Abstract
Previous studies indicated that miRNA plays an important role in human biological processes especially in the field of diseases. However, constrained by biotechnology, only a small part of the miRNA-disease associations has been verified by biological experiment. This impel that more and more researchers pay attention to develop efficient and high-precision computational methods for predicting the potential miRNA-disease associations. Based on the assumption that molecules are related to each other in human physiological processes, we developed a novel structural deep network embedding model (SDNE-MDA) for predicting miRNA-disease association using molecular associations network. Specifically, the SDNE-MDA model first integrating miRNA attribute information by Chao Game Representation (CGR) algorithm and disease attribute information by disease semantic similarity. Secondly, we extract feature by structural deep network embedding from the heterogeneous molecular associations network. Then, a comprehensive feature descriptor is constructed by combining attribute information and behavior information. Finally, Convolutional Neural Network (CNN) is adopted to train and classify these feature descriptors. In the five-fold cross validation experiment, SDNE-MDA achieved AUC of 0.9447 with the prediction accuracy of 87.38% on the HMDD v3.0 dataset. To further verify the performance of SDNE-MDA, we contrasted it with different feature extraction models and classifier models. Moreover, the case studies with three important human diseases, including Breast Neoplasms, Kidney Neoplasms, Lymphoma were implemented by the proposed model. As a result, 47, 46 and 46 out of top-50 predicted disease-related miRNAs have been confirmed by independent databases. These results anticipate that SDNE-MDA would be a reliable computational tool for predicting potential miRNA-disease associations.
Similar content being viewed by others
Introduction
MicroRNAs (miRNAs) are one type of small non-coding RNA with length of 20–25 nucleotides1. They normally influence their target messenger RNAs (mRNAs) by base pairing binding to the 3′ untranslated region (UTR) sites of mRNAs2. These small molecules could function as negative regulator of target gene expression in post-transcriptional3. With the development of molecular biology, increasing miRNAs have been detected4. To date, the famous miRbase database have collected 48,860 mature miRNAs from 271 organisms containing more than 1000 human miRNAs5. In addition, researchers have found that miRNAs are related with multiple significant cell biological activities, involving diffusion, aging, development, death and so on6,7,8,9.
In recent years, an increasing number of experiments have demonstrated that there are close relationships between miRNA with disease10,11,12,13. In particular, miRNAs have been new biomarkers for human cancer, which is important to cancer preventions and treatments14. Therefore, identifying the miRNA-disease associations has gradually become a hot topic in biology15. Early traditional biological experiments identified the disease-related miRNAs by detecting the expression level of miRNAs in biological disease process16. For example, Yohei et al. found that miR-200c could build a molecular link between breast cancer cells and normal cells17. Liu et al. point out that many miRNAs are disordered in cancer and this situation occurs because miRNAs participate in tumorigenesis and function as oncogenes18. Thum et al. reported that miR-21 adjust expression of the ERK-MAP kinase to effect on structure and function of heart19. Traditional experiments achieve high accuracy, while it has the limitations of long experimental time, high cost, and low success rate20. To resolve these issues, for effectively and accurately predict potential miRNA-disease associations, increasing researchers adopted computational model and select the most possible related miRNAs for further traditional biological experiments21.
With the development of biotechnology, some databases were constructed by collecting these biological data. These datasets provide the possibility to classify associations of miRNA-disease through computational methods20,22,23,24,25. Over the years, these methods mostly are according to the assumption that these functionally similar miRNAs tend to be related with semantically similar diseases2,26,27,28. These models could be split into under similarity network models and machine learning models29. For example, Jiang et al.22 presented a computational model to speculate the relationship between miRNA and disease based on a hypergeometric distribution model. This is an early calculation model by fusing multiple sources of information. However, this method built the miRNA-related network by functional similarity, which is limited by the relationship between miRNAs. Based on random walk method, Xuan et al.30 presented MIDP and MIDPE, an extension method of MIDP. MIDP constructed the network by combining the information of each node including similarity, prior information and various ranges of topological structure. This model could effectively reduce noise from data by restarting the walk. Furthermore, You et al.31 proposed PBMDA constructed a heterogeneous graph including three sub-graphs. PBMDA is a depth-first algorithm based on path, which could fully use the topology information of heterogeneous network. In particularly, the priority of new associations between diseases and miRNAs could be identified by evaluating the score of the path. Chen et al.32 proposed a computational method adopted the extreme gradient boosting named EGBMMDA. This is the first learning method based on decision tree for classifying miRNA-disease relationships. EGBMMDA built a comprehensive feature vector by various methods such as statistical, graph theory and matrix factorization. These studies have continually improved the performance of computational method and played an important guiding role in traditional biological experiments33. Therefore, accurately and effectively predict associations between miRNA-disease through computational method become urgently demanded34.
