Abstract
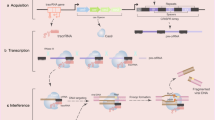
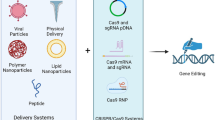
The CRISPR system, as an effective genome editing technology, has been extensively utilized for the construction of disease models in human pluripotent stem cells. Establishment of a gene mutant or knockout stem cell line typically relies on Cas nuclease-generated double-stranded DNA breaks and exogenous templates, which can produce uncontrollable editing byproducts and toxicity. The recently developed adenine base editors (ABE) have greatly facilitated related research by introducing A/T > G/C mutations in the coding regions or splitting sites (AG-GT) of genes, enabling mutant gene knock-in or knock-out without introducing DNA breaks. In this study, we edit the AG bases in exons anterior to achieve gene knockout via the ABE8e-SpRY, which recognizes most expanded protospacer adjacent motif to target the genome. Except for gene-knockout, ABE8e-SpRY can also efficiently establish disease-related A/T-to-G/C variation cell lines by targeting coding sequences. The method we generated is simple and time-saving, and it only takes two weeks to obtain the desired cell line. This protocol provides operating instructions step-by-step for constructing knockout and point mutation cell lines.
Graphical Abstract




Similar content being viewed by others
Code Availability
Not applicable.
Data Availability
Materials related to this paper is available from the corresponding author.
References
Takahashi, K., Tanabe, K., Ohnuki, M., Narita, M., Ichisaka, T., Tomoda, K., & Yamanaka, S. (2007). Induction of pluripotent stem cells from adult human fibroblasts by defined factors. Cell, 131(5), 861–872. https://doi.org/10.1016/j.cell.2007.11.019
Preman, P., Tcw, J., Calafate, S., Snellinx, A., Alfonso-Triguero, M., Corthout, N., Munck, S., Thal, D., Goate, A., De Strooper, B., & Arranz, A. (2021). Human iPSC-derived astrocytes transplanted into the mouse brain undergo morphological changes in response to amyloid-β plaques. Molecular neurodegeneration, 16(1), 68. https://doi.org/10.1186/s13024-021-00487-8
Takahashi, J. (2020). iPS cell-based therapy for Parkinson's disease: A Kyoto trial. Regenerative therapy, 13(18–22). https://doi.org/10.1016/j.reth.2020.06.002
Chang, Y., Li, Y., Bai, R., Wu, F., Ma, S., Saleem, A., Zhang, S., Jiang, Y., Dong, T., Guo, T., Hang, C., Lu, W., Jiang, H., & Lan, F. (2021). hERG-deficient human embryonic stem cell-derived cardiomyocytes for modelling QT prolongation. Stem cell research & therapy, 12(1), 278. https://doi.org/10.1186/s13287-021-02346-1
Jin, M., Xu, R., Wang, L., Alam, M., Ma, Z., Zhu, S., Martini, A., Jadali, A., Bernabucci, M., Xie, P., Kwan, K., Pang, Z., Head, E., Liu, Y., Hart, R., & Jiang, P. (2022). Type-I-interferon signaling drives microglial dysfunction and senescence in human iPSC models of Down syndrome and Alzheimer’s disease. Cell Stem Cell, 29(7), 1135-1153.e1138. https://doi.org/10.1016/j.stem.2022.06.007
Li, X., Lu, W., Li, Y., Wu, F., Bai, R., Ma, S., Dong, T., Zhang, H., Lee, A., Wang, Y., & Lan, F. (2019). MLP-deficient human pluripotent stem cell derived cardiomyocytes develop hypertrophic cardiomyopathy and heart failure phenotypes due to abnormal calcium handling. Cell death & disease, 10(8), 610. https://doi.org/10.1038/s41419-019-1826-4
