Abstract
Oceanic methane from global deep-sea sediment is largely consumed through microbially mediated sulfate-coupled oxidation, resulting in 13C-depleted cell biomass of anaerobic methanotrophic archaea (ANME). The general ecological importance of subseafloor ANME has been well recognized in the last two decades. However, the crucial biochemical pathways for the overall anaerobic oxidation of methane (AOM) still remain enigmatic. Here, methanotrophic pathways were analyzed to trace 13C-depleted amino acid biosynthesis in two clades of ANME (ANME-1 and ANME-2) from the Black Sea. Compound-specific analysis of ANME-dominated microbial mats showed a significant 13C-depletion trend in association with increasing carbon numbers in protein-derived amino acid families (e.g., the pyruvate family in the order of alanine, valine, isoleucine and leucine was down to −114‰). This result indicates a stepwise elongation of 13C-depleted carbon during amino acid biosynthesis. The overall results suggest that intracellular protein amino acids and the most 13C-depleted signature of leucine, which has a specific branched-chain structure, are potentially propagated as isoprenoid precursor molecules into archaeal biosynthesis, resulting in the extremely 13C- and 14C-depleted nature of ANME cells in the deep microbial oasis.
Similar content being viewed by others
Introduction
Microorganisms play a central role in both methane production and consumption in the global carbon cycle. The anaerobic oxidation of methane (AOM) is an important microbial process that controls the release of greenhouse gas from oceanic sediment1. Since the discovery of extremely 13C-depleted lipids produced by modern anaerobic methanotrophic archaea (ANME) in deep-sea sediments2,3,4, we have recognized the 13C-depleted isotopic signatures (ca. −110‰) as an ongoing AOM process from biogeochemical models for lipid biomarker records5,6, even in hydrothermally active seafloor settings7,8. Three phylogenetic groups of anaerobic methanotrophic archaea (ANME) have currently been identified, namely, ANME-1 (with subgroups a and b), ANME-2 (with subgroups a, b, c, and d), and ANME-3, which mediate AOM via sulfate (CH4 + SO42− → HCO3− + HS− + H2O)9, nitrate10, iron, and manganese11. AOM requires methyl coenzyme M reductase12,13, which catalyzes anaerobic methanotrophy through reverse methanogenesis14,15,16. Because obtaining pure cultures in the laboratory and isolating ANME are difficult, the biochemical mechanisms that control the AOM process, especially the pathways leading to 13C-depleted cell biomass, remain largely unknown. To better define the AOM process by focusing on biogeochemistry, we investigated the carbon isotopic composition of amino acids, the fundamental building blocks of proteins, in ANME-1- and ANME-2-dominated mats collected from the northwestern Black Sea (Fig. 1). ANME-1 and ANME-2 are observed in tall reef-like chimney structures (up to ca. 5 m height, 1 m diameter) composed of carbonates and dense microbial biomass (<1010 cells cm−3) (ref.17) where the methane seep rises vertically through the porous calcified interior. Therefore, we conducted compound-specific carbon isotope (13C/12C) analysis of 10 individual amino acids in the form of N-pivaloyl isopropyl ester derivatives and archaeal isoprenoid lipids together with radiocarbon (14C/12C) analysis of ANME cell using an accelerator mass spectrometry.