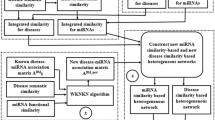
In this study, based on the assumption of molecules are related to each other in human physiological processes, we developed a structural deep network embedding-based model (SDNE-MDA) for predicting miRNA-disease association using molecular association network. The flow chart of SDNE-MDA is shown as Fig. 1. Specifically, we first constructed the molecular association network (MAN)35 by combining multiple different molecules with edges of them. This study extracted behavior information from the heterogeneous network by the structural deep network embedding (SDNE)36, which could maintain the overall structure of large network to the greatest extent. Secondly, SDNE-MDA obtained the miRNA attribute information by the chaos game representation (CGR) algorithm and disease attribute information by disease semantic similarity. After then, we formed the feature descriptor by fusing the behavior information and attribute information of miRNAs and diseases. Finally, these feature descriptors are trained and classified by the CNN to predict miRNA-disease associations. Five-fold cross validation experiment was carried out for SDNE-MDA to verify the performance of prediction and achieved the AUC of 0.9447 with the prediction accuracy of 87.38%. To further evaluate SDNE-MDA, we contrasted the proposed model with two feature extraction models and classifier models. Besides, we carry out SDNE-MDA with three significant human diseases involving breast cancer, kidney cancer and lymphoma. And as a result, 47, 46 and 46 out of top-50 candidate related miRNAs are confirmed by known databases and recent literature, respectively. These experiment result demonstrated that SDNE-MDA is a precisely and effectively computational method for predicting potential associations between miRNA with disease.
Materials and methods
Benchmark database
Human miRNA-disease associations benchmark database HMDD v3.037 was adopted as data support in this paper, which collected 32,281 confirmed miRNA-disease associations, involving 1102 miRNAs and 850 diseases. Here, after data processing, we chose 16,427 known miRNA-disease associations as positive samples including 1023 miRNAs and 850 diseases. What’s more, we defined the adjacency matrix \(AM\) to represent the miRNA-disease associations. When the miRNA \(mi(a)\) have a verified association with the disease \(di(b)\), we set \(AM(mi(a),di(b))=1\), otherwise \(AM(mi(a),di(b))=0\). In this paper, we introduce two other independent databases (dbDEMC38 and miR2Ddisease39) to verified the result of case study.
Molecular associations network
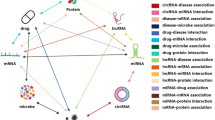
In this study, we combined multiple biological molecular information according the Molecular association network (MAN). The MAN is a heterogeneous information network proposed by Guo et al.40. Currently, this complex network consists of five types of molecular (miRNA, lncRNA, protein, disease, drug) and associations between them. The heterogeneous information network MAN provided a new comprehensive view to explore the complex physiological process and human disease. The structure diagram of molecular association network is as shown in Fig. 2. In this study, we download the information of molecular and associations between them from multiple databases. The number of different molecules is shown in Table 1, and the associations between them are shown in the following Table 2.
