Abstract
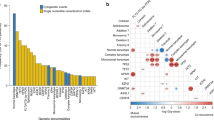
The mutational spectrum and molecular characteristics of acute myelomonocytic lineage leukemia, namely acute myeloid leukemia (AML) French–American–British (FAB) subtypes M4 and M5, are largely unknown. In order to explore the mutational spectrum and prognostic factors of FAB-M4 and -M5, next-generation sequencing (NGS) was performed to screen for mutated genes and fusion genes relevant to the pathogenesis of AML. Of the 63 patients enrolled in the study, 60% had more than three mutated genes. NPM1 had the highest mutation frequency, followed by DNMT3A, FLT3, NRAS, RUNX1, and TET2. Univariate analysis suggested that age ≥60 years was an independent factor for both poor event-free survival (EFS) and overall survival (OS, P = 0.009, 0.002, respectively), MYH11-CBFβ was associated with better EFS and OS (P = 0.029, 0.016, respectively). However, multivariate analysis was not able to identify any independent risk factor for survival in the cohort of FAB-M4 and -M5 patients, including peripheral white blood cell count, bone marrow blast percentage, MYH11-CBFβ, FLT3-ITD, mutations in NPM1 and DNMT3A, and allogeneic hematopoietic stem cell transplantation (allo-HSCT). Our study provided new insight into the mutational spectrum and molecular characteristics of FAB-M4 and -M5. The clinical implications of the genetic signature of FAB-M4 and -M5 need to be further elucidated by larger studies.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout


Similar content being viewed by others
References
Schlenk RF, Döhner H. Genomic applications in the clinic: use in treatment paradigm of acute myeloid leukemia. Hematol Am Soc Hematol Educ Program. 2013;2013:324–30.
Cazzola M, Della Porta MG, Travaglino E, Malcovati L. Classification and prognostic evaluation of myelodysplastic syndromes. Semin Oncol. 2011;38:627–34.
Kohlmann A, Grossmann V, Nadarajah N, Haferlach T. Next-generation sequencing - feasibility and practicality in haematology. Br J Haematol. 2013;160:736–53.
Garraway LA. Genomics-driven oncology: framework for an emerging paradigm. J Clin Oncol. 2013;31:1806–14.
Cancer Genome Atlas Research Network. Genomic and epigenomic landscapes of adult de novo acute myeloid leukemia. New Engl J Med. 2013;368:2059–74.
Papaemmanuil E, Gerstung M, Bullinger L, Gaidzik VI, Paschka P, Roberts ND, et al. Genomic classification and prognosis in acute myeloid leukemia. New Engl J Med. 2016;374:2209–21.
Patel JP, Gönen M, Figueroa ME, Fernandez H, Sun Z, Racevskis J, et al. Prognostic relevance of integrated genetic profiling in acute myeloid leukemia. New Engl J Med. 2012;366:1079–89.
Ofran Y, Rowe JM. Genetic profiling in acute myeloid leukaemia--where are we and what is its role in patient management. Br J Haematol. 2013;160:303–20.
Bennett JM, Catovsky D, Daniel MT, Flandrin G, Galton DA, Gralnick HR, et al. Proposal for the recognition of minimally differentiated acute myeloid leukaemia (AML-M0). Br J Haematol. 1991;78:325–9.
Ley TJ, Miller C, Ding L, Raphael BJ, Mungall AJ, Robertson A, et al. Genomic and epigenomic landscapes of adult de novo acute myeloid leukemia. New Engl J Med. 2013;368:2059–74.
Mardis ER, Ding L, Dooling DJ, Larson DE, McLellan MD, Chen K, et al. Recurring mutations found by sequencing an acute myeloid leukemia genome. New Engl J Med. 2009;361:1058–66.
Estey EH. Acute myeloid leukemia: 2013 update on risk-stratification and management. Am J Hematol. 2013;88:318–27.
Naoe T, Kiyoi H. Gene mutations of acute myeloid leukemia in the genome era. Int J Hematol. 2013;97:165–74.
Grossmann V, Schnittger S, Kohlmann A, Eder C, Roller A, Dicker F, et al. A novel hierarchical prognostic model of AML solely based on molecular mutations. Blood. 2012;120:2963–72.
Walter MJ, Ding L, Shen D, Shao J, Grillot M, McLellan M, et al. Recurrent DNMT3A mutations in patients with myelodysplastic syndromes. Leukemia. 2011;25:1153–8.
Ganguly BB, Kadam NN. Mutations of myelodysplastic syndromes (MDS): an update. Mutat Res/Rev Mutat Res. 2016;769:47–62.
Papaemmanuil E, Gerstung M, Malcovati L, Tauro S, Gundem G, Van Loo P, et al. Clinical and biological implications of driver mutations in myelodysplastic syndromes. Blood. 2013;122:3616–27.
Haferlach T, Nagata Y, Grossmann V, Okuno Y, Bacher U, Nagae G, et al. Landscape of genetic lesions in 944 patients with myelodysplastic syndromes. Leukemia. 2014;28:241–7.
Kurosawa S, Miyawaki S, Yamaguchi T, Kanamori H, Sakura T, Moriuchi Y, et al. Prognosis of patients with core binding factor acute myeloid leukemia after first relapse. Haematologica. 2013;98:1525–31.
Döhner H, Weisdorf DJ, Bloomfield CD. Acute myeloid leukemia. New Engl J Med. 2015;373:1136–52.
Acknowledgements
This work was supported by grants from the National Natural Science Foundation of China (81500118, 61501519), the China Postdoctoral Science Foundation funded project (project No.2016M600443), PLAGH project of Medical Big Data (Project No.2016MBD-025) and Jiangsu Province Postdoctoral Science Foundation funded project (project No.1701184B).
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Rights and permissions
About this article
Cite this article
Cheng, Z., Hu, K., Tian, L. et al. Clinical and biological implications of mutational spectrum in acute myeloid leukemia of FAB subtypes M4 and M5. Cancer Gene Ther 25, 77–83 (2018). https://doi.org/10.1038/s41417-018-0013-6
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41417-018-0013-6