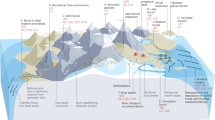
ANME-dominated microbial mats from the Black Sea. (a) Benthic methane seep environment of the Black Sea. The earliest description of the Black Sea mats and carbonate chimneys was reported by Luth and coworkers44. Please see the Supplementary Movie S1 for methane venting from carbonate chimneys. (b) Interior and exterior sections of chimney structure habitat location (image capture during the expedition) showing a pink mat (ANME-1-dominated), black mat, and carbonate precipitate (ANME-2-dominated)30. Photo credit: R/V Meteor cruise M72/1 science party (taken by M. Krüger). In the northwestern Black Sea at the Ukrainian shelf and slope area, a number of active gas seeps (at least 2778 sites)45 occur along the shelf edge near the Crimean Peninsula46,47,48,49,50,51,52. (c) The community structures of the ANME samples were determined by methyl coenzyme M reductase A (mcrA) gene-based clone analyses of the pink mat, black mat, and carbonate samples. ANME-1 and ANME-2 dominate the pink and black sections, respectively. Highly pure ANME-1 was observed in the pink section (Supplementary Fig. S4), whereas there was some diversity in the black and carbonate samples (Supplementary Figs S5 and S6). Based on the lipid analysis of ANME-1, the relative abundance of archaeal lipids was >98% (Supplementary Fig. S4). The isotopic composition of carbon (δ13C, ‰ vs. VPDB) and nitrogen (δ15N, ‰ vs. Air) and the radiocarbon (Δ14C, ‰) data for bulk ANME cell biomass are shown in Fig. 4.
The carbon isotope chemistry of ANME-dominated microbial mats
Mats from this region provide ideal natural enrichment for the study of methane biogeochemistry and microbial anaerobic methanotrophy mediated by modern ANME communities (Supplementary Fig. S1)18,19,20,21. To date, the AOM rate in the microbial reefs of the Black Sea is the fastest observed, with an estimated range of 103–104 nmol cm−3 day−1 (cf. Supplementary Fig. S2)17,21. We observed substantial 13C depletion in the amino acids from the ANME samples (Fig. 2), as low as to −114‰ relative to the Vienna Pee Dee Belemnite (VPDB) international standard (Fig. 3). Assuming the carbon isotopic composition of the substrate methane in the same area to be −50 to −65‰21,22, the isotopic fractionation by the ANME communities associated with amino acid biosynthesis was estimated to be 50–60‰. Methanotrophy by ANME-1 reportedly includes the formation of functionalized one-carbon (C1) compounds, such as methanol, methylamine, and methyl sulfide, from initial methane uptake16. Dual stable isotope probing (D- & 13C-) experiments also suggested that autotrophic carbon fixation from dissolved inorganic carbon (C1) occurred in the ANME-1 community23. Given that amino acid biosynthetic precursors include de novo pathway of pyruvate, phosphoglyceric acid, aspartic acid, α-ketoglutarate, and phosphoenolpyruvate + erythrose-4-phosphate24, we expected biosynthetic amino acids with additional 12C-carbon elongation to produce more 13C-depleted carbon isotopic signatures in ANME communities.
Gas chromatographic separation of 13C-depleted amino acids from ANME-1. Please see the analytical accuracy in Supplementary Fig. S3 and Table S1. Abbreviations: Ala, alanine (underivatized formula, C3H7NO2); Gly, glycine (C2H5NO2); Val, valine (C5H11NO2); Leu, leucine (C6H13NO2); Ile, isoleucine (C6H13NO2); Thr, threonine (C4H9NO3); Asp, aspartic acid (C4H7NO4) and asparagine after hydrolysis; Glu, glutamic acid (C5H9NO4) and glutamine after hydrolysis; Phe, phenylalanine (C9H11NO2); Tyr, tyrosine (C9H11NO3).
δ13C values of amino acids and lipids extracted from ANME mats in the Black Sea. Chemical structure and 13C-depletion of neutral amino acids glycine, alanine, valine, isoleucine, and leucine with carbon numbers (Cn) up to C6. The asterisks (*) represent pyruvate amino acid family members. Abbreviations: PGA, phosphoglyceric acid; Pyr, pyruvate; A, aspartic acid; αkg, α-ketoglutarate; PEP + E4P, phosphoenolpyruvate + erythrose-4-phosphate. The carbon isotopic composition of the ANME-2-dominated black mat and carbonate included the major archaeal C20 isoprenoid (<−116‰, vs. VPDB) (Supplementary Table S1).