Chaos game representation (CGR) algorithm
MiRNA sequences contain a lot of complex information. However, most of the existing sequence feature information extraction algorithms only quantify one of position information and nonlinear information. In order to measure the similarity of these information contained in the miRNA sequences comprehensively. In this study, we chose chaos game representation (CGR)50 to quantize position and nonlinear information to calculate miRNA sequence similarity by pearson coefficient. Firstly, the positions of four nucleotides of miRNA are mapped to Euclidean space by the following formula:
where \({T}_{i}\) is the position of \(i\)th nucleotide, and it is related to the position of the previous nucleotide \({T}_{i-1}\) and the nucleotide coefficient \({G}_{i}\). In this paper, the contribution parameter \(c\) is equal to 0.5 and \({T}_{0}\) is \((0.5, 0.5)\).
Secondly, we divided the CGR space into 64 subspaces as shown in Fig. 3. The attribute information of each subspace \({SS}_{i}\) would be represented by integrating the position information \({X}_{i}, {Y}_{i}\) and nonlinear information \({Z}_{i}\) by the following formula:
where \({num}_{i}\) is the number of points in subspace \({SS}_{i}\).
Finally, each miRNA sequence information could be represented by the descriptor \(m(i)\). And we calculate sequence similarity \({M}_{sim}(m\left(i\right),m(j))\) by Pearson correlation coefficient.
Disease semantic similarity
In this study, the Directed Acyclic Graph (DAG)51 of diseases could be obtained from the Medical Subject Headings (Mesh)52. In the system, a disease \(d(a)\) could be defined by \(DAG(d(a)) = (L(d(a)), E(d(a)))\), where \(L(d(a))\) is a node set including \(d(a)\) and ancestor nodes of \(d(a)\), and \(E(d(a))\) indicates directed edge set of all relationships from ancestor node to child node. The semantic value of \(d(a)\) was contributed by term \(T\) as the formula:
where \(\vartheta\) is a parameter of semantic contribution, and \(\vartheta\) is equal to 0.5 as previous study. Therefore, \(DV\left(D\right)\) of \(D\) could be calculated as follows:
According the assumption that two diseases should have higher similarity if they hold more same parts in DAG, the similarity of the diseases \(d(a)\) with \(d(b)\) could be obtained as follows:
Structural deep network embedding
Since existing network embedding algorithms could not keep the high-order proximity of large-scale networks, this paper adopted the structural deep network embedding (SDNE) to extract the behavior information of miRNAs and diseases. Many existing network embedding models are shallow model (e.g. Laplacian Eigenmaps53, Graph Factorization54), which are unable to validly extract the highly non-linear structural information of network. SDNE is a semi-supervised model for network embedding. For the part of supervised, first-order similarity based on Laplacian matrix would be adopted to preserve local network information. And the part of unsupervised, SDNE used deep autoencoder modeling second-order similarity to save the global network information. Therefore, the loss function of SDNE is divided into two parts, i.e. Laplacian matrix model and Deep autoencoder model.
First-order similarity
To make adjacent nodes of graph closer in the latent space, the loss function of first-order similarity could be obtained as following formula:
where \({s}_{i,j}\) is the adjacency matrix for heterogeneous information network and \({y}_{i}^{(k)}\) indicates the node \(i\) of \(k\)-th layer.
Second-order similarity
For the capturing of global structure information, SDNE construct the deep autoencoder model. Any given \({x}_{i}\) could be convert into the latent representation of \(k\)th layer as:
here \({W}^{\left(k\right)}\) is the \(k\)th layer weight matrix and \({b}^{\left(k\right)}\) as a parameter. According the optimization goal of the autoencoder is to reduce the reconstruction error in input and output, therefore, we could define the loss function as follows:
The adjacency matrices are often very sparse, which means zero elements are far more than non-zero elements. Therefore, the loss function would be optimized as:
where \(\odot\) is the Hadamard product (multiplying the corresponding elements).
