Abstract
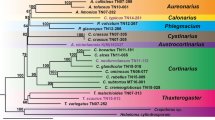
The fragrant orchid Gymnadenia conopsea s.l. is a controversial taxon with two commonly distinguished species, G. conopsea s.str. and G. densiflora. Despite morphological similarity, differentiation between the taxa has been reported for several characters; however, character variation within taxa has obviated a clear consensus. We assessed ITS sequences, microsatellite variation and chromosome numbers on the European scale (1,420 samples) and conducted morphological analyses for 626 samples from Germany. ITS analysis revealed a 2% nucleotide divergence between the taxa, similar to the divergence between other Gymnadenia species. The ITS sequences of G. densiflora form a well-supported monophyletic group sharing a most recent common ancestor with G. nigra and G. austriaca. Thus, G. conopsea and G. densiflora are not sister species, and a species rank is supported for G. densiflora (Wahlenb.) Dietrich and G. conopsea (L.) R.Br. s.str. This was confirmed by the microsatellite analysis, which revealed a strong genetic differentiation between the taxa because of largely non-overlapping sets of alleles. Chromosome numbers showed that G. conopsea was either diploid or tetraploid, whereas G. densiflora was diploid throughout. Morphologically, the taxa differed significantly in the mean value of a number of diagnostic characters. However, a discriminant analysis showed that the morphological variability is substantial, and on the individual level an unequivocal assignment is not possible as 96% of G. conopsea, but only 77% of G. densiflora could be assigned correctly. Further studies are needed on character variation within and among species and ploidy levels to allow for a better identification of the genetically differentiated but morphologically similar taxa.




Similar content being viewed by others
References
Amich F, García-Barriuso M, Bernardos S (2007) Polyploidy and speciation in the orchid flora of the Iberian Peninsula. Bot Helv 117:143–157
Bateman RM, Pridgeon AM, Chase MW (1997) Phylogenetics of subtribe Orchidinae (Orchidoideae, Orchidaceae) based on nuclear ITS sequences. 2. Intrageneric relationships and reclassification to achieve monophyly of Orchis sensu stricto. Lindleyana 12:113–141
Bateman RM, Hollingsworth PM, Preston J, Yi-Bo L, Pridgeon AM, Chase MW (2003) Molecular phylogenetics and evolution of Orchidinae and selected Habenariinae (Orchidaceae). Bot J Linn Soc 142:1–40
Bateman RM, Rudall PJ, James KE (2006) Phylogenetic context, generic affinities and evolutionary origin of the enigmatic Balkan orchid Gymnadenia frivaldii Hampe ex Griseb. Taxon 55:107–118
Bernardos S, Amich F, Gallego F (2003) Karyological and taxonomic notes on Ophrys (Orchidoideae, Orchidaceae) from the Iberian Peninsula. Bot J Linn Soc 142:395–406
Bianco P, D’Emerico S, Medagli P, Ruggiero L (1991) Polyploidy and aneuploidy in Ophrys, Orchis, and Anacamptis (Orchidaceae). Plant Syst Evol 178:235–245
Bisse J (1963) Ein Beitrag zur Kenntnis der deutschen Orchideenflora. Feddes Repert 67:181–189
Bory S, Catrice O, Brown S, Leitch IJ, Gigant R, Chiroleu F, Grisoni M, Duval MF, Besse P (2008) Natural polyploidy in Vanilla planifolia (Orchidaceae). Genome 51:816–826
Campbell VV, Rowe G, Beebee TJC, Hutchings MJ (2002) Isolation and characterization of microsatellite primers for the fragrant orchid Gymnadenia conopsea (L.) R. Brown (Orchidaceae). Conserv Genet 3:209–210
Campbell VV, Rowe G, Beebee TJC, Hutchings MJ (2007) Genetic differentiation amongst fragrant orchids (Gymnadenia conopsea s.l.) in the British Isles. Bot J Linn Soc 155:349–360
Chung MY (2009) Low levels of genetic variation within populations of the four rare orchids Gymnadenia cucullata, Gymnadenia camtschatica, Amitostigma gracile, and Pogonia minor in South Korea: indication of genetic drift and implications for conservation. Plant Syst Evol 281:65–76
Clement M, Posada D, Crandall KA (2000) TCS: a computer program to estimate gene genealogies. Mol Ecol 9:1657–1659
Cozzolino S, Widmer A (2005a) Orchid diversity: an evolutionary consequence of deception? Trends Ecol Evol 20:487–494
Cozzolino S, Widmer A (2005b) The evolutionary basis of reproductive isolation in Mediterranean orchids. Taxon 54:977–985
Cresswell JE (1998) Stabilizing selection and the structural variability of flowers within species. Ann Bot 81:463–473