Branched-chain amino acids in ANME
In the central metabolic pathways of ANME archaea, the pyruvate family includes four major amino acids: alanine, valine, isoleucine, and leucine16. Our data showed a stepwise 13C-depletion trend associated with the carbon numbers of neutral amino acids containing monoamino and monocarboxylic functional groups (Fig. 3), such as glycine (C2), alanine (C3), valine (C5), isoleucine (C6), and leucine (C6) (R2 = 0.91 and R2 = 0.80 in ANME-1 and ANME-2, respectively). Based on the metagenomic analysis of ANME-1, pyruvate serves as the biosynthetic precursor for L-valine (from 2-keto-isovalerate), L-leucine (2-keto-isocaproate), and L-isoleucine (2-keto-3-methylvaletrate), which are synthesized de novo (Fig. 2)16. Valine is synthesized by the addition of a 13C-depleted acetyl group to the alanine carbon skeleton24, resulting in a 13C-depleted biosynthetic flow that eventually progresses to leucine. In contrast, chorismate formed via the shikimate pathway is a precursor for aromatic amino acids with carbon isotopic compositions comparable to phenylalanine and tyrosine in ANME. The formation of tyrosine by enzymatic hydroxylation is the prephenate dominant metabolic pathway16. Of the aspartate amino acid family members, aspartic acid (C4; mixed signal with hydrolyzed asparagine) and threonine (C4) showed consistent carbon isotopic trends in both the ANME-1 and ANME-2 communities (Fig. 3).
The Δ14C values of ANME-1 (−815.3 ± 1.4‰), ANME-2 (−770.4 ± 1.8‰), and carbonate (−855.9 ± 1.6‰) indicated higher 14C depletion than in ambient seawater (Fig. 4). The δ13C and Δ14C cross-plot clearly indicated that venting methane was used directly by ANME cell biomass in the benthic seep chimney. The carbon isotopic order of the present ANME-1 cell biomass was compared with that of oceanic photoautotrophic primary producers because both are end-members in the oceanic carbon cycle (Fig. 5). The carbon isotopic composition of amino acids in representative oceanic photoautotrophs, such as phytoplankton and diatoms25, showed a similar trend to those of neutral amino acids (C2-glycine >C3-alanine >C5-valine >C6-isoleucine >C6-leucine), whereas the ANME communities in this study exhibited a wider carbon isotopic discrimination of the amino acid range than the reference photoautotrophs. Importantly, in both cases, the C6-branched-chain amino acids isoleucine and leucine were the most 13C-depleted carbon skeletons in the biosynthetic pathway.
Carbon isotope ratios (13C/12C and 14C/12C) between photoautotrophs and methanotrophs in the Black Sea. ANME-specific cross-plot of δ13C and Δ14C for the present study, compilation of previous reports using surface chlorophyll-a, dissolved inorganic carbon (DIC) and surface total organic carbon (TOC) at P167 station53, northwestern Black Sea (43°58.88′ N31°30.83′). The same ANME samples (i.e., ANME-dominated mats and the carbonate sample discussed in this study) were used in the radiocarbon analysis.
Carbon isotope ratios (13C/12C) of amino acids. Carbon isotopic composition of neutral amino acids in photoautotrophs (upper diagram: data from ref.25) and methanotrophs (lower diagram: this study). 13C-depletion proceeds through carbon elongation for C2-glycine, C3-alanine, C5-valine, C6-isoleucine, and C6-leucine. Δ13C is defined as the difference of δ13C during these target molecules.
Implications for the fate of 13C-depleted amino acids and lipid synthesis in cell biomass
The biosynthesis of isoprenoid lipids from branched-chain amino acids has been postulated26 and experimentally verified for Euryarchaeota in archaea27,28. Therefore, the branched-chain carbon structures of several amino acids have been considered to be important precursors of branched carbon skeletons and branched alkyl lipids. Among the pyruvate amino acid family, leucine is the product most closely related to lipid synthesis, leading to fundamental C5-isoprenoid precursors followed by isoprenoid unit elongation. Deuterium (2H) probing experiments suggest that partial input from leucine to the mevalonate pathway occurs in archaeal ether lipid synthesis28 and bacterial iso-C15 branched fatty acid synthesis29.