Integrating the first-order similarity and second-order similarity, the finally loss function of SDNE is shown as follows:
where \({L}_{reg}\) is a regularization term, and \(\alpha\) is a parameter to control the loss of the first-order similarity. The regularization term is shown as:
Integration of feature information
In this study, we firstly obtained miRNA sequence similarity and disease semantic similarity and convert them into attribute feature information \({M}_{sim}(i)\), \({D}_{sim}(j)\) of same dimension by stacked autoencoder. The dimension of \({M}_{sim}(i)\) and \({D}_{sim}(j)\) is 64. After then, the behavior feature information of miRNAs \({M}_{b}(i)\) and diseases \({D}_{b}(j)\) were extracted by the structural deep network embedding based on the molecular association network. The dimension of \({M}_{b}(i)\) and \({D}_{b}(j)\) is 128. Finally, a complete sample feature descriptor is constructed by fusing above information based on the HMDD v3.0 database. The feature descriptor was a 384-dimensional vector as follows:
Convolutional neural network algorithm
Convolutional neural network (CNN) is a deep-structured feedforward neural network with convolution calculations. CNN could shift-invariant classify the input information based on layer structure by representation learning capability. With the development of research, CNN has been successfully utilized in bioinformatics55. Therefore, in this paper, we adopted the CNN to train and predict potential miRNA-disease association. Specifically, CNN has a multi-layer structure including input, convolutional layer, pooling layer, fully-connected layer and output as shown in Fig. 4. The input layer is a matrix of all feature descriptor \(FD\left(i,j\right)\) with size \(26284\times 384\). Two convolutional layers \(C1\) and \(C2\) are obtained by 32 filters with \(3\times 1\) convolution kernel and 64 filters with \(3\times 1\) convolution kernel. In this study, we adopted max-pooling \(2\times 1\) kernel to subsample the \(C2\). After repeatedly convolution and pooling, CNN classifies the features from fully-connected layer and output the probability distribution.
Results and discussion
Performance evaluation
In this experiment, we implemented the five-fold cross validation to evaluate the performance of proposed model under HMDD v3.037. These known miRNA-disease pairs would be randomly split into five subsets with no intersection. Each cross validation, one of five subsets would be set as test set and remaining data sets as train set. To avoid the revelation of test data, we constructed the heterogeneous information network by only training data and extract the behavior information. In this study, a class of evaluation criteria were used to assess SDNE-MDA, including accuracy (Acc.), sensitivity (Sen.), specificity (Spec.), precision (Prec.), Matthews Correlation Coefficient (MCC) and area under curve (AUC). As a result, the average Acc, Sen, Spec, Prec, MCC and AUC achieved 87.38%, 87.28%, 87.47%, 87.45%, 74.76% and 0.9447 with standard deviations of 0.44%, 0.93%, 1.01%, 0.82%, 0.88% and 0.0027, respectively as shown in Table 3. In addition, the receiver operating characteristics (ROC) curve and area under precision-recall (PR) curve by SDNE-MDA based on HMDD are shown in Fig. 5.
Comparison with different feature extraction methods
In this study, these nodes in the network could be represented by the attribute and behavior information. Both types of information may influence the result of prediction, so we compared the different feature extraction methods including SDNE-MDA_AI composed of attribute information, SDNE-MDA_BI composed of behavior information and SDNE-MDA composed of both them. In addition, attribute information of other nodes has scarcely effect on prediction of potential miRNA-disease relationships. For reducing the redundancy of model, we only considered the attribute information of miRNAs and diseases. The detail result of comparison between proposed model with different feature extraction models are shown in Table 4. The accuracy of SDNE-MDA is 7.78% and 3.43% higher than that of SDNE-MDA_AI and SDNE-MDA_BI, respectively. In addition, the AUC of proposed model is 0.0811 and 0.0260 higher than SDNE-MDA_AI and SDNE-MDA_BI. The ROC curves and PR curves of three experiments are shown in Fig. 6. These results indicated that integrating the two kind of information to represent the node achieved more distinguished performance.
Comparison with different classifier models
In this study, the CNN was adopted to train and identify potential relationships between miRNA and disease. To further evaluate SDNE-MDA, we compare proposed model with Bagging, Logistic Regression, Naive Bayes and Adaboost classifier model. In this experiment, we implemented the five-fold cross validation in these different classifier models based on the HMDD v3.0. Finally, the proposed model yielded average AUC of 0.9447 based on five-fold cross validation and outperformed Bagging (0.8998), LogisticRegression (0.9270), Naive Bayes (0.8881), Adaboost (0.9226) and MLP (0.9320). The AUC of CNN is 0.0259 higher than the mean AUC of all five model, and the accuracy is 1.60% higher than that of the second highest methods. The detail results of the comparison between SDNE-MDA and other four classifier models are shown in Table 5, and we drew the ROC curves as shown in Fig. 7. Therefore, CNN algorithm is the optimal selection for the proposed model to predicting potential miRNA-disease associations.