Van der Kant, R., Langness, V., Herrera, C., Williams, D., Fong, L., Leestemaker, Y., Steenvoorden, E., Rynearson, K., Brouwers, J., Helms, J., Ovaa, H., Giera, M., Wagner, S., Bang, A., & Goldstein, L. (2019). Cholesterol Metabolism Is a Druggable Axis that Independently Regulates Tau and Amyloid-β in iPSC-Derived Alzheimer’s Disease Neurons. Cell Stem Cell, 24(3), 363-375.e369. https://doi.org/10.1016/j.stem.2018.12.013
Okano, H., Yasuda, D., Fujimori, K., Morimoto, S., & Takahashi, S. (2020). Ropinirole, a New ALS Drug Candidate Developed Using iPSCs. Trends in pharmacological sciences, 41(2), 99–109. https://doi.org/10.1016/j.tips.2019.12.002
Yahata, N., Asai, M., Kitaoka, S., Takahashi, K., Asaka, I., Hioki, H., Kaneko, T., Maruyama, K., Saido, T., Nakahata, T., Asada, T., Yamanaka, S., Iwata, N., & Inoue, H. (2011). Anti-Aβ drug screening platform using human iPS cell-derived neurons for the treatment of Alzheimer’s disease. PLoS ONE, 6(9), e25788. https://doi.org/10.1371/journal.pone.0025788
Ito, T., Kawai, Y., Yasui, Y., Iriguchi, S., Minagawa, A., Ishii, T., Miyoshi, H., Taketo, M., Kawada, K., Obama, K., Sakai, Y., & Kaneko, S. (2021). The therapeutic potential of multiclonal tumoricidal T cells derived from tumor infiltrating lymphocyte-1derived iPS cells. Communications biology, 4(1), 694. https://doi.org/10.1038/s42003-021-02195-x
Ueda, T., Kumagai, A., Iriguchi, S., Yasui, Y., Miyasaka, T., Nakagoshi, K., Nakane, K., Saito, K., Takahashi, M., Sasaki, A., Yoshida, S., Takasu, N., Seno, H., Uemura, Y., Tamada, K., Nakatsura, T., & Kaneko, S. (2020). Non-clinical efficacy, safety and stable clinical cell processing of induced pluripotent stem cell-derived anti-glypican-3 chimeric antigen receptor-expressing natural killer/innate lymphoid cells. Cancer science, 111(5), 1478–1490. https://doi.org/10.1111/cas.14374
Grskovic, M., Javaherian, A., Strulovici, B., & Daley, G. (2011). Induced pluripotent stem cells–opportunities for disease modelling and drug discovery. Nature reviews. Drug discovery, 10(12), 915–929. https://doi.org/10.1038/nrd3577
Krawczyk, E.& Kitlińska, J. (2023). Preclinical Models of Neuroblastoma-Current Status and Perspectives. Cancers 15(13). https://doi.org/10.3390/cancers15133314
Zhou, T., Benda, C., Dunzinger, S., Huang, Y., Ho, J., Yang, J., Wang, Y., Zhang, Y., Zhuang, Q., Li, Y., Bao, X., Tse, H., Grillari, J., Grillari-Voglauer, R., Pei, D., & Esteban, M. (2012). Generation of human induced pluripotent stem cells from urine samples. Nature protocols, 7(12), 2080–2089. https://doi.org/10.1038/nprot.2012.115
Zhang, X. (2013). Cellular reprogramming of human peripheral blood cells. Genomics, proteomics & bioinformatics, 11(5), 264–274. https://doi.org/10.1016/j.gpb.2013.09.001
Burridge, P., Matsa, E., Shukla, P., Lin, Z., Churko, J., Ebert, A., Lan, F., Diecke, S., Huber, B., Mordwinkin, N., Plews, J., Abilez, O., Cui, B., Gold, J., & Wu, J. (2014). Chemically defined generation of human cardiomyocytes. Nature methods, 11(8), 855–860. https://doi.org/10.1038/nmeth.2999
Liang, Z., He, Y., Tang, H., Li, J., Cai, J., & Liao, Y. (2023). Dedifferentiated fat cells: Current applications and future directions in regenerative medicine. Stem cell research & therapy, 14(1), 207. https://doi.org/10.1186/s13287-023-03399-0