The present results and postulated biochemical reaction schemes28,29 imply that the fraction (f) of intracellular 13C-depleted branched-chain amino acids (i.e., leucine as fBCAA) is partially involved in the C5 isoprenoid pathway during membrane lipid synthesis, whereas the other fraction of the intracellular acetoacetyl-CoA pathway (facetyl-CoA) is substantially involved in this process, as expressed in facetyl-CoA + fBCAA = 1 (refs23,24,26,27,28). Alternatively, incorporated extracellular leucine could also produce mevalonate prior to lipid biosynthesis mixotrophically, as suggested by tracer experiments for archaea28. Considering physicochemical influences such as environmental factors, archaeal growth phase, and nutrient profiles27, the branched-chain amino acid transporter and the permease protein were clearly identified during the metagenomic analysis of ANME-1 sample16. Carbon isotopic profiles indicate that leucine and other pyruvate family amino acids potentially play a role in the biosynthesis of the 13C-depleted isoprenoid C5 precursor. The present ANME lipid analysis supports this interpretation by revealing extremely 13C-depleted branched-chain amino acids and isoprenoid lipids (Fig. 3; Supplementary Fig. S4). Similarly, this observation is consistent with previous results showing that 13C-depleted ether lipids (e.g., C20 and C40 isoprenoid units, including hydroxyl ether lipids) are found in ANME layers in the Black Sea19,20,21. Our results advance the biochemical understanding of benthic methane biogeochemistry driven by ANME habitats.
Methods
Sampling location of anaerobic methanotrophic archaea
Anaerobic methanotrophic archaea (ANME)−1- and ANME-2-dominated microbial mats and carbonate samples were collected from the northwestern Black Sea during the R/V Meteor cruise M72/1 (44°46.46′N, 31°59.50′E, depth 235 m) (ref.30). The samples were separated into an ANME-1-dominated pink microbial mat and ANME-2-dominated black mat and carbonate sections20,22. To minimize oxygen contamination, all sampling was performed under a nitrogen atmosphere. The pink mats were separated from the black mats with sterile scalpels. Then, the mats were transferred to glass bottles containing filter-sterilized and Black Sea water. The bottles were sealed with butyl rubber stoppers and plastic screw caps and flushed with methane30.
ANME-specific carbon (13C/12C, 14C/12C) and nitrogen (15N/14N) isotope analysis
Dried samples (ca. 10–30 µg) of pink (ANME-1 dominated), black (ANME-2 dominated), and carbonate sections were transferred to precleaned tin cups prior to isotopic analysis. Carbon and nitrogen isotopic compositions and total carbon and nitrogen contents were determined by using an isotope ratio-monitoring mass spectrometer (Thermo Finnigan Delta Plus XP, Thermo Fisher Scientific) connected to an optimized elemental analyzer (Flash EA1112, CE Instruments)31. Carbon and nitrogen isotopic compositions were expressed using conventional δ notation as follows.
The standard deviations for the carbon and nitrogen isotopic compositions were validated within δ13C < ± 0.2‰ (n = 7) and δ15N < ± 0.3‰ (n = 7) with the standard reagent BG-T (ref.32). ANME samples were also prepared for radiocarbon measurement by single-stage accelerator mass spectrometry at the Atmosphere and Ocean Research Institute, The University of Tokyo33,34. The Δ14C notation is defined as
Cross-plots of δ13C and Δ14C are shown in Supplementary Table S1 and Fig. 4.