Comparison with related work
An increasing number of researchers have focused on the prediction of miRNA-disease associations, and a mass of model have been proposed. To further evaluate the predictive performance of our method, the SDNE-MDA was compared with six state-of-the-art classical methods under five-fold cross validation, including RWRMDA56, MTDN57, EGBMMDA32, LMTRDA58, DBMDA59 and PBMDA31. Since these algorithms have not calculated multiple evaluation criteria, we only compare the AUC on the terms of five-fold cross validation based HMDD database. The detail results of the comparison between SDNE-MDA and other six related works are shown in Table 6. The proposed method is 0.0399 higher than the average AUC of all algorithms, and 0.0275 higher than that of the second highest methods. This is mainly due to SDNE-MDA integrated two types of information of miRNAs and diseases, and extract the feature more comprehensively. Therefore, the proposed model is an effective and reliable computational tool for predicting potential miRNA-disease associations.
Case studies
For further evaluating the prediction ability of SDNE-MDA, we implemented case studies based on three significant human diseases (Breast Neoplasms, Kidney Neoplasms, Lymphoma). In this study, these known miRNA-disease associations based on HMDD v3.0 database would be the training set. To avoid the overlap in the train data and prediction list, the test set is the unknown relationship pairs between three diseases and all possible miRNAs. As a result, 47, 46 and 46 of top-50 candidate related miRNAs were confirmed by independent databases. Therefore, SDNE-MDA is a feasible and reliable model for predicting potential relationships between miRNA and disease.
Breast Neoplasms is the most universal neoplasms in female and the risk of breast cancer is up to 13% in the United States. Although men may also develop breast cancer, 99% of patients are women. There are approximately 276,480 novel cases in women and 42,170 were die from breast cancer in 202060. In previous few years, studies had indicated the expression level of miRNA have strong impact to growth and division of breast tumor cell61. Therefore, we implemented a case study of Breast Neoplasms-miRNA associations by SDNE-MDA. In the prediction list shown as Table 7, 47 of top 50 predicted Breast Neoplasms related miRNAs were verified based on independent databases.
Kidney Neoplasms is a novel cancer with higher adult incidence60. In the past few years, however, morbidity and mortality of kidney neoplasms have been increasing. There are about 73,750 novel cases in kidney neoplasms with about 45,520 in male and about 28,230 in female in United States and about 14,830 deaths for this cancer (9860 men and 4970 women) in 2020. Recently, increasing researchers have indicated miRNAs are related with kidney neoplasms62. Thus, we take Kidney Neoplasms as a case study for SDNE-MDA and prioritize the candidate miRNAs. In the prediction list shown as Table 8, 46 of top-50 potential kidney neoplasms-related miRNAs were confirmed by independent databases.
Lymphoma is one of the most common malignant cancers (~ 4% of all new cancer) especially in teenagers in United States60. Lymphoma mainly contains two types of Hodgkin Lymphoma (HL) and non-Hodgkin Lymphoma (NHL). In 2020, it is estimated that about 85,720 new cases of Lymphoma (47,070 of men and 38,650 of women) and 20,910 deaths for HL and NHL (12,030 of men and 8,880 of women). Therefore, we implemented SDNE-MDA to prioritize possible miRNAs for Lymphoma based on HMDD v3.0. As shown in Table 9, 46 out of top 50 predicted Lymphoma candidate miRNAs were verified by independent databases.
