Abstract
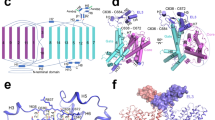
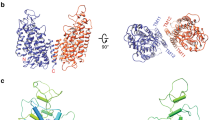
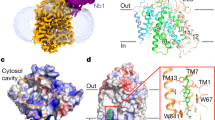
Urea is highly concentrated in the mammalian kidney to produce the osmotic gradient necessary for water re-absorption. Free diffusion of urea across cell membranes is slow owing to its high polarity, and specialized urea transporters have evolved to achieve rapid and selective urea permeation. Here we present the 2.3 Å structure of a functional urea transporter from the bacterium Desulfovibrio vulgaris. The transporter is a homotrimer, and each subunit contains a continuous membrane-spanning pore formed by the two homologous halves of the protein. The pore contains a constricted selectivity filter that can accommodate several dehydrated urea molecules in single file. Backbone and side-chain oxygen atoms provide continuous coordination of urea as it progresses through the filter, and well-placed α-helix dipoles provide further compensation for dehydration energy. These results establish that the urea transporter operates by a channel-like mechanism and reveal the physical and chemical basis of urea selectivity.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Sebbane, F. et al. The Yersinia pseudotuberculosis Yut protein, a new type of urea transporter homologous to eukaryotic channels and functionally interchangeable in vitro with the Helicobacter pylori UreI protein. Mol. Microbiol. 45, 1165–1174 (2002)
Weeks, D. L., Eskandari, S., Scott, D. R. & Sachs, G. A H+-gated urea channel: the link between Helicobacter pylori urease and gastric colonization. Science 287, 482–485 (2000)
Hediger, M. A. et al. Structure, regulation and physiological roles of urea transporters. Kidney Int. 49, 1615–1623 (1996)
Sands, J. M. Mammalian urea transporters. Annu. Rev. Physiol. 65, 543–566 (2003)
Bagnasco, S. M. Role and regulation of urea transporters. Pflugers Arch. 450, 217–226 (2005)
Finkelstein, A. Water and nonelectrolyte permeability of lipid bilayer membranes. J. Gen. Physiol. 68, 127–135 (1976)
Valladares, A., Montesinos, M. L., Herrero, A. & Flores, E. An ABC-type, high-affinity urea permease identified in cyanobacteria. Mol. Microbiol. 43, 703–715 (2002)
Kojima, S., Bohner, A. & von Wiren, N. Molecular mechanisms of urea transport in plants. J. Membr. Biol. 212, 83–91 (2006)
You, G. et al. Cloning and characterization of the vasopressin-regulated urea transporter. Nature 365, 844–847 (1993)
MacIver, B., Smith, C. P., Hill, W. G. & Zeidel, M. L. Functional characterization of mouse urea transporters UT-A2 and UT-A3 expressed in purified Xenopus laevis oocyte plasma membranes. Am. J. Physiol. Renal Physiol. 294, F956–F964 (2008)
Raunser, S. et al. Oligomeric structure and functional characterization of the urea transporter from Actinobacillus pleuropneumoniae . J. Mol. Biol. 387, 619–627 (2009)
Zhao, D., Sonawane, N. D., Levin, M. H. & Yang, B. Comparative transport efficiencies of urea analogues through urea transporter UT-B. Biochim. Biophys. Acta 1768, 1815–1821 (2007)
Mannuzzu, L. M., Moronne, M. M. & Macey, R. I. Estimate of the number of urea transport sites in erythrocyte ghosts using a hydrophobic mercurial. J. Membr. Biol. 133, 85–97 (1993)
Minocha, R., Studley, K. & Saier, M. H. The urea transporter (UT) family: bioinformatic analyses leading to structural, functional, and evolutionary predictions. Receptors Channels 9, 345–352 (2003)
Chou, C. L. & Knepper, M. A. Inhibition of urea transport in inner medullary collecting duct by phloretin and urea analogues. Am. J. Physiol. 257, F359–F365 (1989)