Compound-specific isotope analysis (CSIA) of protein amino acids
CSIA of protein amino acids was performed. After acid hydrolysis of the ANME samples with 6 M HCl (110 °C, 12 h), the amino acid fraction was separated into a hydrophilic fraction for derivatization to N-pivaloyl isopropyl esters35,36,37,38 and a lipophilic fraction (hexane/dichloromethane, 6:5, v/v) for further lipid analysis. To eliminate organic and inorganic matrix effects and to improve the baseline resolution and accuracy of CSIA, the amino acid fraction was pretreated by cation-exchange column chromatography (AG-50W-X8 resin; 200–400 mesh, Bio-Rad Laboratories; cf. δ15N profiles in ref.38). Supplementary Fig. S3 shows the validation of the carbon isotopic composition of N-pivaloyl isopropyl esters of amino acids before and after column chromatography. Carbon isotopic analyses by online gas chromatography/combustion/isotope ratio mass spectrometry (GC/C/IRMS) were performed with an IRMS (Finnigan Delta Plus XP, Thermo Fisher Scientific) combined with a GC (6890 N, Agilent Technologies) with a capillary column (Ultra2, Agilent Technologies; 25 m × 0.32 mm; film thickness, 0.52 μm) in combustion and reduction furnaces36,37. The GC heating program was as follows: 3 min at 40 °C, 40–110 °C at a rate of 15 °C min−1, 110–150 °C at a rate of 3 °C min−1, 150–220 °C at a rate of 6 °C min−1, and an isothermal hold at 220 °C for 17.3 min. We occasionally used an HP-INNOWAX column (Agilent Technologies; 30 m × 0.32 mm; film thickness, 0.50 μm), a general purpose dimethylpolysiloxane column (HP-1, Agilent Technologies; 30 m × 0.32 μm; film thickness, 1.0 μm), and a joint column with DB-23 (30 m x 0.32 mm i.d., 0.25 μm, Agilent Technologies) and Ultra2 for threonine (Thr), tyrosine (Tyr), and aspartic acid (Asp), respectively, to improve baseline resolution37.
Compound-specific isotope analysis (CSIA) of archaeal lipids
Archaeal lipids were assessed in the same ANME samples (i.e., C20 and C40 isoprenoids, Supplementary Fig. S4) using an improved method39. After separation of the lipid fraction, an ether cleavage treatment40,41 was performed with 57 wt % HI (in H2O) in a reaction vial with a PTFE-lined cap at 110 °C for 4 h. After the addition of 5 wt % NaCl aqueous solution (5 mL) and n-hexane (5 mL), the n-hexane fraction was recovered by liquid/liquid extraction (×3). Then, n-hexane (5 mL) and PtO2 (5 mg) were added to the sample tube, and hydrogenation was performed by gentle H2 gas bubbling at room temperature for 30 min. Finally, the n-hexane fraction was recovered for the final fraction prior to gas chromatography with flame ionization detection (GC/FID; 6890, Agilent Technologies) and an HP-5 column (Agilent Technologies; 30 m × 0.32 mm i.d., 0.52 μm film thickness). Based on the lipid analysis of the ANME-1 sample, the relative abundance of archaeal lipids (>98%; Supplementary Fig. S4) was greater than that of bacterial alkyl lipids. The key enzymatic process of anaerobic methanotrophy by methyl-coenzyme M reductase (mcr) with coenzyme factor 430 (F430) has been validated previously, together with the CSIA of isolated F430 (ref.42). The results from lipids and F430 are consistent with evidence of a nonsyntrophic AOM process driven by ANME alone43.
References
Reeburgh, W. S. Oceanic methane biogeochemistry. Chem. Rev. 107, 486–513 (2007).
Hinrichs, K. U., Hayes, J. M., Sylva, S. P., Brewer, P. G. & deLong, E. F. Methane-consuming archaebacteria in marine sediments. Nature 398, 802–805 (1999).
Elvert, M., Suess, E. & Whiticar, M. J. Anaerobic methane oxidation associated with marine gas hydrates: superlight C-isotopes from saturated and unsaturated C20 and C25 irregular isoprenoids. Naturwissenschaften 86, 295–300 (1999).
Orphan, V. J., House, C. H., Hinrichs, K. U., McKeegan, K. D. & deLong, E. F. Methane-consuming archaea revealed by directly coupled isotopic and phylogenetic analysis. Science 293, 484–487 (2001).