Tani, S., Okada, H., Chung, U., Ohba, S.,& Hojo, H. (2021). The Progress of Stem Cell Technology for Skeletal Regeneration. International journal of molecular sciences, 22(3). https://doi.org/10.3390/ijms22031404
Escribá, R., Larrañaga-Moreira, J., Richaud-Patin, Y., Pourchet, L., Lazis, I., Jiménez-Delgado, S., Morillas-García, A., Ortiz-Genga, M., Ochoa, J., Carreras, D., Pérez, G., de la Pompa, J., Brugada, R., Monserrat, L., Barriales-Villa, R., & Raya, A. (2023). iPSC-Based Modeling of Variable Clinical Presentation in Hypertrophic Cardiomyopathy. Circulation research, 133(2), 108–119. https://doi.org/10.1161/circresaha.122.321951
Li, C., Chen, S., Zhou, Y., Zhao, Y., Liu, P.,& Cai, J. (2018). Application of induced pluripotent stem cell transplants: Autologous or allogeneic? Life sciences, 212(145–149). https://doi.org/10.1016/j.lfs.2018.09.057
Hook, G., Reinheckel, T., Ni, J., Wu, Z., Kindy, M., Peters, C., & Hook, V. (2022). Cathepsin B Gene Knockout Improves Behavioral Deficits and Reduces Pathology in Models of Neurologic Disorders. Pharmacological reviews, 74(3), 600–629. https://doi.org/10.1124/pharmrev.121.000527
Vermersch, E., Jouve, C., & Hulot, J. (2020). CRISPR/Cas9 gene-editing strategies in cardiovascular cells. Cardiovascular research, 116(5), 894–907. https://doi.org/10.1093/cvr/cvz250
Liu, N., & Olson, E. (2022). CRISPR Modeling and Correction of Cardiovascular Disease. Circulation research, 130(12), 1827–1850. https://doi.org/10.1161/circresaha.122.320496
Boutin, J., Cappellen, D., Rosier, J., Amintas, S., Dabernat, S., Bedel, A., & Moreau-Gaudry, F. (2022). ON-Target Adverse Events of CRISPR-Cas9 Nuclease: More Chaotic than Expected. The CRISPR journal, 5(1), 19–30. https://doi.org/10.1089/crispr.2021.0120
Ihry, R., Worringer, K., Salick, M., Frias, E., Ho, D., Theriault, K., Kommineni, S., Chen, J., Sondey, M., Ye, C., Randhawa, R., Kulkarni, T., Yang, Z., McAllister, G., Russ, C., Reece-Hoyes, J., Forrester, W., Hoffman, G., Dolmetsch, R., & Kaykas, A. (2018). p53 inhibits CRISPR-Cas9 engineering in human pluripotent stem cells. Nature medicine, 24(7), 939–946. https://doi.org/10.1038/s41591-018-0050-6
Mali, P., Yang, L., Esvelt, K., Aach, J., Guell, M., DiCarlo, J., Norville, J., & Church, G. (2013). RNA-guided human genome engineering via Cas9. Science (New York, N.Y.), 339(6121), 823–826. https://doi.org/10.1126/science.1232033
Park, J., Park, M., Lee, S., Kim, D., Kim, K., Jang, H., & Cha, H. (2023). Gene editing with “pencil” rather than “scissors” in human pluripotent stem cells. Stem cell research & therapy, 14(1), 164. https://doi.org/10.1186/s13287-023-03394-5
Komor, A., Kim, Y., Packer, M., Zuris, J., & Liu, D. (2016). Programmable editing of a target base in genomic DNA without double-stranded DNA cleavage. Nature, 533(7603), 420–424. https://doi.org/10.1038/nature17946
Porto, E., Komor, A., Slaymaker, I., & Yeo, G. (2020). Base editing: Advances and therapeutic opportunities. Nature reviews. Drug discovery, 19(12), 839–859. https://doi.org/10.1038/s41573-020-0084-6
Walton, R., Christie, K., Whittaker, M., & Kleinstiver, B. (2020). Unconstrained genome targeting with near-PAMless engineered CRISPR-Cas9 variants. Science (New York, N.Y.), 368(6488), 290–296. https://doi.org/10.1126/science.aba8853
