Abstract
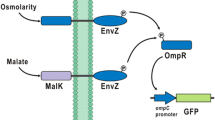
Escherichia coli does not have the methanol sensing apparatus, was engineered to sense methanol by employing chimeric two-component system (TCS) strategy. A chimeric FlhS/EnvZ (FlhSZ) chimeric histidine kinase (HK) was constructed by fusing the sensing domain of Paracoccus denitrificans FlhS with the catalytic domain of E. coli EnvZ. The constructed chimeric TCS FlhSZ/OmpR could sense methanol by the expression of ompC and gfp gene regulated by ompC promoter. Real-time quantitative PCR analysis and GFP-based fluorescence analysis showed the dynamic response of the chimeric TCS to methanol. The expression of ompC and the gfp fluorescence was maximum at 0.01 and 0.5% of methanol, respectively. These results suggested that E. coli was successfully engineered to sense methanol by the introduction of chimeric HK FlhSZ. This strategy can be employed for the construction of several chimeric TCS based bacterial biosensors for the development of biochemical producing recombinant microorganisms.
Similar content being viewed by others
References
California Environmental Protection Agency (CalEPA) (1999) Air Toxics Hot Spots Program Risk Assessment Guidelines: Part III. Technical Support Document for the Determination of Noncancer Chronic Reference Exposure Levels. SRP Draft. Office of Environmental Health Hazard Assessment, Berkeley, CA.
Ed. Budavari, S. and N. Rahway (1989) The Merck Index. An Encyclopedia of Chemicals, Drugs, and Biologicals.
Stock, A. M., V. L. Robinson, and P. N. Goudreau (2000) Two-Component Signal Transduction. Annu. Rev. Biochem. 69: 183–215.
West, A. H. and A. M. Stock (2001) Histidine kinases and response regulator proteins in two-component signaling systems. Trends in Biochem. Sci. 26: 369–376.
Utsumi, R., R. Brissette, A. Rampersaud, S. Forst, K. Oosawa, and M. Inouye (1989) Activation of bacterial porin gene expression by a chimeric signal transducer in response to aspartate. Sci. 245: 1246–1249.
Harms, N., W. N. M. Reijnders, S. Koning, and R. J. M. van Spanning (2001) Two-component system that regulates methanol and formaldehyde oxidation in Paracoccus denitrificans. J. Bacteriol. 183: 664–670.
Harms, N., J. Ras, W. N. Reijnders, R. J. van Spanning, and A. H. Stouthamer (1996) S-formylglutathione hydrolase of Paracoccus denitrificans is homologous to human esterase D: A universal pathway for formaldehyde detoxification? J. Bacteriol. 178: 6296–6299.
Ras, J., P. W. Van Ophem, W. N. Reijnders, R. J. Van Spanning, J. A. Duine, A. H. Stouthamer, and N. Harms (1995) Isolation, sequencing, and mutagenesis of the gene encoding NAD-and glutathione-dependent formaldehyde dehydrogenase (GDFALDH) from Paracoccus denitrificans, in which GD-FALDH is essential for methylotrophic growth. J. Bacteriol. 177: 247–251.
Baslé, A., G. Rummel, P. Storici, J. P. Rosenbusch, and T. Schirmer (2006) Crystal structure of Osmoporin OmpC from E. coli at 2.0 Å. J. Mol. Biol. 362: 933–942.
Eswar, N., B. Webb, M. A. Marti-Renom, M. S. Madhusudhan, D. Eramian, M. -Y. Shen, U. Pieper, and A. Sali (2007) Comparative protein structure modeling using MODELLER. Curr. Protoc. Protein Sci., John Wiley & Sons, Inc., City.
Bhattacharya, D. and J. Cheng (2013) 3Drefine: Consistent protein structure refinement by optimizing hydrogen bonding network and atomic-level energy minimization. Proteins 81: 119–131.
Eleaume, H. and S. Jabbouri (2004) Comparison of two standardisation methods in real-time quantitative RT-PCR to follow Staphylococcus aureus genes expression during in vitro growth. J. Microbiol. Meth. 59: 363–370.
Maruthamuthu, M., I. Ganesh, S. Ravikumar, and S. Hong (2015) Evaluation of zraP gene expression characteristics and construction of a lead (Pb) sensing and removal system in a recombinant Escherichia coli. Biotechnol. Lett. 37: 659–664.
Mayson, B. E., D. J. Kilburn, B. L. Zamost, C. K. Raymond, and G. J. Lesnicki (2003) Effects of methanol concentration on expression levels of recombinant protein in fed-batch cultures of Pichia methanolica. Biotechnol. Bioeng. 81: 291–298.
Ganesh, I., S. Ravikumar, I.-K. Yoo, and S. Hong (2015) Construction of malate-sensing Escherichia coli by introduction of a novel chimeric two-component system. Bioproc. Biosyst. Eng. 38: 797–804.
Wen, G., X. Wen, S. Shuang, and M. M. F. Choi (2014) Wholecell biosensor for determination of methanol. Sensors and Actuators B: Chem. 201: 586–591.
Zhao, C., H. Li, J. Sheng, L. Chen, F. Li, S. Yang, C. Dong, and M. M. F. Choi (2009) Isolation of a Methylobacterium organophilium strain, and its application to a methanol biosensor. Microchim. Acta 167: 67–73.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Selvamani, V., Ganesh, I., Maruthamuthu, M.k. et al. Engineering chimeric two-component system into Escherichia coli from Paracoccus denitrificans to sense methanol. Biotechnol Bioproc E 22, 225–230 (2017). https://doi.org/10.1007/s12257-016-0484-y
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12257-016-0484-y