Hoehler, T. M., Alperin, M. J., Albert, D. B. & Martens, C. S. Field and laboratory studies of methane oxidation in an anoxic marine sediment: evidence for a methanogen-sulfate reducer consortium. Global. Biogeochem. Cycles 8, 451–463 (1994).
Summons, R. E., Jahnke, L. L. & Roksandic, Z. Carbon isotopic fractionation in lipids from methanotrophic bacteria: relevance for interpretation of the geochemical record of biomarkers. Geochim. Cosmochim. Acta 58, 2853–2863 (1994).
Teske, A. et al. Microbial diversity of hydrothermal sediments in the Guaymas Basin: evidence for anaerobic methanotrophic communities. Appl. Environ. Microbiol. 68, 1994–2007 (2002).
Schouten, S., Wakeham, S. G., Hopmans, E. C. & Sinninghe Damsté, J. S. Biogeochemical evidence that thermophilic archaea mediate the anaerobic oxidation of methane. Appl. Environ. Microbiol. 69, 1680–1686 (2003).
Boetius, A. et al. A marine microbial consortium apparently mediating anaerobic oxidation of methane. Nature 407, 623–626 (2000).
Haroon, M. F. et al. Anaerobic oxidation of methane coupled to nitrate reduction in a novel archaeal lineage. Nature 500, 567–570 (2013).
Beal, E. J., House, C. H. & Orphan, V. J. Manganese- and iron-dependent marine methane oxidation. Science 325, 184–187 (2009).
Krüger, M. et al. A conspicuous nickel protein in microbial mats that oxidize methane anaerobically. Nature 426, 878–881 (2003).
Shima, S. et al. Structure of a methyl-coenzyme M reductase from Black Sea mats that oxidize methane anaerobically. Nature 481, 98–101 (2012).
Hallam, S. J. et al. Reverse methanogenesis: testing the hypothesis with environmental genomics. Science 305, 1457–1462 (2004).
Scheller, S., Goenrich, M., Boecher, R., Thauer, R. K. & Jaun, B. The key nickel enzyme of methanogenesis catalyses the anaerobic oxidation of methane. Nature 465, 606–608 (2010).
Meyerdierks, A. et al. Metagenome and mRNA expression analyses of anaerobic methanotrophic archaea of the ANME-1 group. Environ. Microbiol. 12, 422–439 (2010).
Treude, T. et al. Consumption of methane and CO2 by methanotrophic microbial mats from gas seeps of the anoxic Black Sea. Appl. Environ. Microbiol. 73, 2271–2283 (2007).
Reeburgh, W. S. et al. Black Sea methane geochemistry. Deep Sea Res. Part 1 Oceanogr. 38, S1189–S1210 (1991).
Thiel, V. et al. Molecular signals for anaerobic methane oxidation in Black Sea seep carbonates and a microbial mat. Marine Chem. 73, 97–112 (2001).
Blumenberg, M., Seifert, R., Reitner, J., Pape, T. & Michaelis, W. Membrane lipid patterns typify distinct anaerobic methanotrophic consortia. PNAS 101, 11111–11116 (2004).
Michaelis, W. et al. Microbial reefs in the Black Sea fueled by anaerobic oxidation of methane. Science 297, 1013–1015 (2002).
Schmale, O., Beaubien, S. E., Rehder, G., Greinert, J. & Lombardi, S. Gas seepage in the Dnepr paleo-delta area (NW-Black Sea) and its regional impact on the water column methane cycle. J. Marine Syst. 80, 90–100 (2010).
Kellermann, M. Y. et al. Autotrophy as a predominant mode of carbon fixation in anaerobic methane-oxidizing microbial communities. PNAS 109, 19321–19326 (2012).
Hayes, J. M. Fractionation of carbon and hydrogen isotopes in biosynthetic processes. Rev. Mineral. Geochem. 43, 225–277 (2001).