Quick, M. & Javitch, J. A. Monitoring the function of membrane transport proteins in detergent-solubilized form. Proc. Natl Acad. Sci. USA 104, 3603–3608 (2007)
Shi, L., Quick, M., Zhao, Y., Weinstein, H. & Javitch, J. A. The mechanism of a neurotransmitter:sodium symporter—inward release of Na+ and substrate is triggered by substrate in a second binding site. Mol. Cell 30, 667–677 (2008)
von Heijne, G. & Gavel, Y. Topogenic signals in integral membrane proteins. Eur. J. Biochem. 174, 671–678 (1988)
Shayakul, C., Steel, A. & Hediger, M. A. Molecular cloning and characterization of the vasopressin-regulated urea transporter of rat kidney collecting ducts. J. Clin. Invest. 98, 2580–2587 (1996)
Bradford, A. D. et al. 97- and 117-kDa forms of collecting duct urea transporter UT-A1 are due to different states of glycosylation. Am. J. Physiol. Renal Physiol. 281, F133–F143 (2001)
Lucien, N. et al. Antigenic and functional properties of the human red blood cell urea transporter hUT-B1. J. Biol. Chem. 277, 34101–34108 (2002)
Yang, B. & Verkman, A. S. Analysis of double knockout mice lacking aquaporin-1 and urea transporter UT-B. Evidence for UT-B-facilitated water transport in erythrocytes. J. Biol. Chem. 277, 36782–36786 (2002)
Imai, Y. N., Inoue, Y., Nakanishi, I. & Kitaura, K. Amide–π interactions between formamide and benzene. J. Comput. Chem. 30, 2267–2276 (2009)
Doyle, D. A. et al. The structure of the potassium channel: molecular basis of K+ conduction and selectivity. Science 280, 69–77 (1998)
Dutzler, R., Campbell, E. B., Cadene, M., Chait, B. T. & MacKinnon, R. X-ray structure of a ClC chloride channel at 3.0 Å reveals the molecular basis of anion selectivity. Nature 415, 287–294 (2002)
Terris, J. M., Knepper, M. A. & Wade, J. B. UT-A3: localization and characterization of an additional urea transporter isoform in the IMCD. Am. J. Physiol. Renal Physiol. 280, F325–F332 (2001)
Blount, M. A., Klein, J. D., Martin, C. F., Tchapyjnikov, D. & Sands, J. M. Forskolin stimulates phosphorylation and membrane accumulation of UT-A3. Am. J. Physiol. Renal Physiol. 293, F1308–F1313 (2007)
Khademi, S. et al. Mechanism of ammonia transport by Amt/MEP/Rh: structure of AmtB at 1.35 Å. Science 305, 1587–1594 (2004)
Zheng, L., Kostrewa, D., Berneche, S., Winkler, F. K. & Li, X. D. The mechanism of ammonia transport based on the crystal structure of AmtB of Escherichia coli . Proc. Natl Acad. Sci. USA 101, 17090–17095 (2004)
Murata, K. et al. Structural determinants of water permeation through aquaporin-1. Nature 407, 599–605 (2000)
Sui, H., Han, B. G., Lee, J. K., Walian, P. & Jap, B. K. Structural basis of water-specific transport through the AQP1 water channel. Nature 414, 872–878 (2001)
Fu, D. et al. Structure of a glycerol-conducting channel and the basis for its selectivity. Science 290, 481–486 (2000)
Levin, M. H., de la Fuente, R. & Verkman, A. S. Urearetics: a small molecule screen yields nanomolar potency inhibitors of urea transporter UT-B. FASEB J. 21, 551–563 (2007)
Shayakul, C. & Hediger, M. A. The SLC14 gene family of urea transporters. Pflugers Arch. 447, 603–609 (2004)
Zhang, C., Sands, J. M. & Klein, J. D. Vasopressin rapidly increases phosphorylation of UT-A1 urea transporter in rat IMCDs through PKA. Am. J. Physiol. Renal Physiol. 282, F85–F90 (2002)
Punta, M. et al. Structural genomics target selection for the New York Consortium on Membrane Protein Structure. J. Struct. Funct. Genomics 10.1007/s10969-009-9071-1 (in press)