Delforge P (2006) Orchids of Europe, North Africa and the Middle East, 3rd edn. A&C Black Publishers Ltd., London
Development Core Team R (2009) R: a language and environment for statistical computing. R Foundation for Statistical Computing, Vienna, Austria
Devos N, Oh S-H, Raspé O, Jacquemart AL, Manos PS (2005) Nuclear ribosomal DNA sequence variation and evolution of spotted marsh-orchids (Dactylorhiza maculata group). Mol Phylogenet Evol 36:568–580
Dressler RL (1987) Die Orchideen: Biologie und Systematik der Orchidaceae. Eugen Ulmer GmbH & Co., Stuttgart
Dworschak W (2002) Gliederung der verschiedenen Erscheinungsformen der Mücken-Händelwurz in Südbayern. In: Naturwissenschaftlicher Verein Wuppertal e.V. (Ed.) Jahresberichte des Naturwissenschaftlichen Vereins Wuppertal e.V., Heft 55, Wuppertal, pp 27–45
Gill DE (1989) Fruiting failure, pollinator inefficiency and speciation in orchids. In: Otte D, Endler JA (eds) Speciation and its consequences. Sinauer Associates Inc., pp 458–481
Greilhuber J, Ehrendorfer F (1975) Chromosome numbers and evolution in Ophrys (Orchidaceae). Plant Syst Evol 124:125–138
Guindon S, Gascuel O (2003) A simple, fast, and accurate algorithm to estimate large phylogenies by maximum likelihood. Syst Biol 52:696–704
Gustafsson S (2000) Patterns of genetic variation in Gymnadenia conopsea, the fragrant orchid. Mol Ecol 9:1863–1872
Gustafsson S, Lönn M (2003) Genetic differentiation and habitat preference of flowering-time variants within Gymnadenia conopsea. Heredity 91:284–292
Gustafsson S, Sjögren-Gulve P (2002) Genetic diversity in the rare orchid, Gymnadenia odoratissima and a comparison with the more common congener, G. conopsea. Conserv Genet 3:225–234
Gustafsson S, Thorén PA (2001) Microsatellite loci in Gymnadenia conopsea, the fragrant orchid. Mol Ecol Notes 1:81–82
Hedrén M, Klein E, Teppner H (2000) Evolution of polyploids in the European orchid genus Nigritella: evidence from allozyme data. Phyton-Ann Rei Bot A 40:239–275
Hedrén M, Fay MF, Chase MW (2001) Amplified fragment length polymorphisms (AFLP) reveal details of polyploid evolution in Dactylorhiza (Orchidaceae). Am J Bot 88:1868–1880
Hill T, Lewicki P (2007) STATISTICS methods and applications. StatSoft, Tulsa
Jersáková J, Castro S, Sonk N, Milchreit K, Schödelbauerová I, Tolasch T, Dötterl S (2010) Absence of pollinator-mediated premating barriers in mixed-ploidy populations of Gymnadenia conopsea s.l. (Orchidaceae). Evol Ecol 24:1199–1218
Juillet N, Delle-Vedove R, Dormont L, Schatz B, Pailler T (2010) Differentiation in a tropical deceptive orchid: colour polymorphism and beyond. Plant Syst Evol 289:213–221
Lee SS (2002) A review of orchid mycorrhizae in Korea. The Plant Pathol J 18:169–178
Librado P, Rozas J (2009) DnaSP v5: a software for comprehensive analysis of DNA polymorphism data. Bioinformatics 25:1451–1452
Mallet J (2005) Hybridization as an invasion of the genome. Trends Ecol Evol 20:229–237
Marhold K, Jongepierová I, Krahulcová A, Kučera J (2005) Morphological and karyological differentiation of Gymnadenia densiflora and G. conopsea in the Czech Republic and Slovakia. Preslia 77:159–176
Micheneau C, Johnson SD, Fay MF (2009) Orchid pollination: from Darwin to the present day. Bot J Linn Soc 161:1–19
Mrkvicka AC (1993) Statistische Untersuchungen an Gymnadenia conopsea (L.) R. Br. s.l. Mitt Bl Arbeitskr Heim Orch Baden-Württ 25:361–367
Nei M, Kumar S (2000) Molecular evolution and phylogenetics. Oxford University Press, New York
Nordström S, Hedrén M (2008) Genetic differentiation and postglacial migration of the Dactylorhiza majalis ssp. traunsteineri/lapponica complex into Fennoscandia. Plant Syst Evol 276:73–87
Otero JT, Flanagan NS (2006) Orchid diversity—beyond deception. Trends Ecol Evol 21:64–65
Peakall R (2007) Speciation in the Orchidaceae: confronting the challenges. Mol Ecol 16:2834–2837
Posada D (2008) jModelTest: phylogenetic model averaging. Mol Biol Evol 25:1253–1256
Ramsey J, Schemske DW (1998) Pathways, mechanisms, and rates of polyploid formation in flowering plants. Annu Rev Ecol Syst 29:467–501
Salzmann CC, Nardella AM, Cozzolino S, Schiestl FP (2007) Variability in floral scent in rewarding and deceptive orchids: The signature of pollinator-imposed selection? Ann Bot 100:757–765