Keil, R. G. & Fogel, M. L. Reworking of amino acid in marine sediments: stable carbon isotopic composition of amino acids in sediments along the Washington coast. Limnol. Oceanogr. 46, 14–23 (2001).
Eglinton, G. & Hamilton, R. J. Leaf epicuticular waxes. Science 156, 1322–1335 (1967).
Whitman, W. B., Ankwanda, E. & Wolfe, R. S. Nutrition and carbon metabolism of Methanococcus voltae. J. Bacteriol. 149, 852–863 (1982).
Yamauchi, N. The pathway of leucine to mevalonate in halophilic archaea: efficient incorporation of leucine into isoprenoidal lipid with the involvement of isovaleryl-CoA dehydrogenase in Halobacterium salinarum. Biosci. Biotechnol. Biochem. 74, 443–446 (2010).
Beck, H. C., Hansen, A. M. & Lauritsen, F. R. Catabolism of leucine to branched-chain fatty acids in Staphylococcus xylosus. J. Appl. Microbiol. 96, 1185–1193 (2004).
Siegert, M. et al. The nitrogen cycle in anaerobic methanotrophic mats of the Black Sea is linked to sulfate reduction and biomass decomposition. FEMS Microbiol. Ecol. 86, 231–245 (2013).
Ogawa, N. O., Nagata, T., Kitazato, H. & Ohkouchi, N. Ultrasensitive elemental analyzer/isotope ratio mass spectrometer for stable nitrogen and carbon isotope analyses. Earth, Life and Isotopes. Kyoto Univ. Press, 339–353 (2010).
Tayasu, I. et al. New organic reference materials for carbon- and nitrogen- stable isotope ratio measurements provided by Center for Ecological Research, Kyoto University, and Institute ofBiogeosciences, Japan Agency for Marine-Earth Science and Technology. Limnol. 12, 261–266 (2011).
Yamane, M. et al. Compound-specific 14C dating of IODP Expedition 318 core U1357A obtained off the Wilkes Land coast, Antarctica. Radiocarbon 56, 1009–1017 (2014).
Yokoyama, Y. et al. Developing ultra small-scale radiocarbon sample measurement at the University of Tokyo. Radiocarbon 52, 310–318 (2010).
Metges, C. C., Petzke, K. J. & Hennig, U. Gas chromatography combustion isotope ratio mass spectrometric comparison of N-acetyl- and N-pivaloyl amino acid esters to measure N-15 isotopic abundances in physiological samples. J. Mass Spectrometry 31, 367–376 (1996).
Chikaraishi, Y. et al. Determination of aquatic food-web structure based on compound-specific nitrogen isotopic composition of amino acids. Limnol. Oceanogr: Methods 7, 740–750 (2009).
Chikaraishi, Y., Takano, Y., Ogawa, N.O. & Ohkouchi, N. Instrumental optimization for compound-specific nitrogen isotope analysis of amino acids by gas chromatography/combustion/isotope ratio mass spectrometry. Earth, Life, and Isotopes, Kyoto Univ. Press, pp. 367–386 (2010).
Takano, Y. et al. Isolation and desalting with cation-exchange chromatography for compound-specific nitrogen isotope analysis of amino acids. Rapid Commun. Mass Spectrom. 24, 2317–2323 (2010).
Takano, Y. et al. Sedimentary membrane lipids recycled by deep-sea benthic archaea. Nature Geosci. 3, 858–861 (2010).
Schouten, S. et al. Structural characterization, occurrence and fate of archaeal ether-bound acyclic and cyclic biphytanes and corresponding diols in sediments. Org. Geochem. 29, 1305–1319 (1998).
Elvert, M., Suess, E., Greinert, J. & Whiticar, M. J. Archaea mediating anaerobic methane oxidation in deep-sea sediments at cold seeps of the eastern Aleutian subduction zone. Org. Geochem. 31, 1175–1187 (2000).
Kaneko, M. et al. Quantitative analysis of coenzyme F430 in environmental samples: a new diagnostic tool for methanogenesis and anaerobic methane oxidation. Anal. Chem. 86, 3633–3638 (2014).