Conclusion
In previous few years, accumulating number of researches demonstrated that miRNAs have closely link with diseases. Various of biological experiments and computational methods are committed to classify the association of them. In this paper, we proposed a structural deep network embedding-based model SDNE-MDA to predict miRNA-disease associations. This model constructed a complex network MAN by fusing miRNAs, diseases and three related molecular (lncRNA, drug and protein) with their relationships. Through the comprehensive heterogeneous information network, potential miRNA-disease associations could be predicted more accurate and efficient. And CNN is utilized to train and classify the potential miRNA-disease associations. Compared with other classifiers and feature extraction models, SDNE-MDA showed outstanding performance. In addition, case studies were implemented on three significant human disease for further validate performance of SDNE-MDA. As a result, 47, 46 and 46 of top-50 predicted miRNAs have been confirmed by independent databases. These results demonstrated that SDNE-MDA is a reliable computational tool for predicting miRNA-disease associations.
References
Kloosterman, W. P. & Plasterk, R. H. A. The diverse functions of microRNAs in animal development and disease. Dev. Cell 11, 441–450 (2006).
Ji, B.-Y. et al. Predicting miRNA-disease association from heterogeneous information network with GraRep embedding model. Sci. Rep. 10, 6658 (2020).
Ines, A. G. & Miska, E. A. MicroRNA functions in animal development and human disease. Development 132, 4653–4662 (2005).
Guo, Z.-H. et al. A learning based framework for diverse biomolecule relationship prediction in molecular association network. Commun. Biol. 3, 1–9 (2020).
Kozomara, A., Birgaoanu, M. & Griffiths-Jones, S. miRBase: From microRNA sequences to function. Nucleic Acids Res. 47, D155–D162 (2018).
Cheng, A. M., Byrom, M. W., Jeffrey, S. & Ford, L. P. Antisense inhibition of human miRNAs and indications for an involvement of miRNA in cell growth and apoptosis. Nucleic Acids Res. 33, 1290–1297 (2005).
Xantha, K. & Victor, A. Developmental biology. Encountering microRNAs in cell fate signaling. Science 310, 1288–1289 (2005).
Miska, E. A. How microRNAs control cell division, differentiation and death. Curr. Opin. Genet. Dev. 15, 563–568 (2005).
Xu, P., Guo, M. & Hay, B. A. MicroRNAs and the regulation of cell death. Trends Genet. 20, 617–624 (2004).
Ramiro, G., Guido, M. & Croce, C. M. Targeting microRNAs in cancer: Rationale, strategies and challenges. Nat. Rev. Drug Discov. 9, 775–789 (2010).
Farazi, T. A., Spitzer, J. I., Pavel, M. & Thomas, T. miRNAs in human cancer. J. Pathol. 223, 102–115 (2015).
You, Z.-H. et al. PRMDA: Personalized recommendation-based miRNA-disease association prediction. Oncotarget 8, 85568 (2017).
Wang, L. et al. Using two-dimensional principal component analysis and rotation forest for prediction of protein–protein interactions. Sci. Rep. 8, 12874 (2018).
Bartels, C. L. & Tsongalis, G. J. MicroRNAs: Novel biomarkers for human cancer. Clin. Chem. 55, 623–631 (2009).
Zheng, K. et al. MLMDA: A machine learning approach to predict and validate microRNA-disease associations by integrating of heterogenous information sources. J. Transl. Med. 17, 1–14 (2019).
Chen, X., Xie, D., Zhao, Q. & You, Z.-H. MicroRNAs and complex diseases: From experimental results to computational models. Brief. Bioinform. 20, 515–539 (2019).
Yohei, S. et al. Downregulation of miRNA-200c links breast cancer stem cells with normal stem cells. Cell 138, 592–603 (2009).
Liu, B. et al. MiR-26a enhances metastasis potential of lung cancer cells via AKT pathway by targeting PTEN. BBA Mol. Basis Disease 1822, 1692–1704 (2012).
Thum, T. et al. MicroRNA-21 contributes to myocardial disease by stimulating MAP kinase signalling in fibroblasts. Nature 456, 980–984 (2008).
Chen, X. et al. WBSMDA: Within and between score for miRNA-disease association prediction. Sci. Rep. 6, 21106 (2016).
Weidhaas, J. Using microRNAs to understand cancer biology. Lancet Oncol. 11, 136–146 (2010).