Otwinowski, Z. & Minor, W. Processing of X-ray diffraction data collected in oscillation mode. Methods Enzymol. 276, 307–326 (1997)
Pape, T. & Schneider, T. R. HKL2MAP: a graphical user interface for macromolecular phasing with SHELX. J. Appl. Crystallogr. 37, 843–844 (2004)
Terwilliger, T. C. & Berendzen, J. Automated MAD and MIR structure solution. Acta Crystallogr. D 55, 849–861 (1999)
McCoy, A. J. et al. Phaser crystallographic software. J. Appl. Crystallogr. 40, 658–674 (2007)
Emsley, P. & Cowtan, K. Coot: model-building tools for molecular graphics. Acta Crystallogr. D 60, 2126–2132 (2004)
Collaborative Computational Project number 4 The CCP4 suite: programs for protein cystallography. Acta Crystallogr. D 50, 760–763 (1994)
Potterton, L. et al. Developments in the CCP4 molecular-graphics project. Acta Crystallogr. D 60, 2288–2294 (2004)
Lebedev, A. A., Vagin, A. A. & Murshudov, G. N. Model preparation in MOLREP and examples of model improvement using X-ray data. Acta Crystallogr. D 64, 33–39 (2008)
Vagin, A. A. et al. REFMAC5 dictionary: organization of prior chemical knowledge and guidelines for its use. Acta Crystallogr. D 60, 2184–2195 (2004)
Winn, M. D., Isupov, M. N. & Murshudov, G. N. Use of TLS parameters to model anisotropic displacements in macromolecular refinement. Acta Crystallogr. D 57, 122–133 (2001)
Brünger, A. T. et al. Crystallography & NMR system: A new software suite for macromolecular structure determination. Acta Crystallogr. D 54, 905–921 (1998)
Acknowledgements
We thank R. MacKinnon for advice and support throughout the project, C. Miller and E. Gouaux for comments on the manuscript, J. Love and NYCOMPS for cloning and the initial screen of protein expression levels, Y. Pan for messenger RNA preparation and oocyte injection, and J. Weng and Y. Cao for crystal screening and data collection at the synchrotrons. Data for this study were measured at beamlines X4A, X4C, X25 and X29 of the NSLS and the NE-CAT 24ID-E at the Advanced Photon Source. This work was supported by the US National Institutes of Health (HL086392 to M.Z. and T32HL087745 to E.J.L.), the NYCOMPS that is supported by the NIH Protein Structure Initiatives PSI-II (GM075026 to W. A. Hendrickson), and the American Heart Association (0630148N to M.Z.). M.Z. is a Pew Scholar in Biomedical Sciences.
Author Contributions E.J.L. and M.Z. conceived and designed the experiments. E.J.L. purified and crystallized the protein; M.Q. performed and analysed the radiotracer flux and SPA binding assays; E.J.L. and M.Z. collected and processed the X-ray data, solved the structure, and wrote the paper.
Author information
Authors and Affiliations
Corresponding author
Supplementary information
Supplementary Information
This file contains Supplementary Table 1 and Supplementary Figures S1-S6 with Legends. (PDF 9066 kb)
Rights and permissions
About this article
Cite this article
Levin, E., Quick, M. & Zhou, M. Crystal structure of a bacterial homologue of the kidney urea transporter . Nature 462, 757–761 (2009). https://doi.org/10.1038/nature08558
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature08558
This article is cited by
-
A critical analysis of urea transporter B inhibitors: molecular fingerprints, pharmacophore features for the development of next-generation diuretics
Molecular Diversity (2022)
-
Urea-aromatic interactions in biology
Biophysical Reviews (2020)
-
SLC14A1: a novel target for human urothelial cancer
Clinical and Translational Oncology (2017)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.