Vicencio, J., Sánchez-Bolaños, C., Moreno-Sánchez, I., Brena, D., Vejnar, C., Kukhtar, D., Ruiz-López, M., Cots-Ponjoan, M., Rubio, A., Melero, N., Crespo-Cuadrado, J., Carolis, C., Pérez-Pulido, A., Giráldez, A., Kleinstiver, B., Cerón, J., & Moreno-Mateos, M. (2022). Genome editing in animals with minimal PAM CRISPR-Cas9 enzymes. Nature communications, 13(1), 2601. https://doi.org/10.1038/s41467-022-30228-4
Li, J., Xu, R., Qin, R., Liu, X., Kong, F., & Wei, P. (2021). Genome editing mediated by SpCas9 variants with broad non-canonical PAM compatibility in plants. Molecular plant, 14(2), 352–360. https://doi.org/10.1016/j.molp.2020.12.017
Ren, Q., Sretenovic, S., Liu, S., Tang, X., Huang, L., He, Y., Liu, L., Guo, Y., Zhong, Z., Liu, G., Cheng, Y., Zheng, X., Pan, C., Yin, D., Zhang, Y., Li, W., Qi, L., Li, C., Qi, Y., & Zhang, Y. (2021). PAM-less plant genome editing using a CRISPR-SpRY toolbox. Nature plants, 7(1), 25–33. https://doi.org/10.1038/s41477-020-00827-4
Xu, Z., Kuang, Y., Ren, B., Yan, D., Yan, F., Spetz, C., Sun, W., Wang, G., Zhou, X., & Zhou, H. (2021). SpRY greatly expands the genome editing scope in rice with highly flexible PAM recognition. Genome biology, 22(1), 6. https://doi.org/10.1186/s13059-020-02231-9
Zhang, C., Wang, Y., Wang, F., Zhao, S., Song, J., Feng, F., Zhao, J., & Yang, J. (2021). Expanding base editing scope to near-PAMless with engineered CRISPR/Cas9 variants in plants. Molecular plant, 14(2), 191–194. https://doi.org/10.1016/j.molp.2020.12.016
Asano, Y., Yamashita, K., Hasegawa, A., Ogasawara, T., Iriki, H., & Muramoto, T. (2021). Knock-in and precise nucleotide substitution using near-PAMless engineered Cas9 variants in Dictyostelium discoideum. Scientific reports, 11(1), 11163. https://doi.org/10.1038/s41598-021-89546-0
Evans, B.& Bernstein, D. (2021). SpRY Cas9 Can Utilize a Variety of Protospacer Adjacent Motif Site Sequences To Edit the Candida albicans Genome. mSphere, 6(3), https://doi.org/10.1128/mSphere.00303-21
Alves, C., Ha, L., Yaworski, R., Sutton, E., Lazzarotto, C., Christie, K., Reilly, A., Beauvais, A., Doll, R., de la Cruz, D., Maguire, C., Swoboda, K., Tsai, S., Kothary, R., & Kleinstiver, B. (2024). Optimization of base editors for the functional correction of SMN2 as a treatment for spinal muscular atrophy. Nature biomedical engineering, 8(2), 118–131. https://doi.org/10.1038/s41551-023-01132-z
Berget, S., Moore, C., & Sharp, P. (1977). Spliced segments at the 5’ terminus of adenovirus 2 late mRNA. Proceedings of the National Academy of Sciences of the United States of America, 74(8), 3171–3175. https://doi.org/10.1073/pnas.74.8.3171
Sharp, P. (2005). The discovery of split genes and RNA splicing. Trends in biochemical sciences, 30(6), 279–281. https://doi.org/10.1016/j.tibs.2005.04.002
Musunuru, K., Chadwick, A., Mizoguchi, T., Garcia, S., DeNizio, J., Reiss, C., Wang, K., Iyer, S., Dutta, C., Clendaniel, V., Amaonye, M., Beach, A., Berth, K., Biswas, S., Braun, M., Chen, H., Colace, T., Ganey, J., Gangopadhyay, S., … Kathiresan, S. (2021). In vivo CRISPR base editing of PCSK9 durably lowers cholesterol in primates. Nature, 593(7859), 429–434. https://doi.org/10.1038/s41586-021-03534-y
