the UK Biotechnology and Biological Sciences Research Council and the Wellcome Trust.
Author information
Authors and Affiliations
Contributions
I.S.R. and I.C. designed experiments, created Hv1 point mutations and performed electrophysiological experiments; Y.M. and Z.A.S. created Hv1 models and performed molecular dynamics simulations; D.E.C. and M.S.P.S. directed research activities; I.S.R., Y.M., Z.A.S., M.S.P.S. and D.E.C. wrote the paper.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–5 and Supplementary Tables 1–3 (PDF 15136 kb)
Rights and permissions
About this article
Cite this article
Ramsey, I., Mokrab, Y., Carvacho, I. et al. An aqueous H+ permeation pathway in the voltage-gated proton channel Hv1. Nat Struct Mol Biol 17, 869–875 (2010). https://doi.org/10.1038/nsmb.1826
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nsmb.1826
This article is cited by
-
Transient water wires mediate selective proton transport in designed channel proteins
Nature Chemistry (2023)
-
Construction of angstrom-scale ion channels with versatile pore configurations and sizes by metal-organic frameworks
Nature Communications (2023)
-
Inhibiting Hv1 channel in peripheral sensory neurons attenuates chronic inflammatory pain and opioid side effects
Cell Research (2022)
-
Sperm ion channels and transporters in male fertility and infertility
Nature Reviews Urology (2021)
-
Thermodynamics and Mechanism of the Membrane Permeation of Hv1 Channel Blockers
The Journal of Membrane Biology (2021)