Jiang, Q. et al. Prioritization of disease microRNAs through a human phenome-microRNAome network. BMC Syst. Biol. 4, S2 (2010).
Xuan, P. et al. Correction: Prediction of microRNAs associated with human diseases based on weighted k most similar neighbors. PLoS ONE 8, e70204 (2013).
Chen, X. et al. HGIMDA: Heterogeneous graph inference for miRNA-disease association prediction. Oncotarget 7, 65257 (2016).
Wang, L., Wang, H.-F., Liu, S.-R., Yan, X. & Song, K.-J. Predicting protein–protein interactions from matrix-based protein sequence using convolution neural network and feature-selective rotation forest. Sci. Rep. 9, 9848 (2019).
Huang, Z.-A. et al. PBHMDA: Path-based human microbe-disease association prediction. Front. Microbiol. 8, 233 (2017).
Chen, X. Predicting lncRNA-disease associations and constructing lncRNA functional similarity network based on the information of miRNA. Sci. Rep. 5, 13186 (2015).
Pasquier, C. & Gardès, J. Prediction of miRNA-disease associations with a vector space model. Sci. Rep. 6, 27036 (2016).
Li, J.-Q., Rong, Z.-H., Chen, X., Yan, G.-Y. & You, Z.-H. MCMDA: Matrix completion for MiRNA-disease association prediction. Oncotarget 8, 21187 (2017).
Ping, X. et al. Prediction of potential disease-associated microRNAs based on random walk. Bioinformatics 31, 1805–1815 (2015).
You, Z. H. et al. PBMDA: A novel and effective path-based computational model for miRNA-disease association prediction. Plos Computat. Biol. 13, e1005455 (2017).
Chen, X., Huang, L., Xie, D. & Zhao, Q. EGBMMDA: Extreme gradient boosting machine for MiRNA-disease association prediction. Cell Death Dis. 9, 3 (2018).
Huang, Y.-A. et al. EPMDA: An expression-profile based computational model for microRNA-disease association prediction. Oncotarget 8, 87033 (2017).
Chen, X., Cheng, J.-Y. & Yin, J. Predicting microRNA-disease associations using bipartite local models and hubness-aware regression. RNA Biol. 15, 1192–1205 (2018).
Guo, Z.-H., Yi, H.-C. & You, Z.-H. Construction and comprehensive analysis of a molecular association network via lncRNA–miRNA–disease–drug–protein graph. Cells, 8(8), 866 (2019).
Wang, D., Peng, C. & Zhu, W. Structural deep network embedding. In Proceedings of the 22nd ACM SIGKDD international conference on Knowledge discovery and data mining, 1225–1234 (2016).
Huang, Z. et al. HMDD v3.0: A database for experimentally supported human microRNA-disease associations. Nucleic Acids Res. 47, D1013–D1017 (2018).
Yang, Z. et al. dbDEMC 2.0: Updated database of differentially expressed miRNAs in human cancers. Nucleic Acids Res. 45, D812–D818 (2017).
Jiang, Q. et al. miR2Disease: A manually curated database for microRNA deregulation in human disease. Nucleic Acids Res. 37, D98-104 (2009).
Guo, Z.-H., et al. Integrative construction and analysis of molecular association network in human cells by fusing node attribute and behavior information. Mol. Therapy-Nucleic Acids 19, 498–506 (2020).
Zhou, H. et al. HMDD v3. 0: a database for experimentally supported human microRNA–disease associations. Nucleic Acids Res. 47(D1), D1013–D1017 (2018).
Chou, C.-H., et al. miRTarBase update 2018: a resource for experimentally validated microRNA-target interactions. Nucleic Acids Res. 46(D1), D296–D302 (2017).
Wishart, D. S. et al. DrugBank 5.0: A major update to the DrugBank database for 2018. Nuclc Acids Res. 46, D1074 (2018).
Chen, G. et al. LncRNADisease: A database for long-non-coding RNA-associated diseases. Nuclc Acids Res. 41, D983–D986 (2013).