Scacchi R, de Angelis G (1989) Isoenzyme polymorphisms in Gymnaedenia conopsea and its inferences for systematics within this species. Biochem Syst Ecol 17:25–33
Schiestl FP, Schlüter PM (2009) Floral isolation, specialized pollination, and pollinator behavior in orchids. Annu Rev Entomol 54:425–446
Schmeil O (1996) Flora von Deutschland, 90. Auflage. Quelle & Meyer Bestimmungsbücher, Wiesbaden
Schwarzacher T, Ambros P, Schweizer D (1980) Application of giemsa banding to orchid karyotype analysis. Plant Syst Evol 134:293–297
Scopece G, Widmer A, Cozzolino S (2008) Evolution of postzygotic reproductive isolation in a guild of deceptive orchids. Am Nat 171:315–326
Soliva M, Widmer A (1999) Genetic and floral divergence among sympatric populations of Gymnadenia conopsea s.l. (Orchideaceae) with different flowering phenology. Int J Plant Sci 160:897–905
Ståhlberg D (2009) Habitat differentiation, hybridization and gene flow patterns in mixed populations of diploid and autotetraploid Dactylorhiza maculata s.l. (Orchidaceae). Evol Ecol 23:295–328
Stark C, Babik W, Durka W (2009) Fungi from the roots of the common terrestrial orchid Gymnadenia conopsea. Mycol Res 113:952–959
Stephens M, Smith NJ, Donnelly P (2001) A new statistical method for haplotype reconstruction from population data. Am J Hum Genet 68:978–989
Swarts ND, Dixon KW (2009) Terrestrial orchid conservation in the age of extinction. Ann Bot 104:543–556
Tamura K, Dudley J, Nei M, Kumar S (2007) MEGA4: molecular evolutionary genetics analysis (MEGA) software version 4.0. Mol Biol Evol 24:1596–1599
Taylor DL, Bruns TD, Szaro TM, Hodges SA (2003) Divergence in mycorrhizal specialization within Hexalectris spicata (Orchidaceae), a nonphotosynthetic desert orchid. Am J Bot 90:1168–1179
Taylor DL, Bruns TD, Hodges SA (2004) Evidence for mycorrhizal races in a cheating orchid. Proc R Soc Lond B Biol Sci 271:35–43
Thompson JN (1994) The coevolutionary process. The University of Chicago Press, Chicago
Trávníček P, Kubátová B, Čurn V, Rauchová J, Krajníková E, Jersáková J, Suda J (2011) Remarkable coexistence of multiple cytotypes of the Gymnadenia conopsea aggregate (the fragrant orchid): evidence from flow cytometry. Ann Bot 107:77–87
Tutin TG, Heywood VH, Burges NA, Valentine DH, Walters SM, Webb DA (1980) Flora Europaea Cambridge University Press, Cambridge
van den Berg C, Higgins WE, Dressler RL, Whitten WM, Soto-Arenas MA, Chase MW (2009) A phylogenetic study of Laeliinae (Orchidaceae) based on combined nuclear and plastid DNA sequences. Ann Bot 104:417–430
Vereecken NJ, Dafni A, Cozzolino S (2010) Pollination syndromes in Mediterranean orchids-implications for speciation, taxonomy and conservation. Bot Rev 76:220–240
Vöth W, Sontag S (2006) Die intraspezifischen Varietäten der Gymnadenia conopsea (L.). R Br J Eur Orch 38:581–624
White TJ, Bruns T, Lee S, Taylor J (1990) Amplification and direct sequencing of fungal ribosomal RNA genes for phylogenetics. In: Innis MA, Gelfand DH, Sninsky JJ, White TJ (eds) PCR protocols: a guide to methods and applications. Academic Press, Inc., San Diego, pp 315–322
Winterfeld G, Röser M (2007) Chromosomal localization and evolution of satellite DNAs and heterochromatin in grasses (Poaceae), especially tribe Aveneae. Plant Syst Evol 264:75–100
Acknowledgments
We are grateful to local AHOs (Arbeitskreis Heimische Orchideen) and their members for their support during this work. Special thanks to K. Heyde, W. Mohrman, P. Rode, J. Passin, P. Steinfeld and F. Opitz for locating plant populations; and to W. Dworschak, O. Gerbaud, A. Hoch, H. Kerschbaumsteiner, S. Klemich, P. Kuss and M. L. Perko for providing samples. Thanks to Jan Hanspach for statistical advice and two anonymous reviewers for valuable comments. This work was supported by the Federal Ministry of Education and Research (BMBF-FKz 01LC0018A) within the framework of BIOLOG-SUBICON (Successional Change and Biodiversity Conservation).
Author information
Authors and Affiliations
Corresponding author
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Stark, C., Michalski, S.G., Babik, W. et al. Strong genetic differentiation between Gymnadenia conopsea and G. densiflora despite morphological similarity. Plant Syst Evol 293, 213–226 (2011). https://doi.org/10.1007/s00606-011-0439-x
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00606-011-0439-x