Li, L., Cao, Y., Zhao, F., Mao, B., Ren, X., Wang, Y., Guan, Y., You, Y., Li, S., Yang, T.,& Zhao, X. (2019). Validation and Classification of Atypical Splicing Variants Associated With Osteogenesis Imperfecta. Frontiers in genetics, 10(979). https://doi.org/10.3389/fgene.2019.00979
Liu, M., Cardilla, A., Ngeow, J., Gong, X.,& Xia, Y. (2022). Studying Kidney Diseases Using Organoid Models. Frontiers in cell and developmental biology, 10(845401). https://doi.org/10.3389/fcell.2022.845401
Saleem, A., Abbas, M., Wang, Y., & Lan, F. (2022). hPSC gene editing for cardiac disease therapy. Pflugers Archiv : European journal of physiology, 474(11), 1123–1132. https://doi.org/10.1007/s00424-022-02751-2
Zhou, Z., Ma, X., & Zhu, S. (2020). Recent advances and potential applications of human pluripotent stem cell-derived pancreatic β cells. Acta biochimica et biophysica Sinica, 52(7), 708–715. https://doi.org/10.1093/abbs/gmaa047
Qi, T., Wu, F., Xie, Y., Gao, S., Li, M., Pu, J., Li, D., Lan, F.,& Wang, Y. (2020). Base Editing Mediated Generation of Point Mutations Into Human Pluripotent Stem Cells for Modeling Disease. Frontiers in cell and developmental biology, 8(590581). https://doi.org/10.3389/fcell.2020.590581
Xie, G., Lin, S., Wu, F.& Liu, J. (2023). Nanomaterial-based ophthalmic drug delivery. Advanced drug delivery reviews, 200(115004). https://doi.org/10.1016/j.addr.2023.115004
Nandy, K., Babu, D., Rani, S., Joshi, G., Ijee, S., George, A., Palani, D., Premkumar, C., Rajesh, P., Vijayanand, S., David, E., Murugesan, M., & Velayudhan, S. (2023). Efficient gene editing in induced pluripotent stem cells enabled by an inducible adenine base editor with tunable expression. Scientific reports, 13(1), 21953. https://doi.org/10.1038/s41598-023-42174-2
Wang, J., Zhu, H., Gan, J., Liang, G., Li, L.,& Zhao, Y. (2023). Engineered mRNA Delivery Systems for Biomedical Applications. Advanced materials (Deerfield Beach, Fla.), e2308029. https://doi.org/10.1002/adma.202308029
Van der Wiel, I., Cheng, J., Koukiekolo, R., Lyn, R., Stevens, N., O’Connor, N., Turro, N., & Pezacki, J. (2009). FLEth RNA intercalating probe is a convenient reporter for small interfering RNAs. Journal of the American Chemical Society, 131(29), 9872–9873. https://doi.org/10.1021/ja902636m
Funding
We gratefully acknowledge funding support from the National Natural Science Foundation of China (82070258, 81970205), Shenzhen Fundamental Research Program (ZDSYS20200923172000001), China Postdoctoral Science Foundation (2022M720498).
Author information
Authors and Affiliations
Contributions
MSH and LF designed this study; CY and ZYS performed the majority of cell experiments and data analysis; CY and ZYS provided the manuscript preparation. All authors read and approved the final manuscript.
Corresponding author
Ethics declarations
Ethics Approval
Not applicable.
Consent for Publication
Not applicable.
Consent to Participate
Not applicable.
Competing Interests
The authors indicate no potential conflicts of interest.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Chang, Y., Lan, F., Zhang, Y. et al. Crispr-Based Editing of Human Pluripotent Stem Cells for Disease Modeling. Stem Cell Rev and Rep (2024). https://doi.org/10.1007/s12015-024-10713-7
Accepted:
Published:
DOI: https://doi.org/10.1007/s12015-024-10713-7