Milucka, J. et al. Zero-valent sulfur is a key intermediate in marine methane oxidation. Nature 491, 541–546 (2012).
Luth, C., Luth, U., Gebruk, A. V. & Thiel, H. Methane gas seeps along the oxic/anoxic gradient in the Black Sea: manifestations, biogenic sediment compounds and preliminary results on benthic ecology. Marine Ecol. 20, 221–249 (1999).
Naudts, L. et al. Geological and morphological setting of 2778 methane seeps in the Dnepr paleo-delta, northwestern Black Sea. Marine Geol. 227, 177–199 (2006).
Whiticar, M. J., Faber, E. & Schoell, M. Biogenic methane formation in marine and freshwater environments: CO2 reduction vs. acetate fermentation: Isotope evidence. Geochim. Cosmochim. Acta 50, 693–709 (1986).
Peckmann, J. et al. Methane-derived carbonates and authigenic pyrite from the northwestern Black Sea. Marine Geol. 177, 129–150 (2001).
Treude, T. et al. Subsurface microbial methanotrophic mats in the Black Sea. Appl. Environ. Microbiol. 71, 6375–6378 (2005).
Lüdmann, T. et al. Heat flow and quantity of methane deduced from a gas hydrate field in the vicinity of the Dnieper Canyon, northwestern Black Sea. Geo-Marine Lett. 24, 182–193 (2004).
Pape, T. et al. Marine methane biogeochemistry of the Black Sea: A review. In Links between Geological Processes, Microbial Activities & Evolution ofLife, Springer, 281–311 (2008).
Naudts, L. et al. Anomalous sea-floor backscatter patterns in methane venting areas, Dnepr paleo-delta, NW Black Sea. Marine Geol. 251, 253–267 (2008).
Bertram, S. et al. Methanogenic capabilities of ANME archaea deduced from 13C-labelling approaches. Environ. Microbiol. 15, 2384–2393 (2013).
Kusch, S. et al. Implications for chloro-and pheopigment synthesis and preservation from combined compound-specific δ13C, δ15N, and Δ14C analysis. Biogeosci. 7, 4105–4118 (2010).
Acknowledgements
We thank Dr. R. Seifert and Prof. Dr. W. Michaelis (Univ. Hamburg) and the shipboard party of the R/V Meteor cruise (M72/1: Geochemical cycling and microbial life at active gas seeps) for providing the samples from the Black Sea. We thank Dr. M. Elvert (Univ. Bremen) and Dr. G. Wegener (MPI Bremen) for providing constructive and critical comments. We also thank Y. Saito, Y. Sasaki (JAMSTEC) for technical support and Dr. Y. Takaki (JAMSTEC) for the discussion of genomic sequence. The study was supported in part by the Japan Society for the Promotion of Science (JSPS: No. 16H04083, 15H05332) and the Submarine Resources Research Project in JAMSTEC.
Author information
Authors and Affiliations
Contributions
Y.T. performed the chemical analysis and wrote the paper; Y.C. and N.O.O. performed the IRMS analysis with the carbon isotope standard reagents; M.K. performed the analysis of coenzyme factor 430 derived from the glutamic acid potential precursor; H.I. performed the phylogenic molecular analysis; Y.M. and Y.Y. performed the 14C-AMS analysis; M.K. performed the deep-sea field survey and the ANME sampling; and Y.T., Y.C., and N.O. designed the study. All the authors discussed the results and commented on the manuscript.
Corresponding author
Ethics declarations
Competing Interests
The authors declare no competing interests.
Additional information
Publisher's note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Takano, Y., Chikaraishi, Y., Imachi, H. et al. Insight into anaerobic methanotrophy from 13C/12C- amino acids and 14C/12C-ANME cells in seafloor microbial ecology. Sci Rep 8, 14070 (2018). https://doi.org/10.1038/s41598-018-31004-5
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-018-31004-5
Keywords
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.