Miao, Y., Liu, W., Zhang, Q. & Guo, A. lncRNASNP2: An updated database of functional SNPs and mutations in human and mouse lncRNAs. Nucleic Acids Res. 46, D276–D280 (2018).
Szklarczyk, D. et al. The STRING database in 2017: quality-controlled protein–protein association networks, made broadly accessible. Nucleic Acids Res. 45, gkw937 (2017).
Cheng, L. et al. LncRNA2Target v2.0: A comprehensive database for target genes of lncRNAs in human and mouse. Nucleic Acids Res. 47, D140–D144 (2019).
Davis, A. P. et al. The Comparative Toxicogenomics Database: Update 2019. Nucleic Acids Res. 47, D948–D954 (2019).
Janet, P. et al. DisGeNET: A comprehensive platform integrating information on human disease-associated genes and variants. Nucleic Acids Res. D833–D839 (2017).
Jeffrey, H. J. Chaos game representation of gene structure. Nucleic Acids Res. 18, 2163–2170 (1990).
Kalisch, M. & Buehlmann, P. Estimating high-dimensional directed acyclic graphs with the PC-algorithm. J. Mach. Learn. Res. 8, 613–636 (2012).
Lipscomb, C. E. Medical subject headings (MeSH). Bull. Med. Libr. Assoc. 88, 265 (2000).
Belkin, M. & Niyogi, P. Laplacian eigenmaps for dimensionality reduction and data representation. Neural Comput. 15(6), 1373–1396 (2003).
Ahmed, A., Shervashidze, N., Narayanamurthy, S., Josifovski, V. & Smola, A. J. Distributed large-scale natural graph factorization. In Proceedings of the 22nd international conference on World Wide Web, 37–48 (2013).
Wang, L., You, Z.-H., Huang, Y.-A., Huang, D.-S. & Chan, K. C. An efficient approach based on multi-sources information to predict circRNA-disease associations using deep convolutional neural network. Bioinformatics 36, 4038–4046 (2020).
Chen, X., Liu, M. X. & Yan, G. Y. RWRMDA: Predicting novel human microRNA-disease associations. Mol. BioSyst. 8, 2792–2798 (2012).
Xu, J. et al. Prioritizing candidate disease miRNAs by topological features in the miRNA target-dysregulated network: Case study of prostate cancer. Mol. Cancer Ther. 10, 1857–1866 (2011).
Wang, L. et al. LMTRDA: Using logistic model tree to predict MiRNA-disease associations by fusing multi-source information of sequences and similarities. PLoS Computat. Biol. 15, e1006865 (2019).
Zheng, K. et al. Dbmda: A unified embedding for sequence-based miRNA similarity measure with applications to predict and validate miRNA-disease associations. Mol. Therapy-Nucleic Acids 19, 602–611 (2020).
Siegel, R. L., Miller, K. D. & Jemal, A. Cancer statistics, 2019. CA Cancer J. Clin. 69, 7–34 (2019).
Iorio, M. V. et al. MicroRNA gene expression deregulation in human breast cancer. Can. Res. 65, 7065–7070 (2005).
Muhamed Ali, A. et al. A machine learning approach for the classification of kidney cancer subtypes using miRNA genome data. Mol. Therapy-Nucleic Acids 8, 2422 (2018).
Acknowledgements
The authors would like to thank all anonymous reviewers for their constructive advice.
Funding
This work is supported in part by the National Natural Science Foundation of China, under Grant 61702444, in part by the West Light Foundation of The Chinese Academy of Sciences, under Grant 2018-XBQNXZ-B-008, in part by the Chinese Postdoctoral Science Foundation, under Grant 2019M653804, in part by the Tianshan Youth—Excellent Youth, under Grant 2019Q029, in part by the Qingtan scholar talent project of Zaozhuang University.
Author information
Authors and Affiliations
Contributions
H.L., H.C., Z.Y. and L.W. conceived the algorithm, carried out analyses, prepared the data sets, carried out experiments, and wrote the manuscript. S.S., X.Y. and J.Y. analyzed experiments. All authors reviewed the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Li, HY., Chen, HY., Wang, L. et al. A structural deep network embedding model for predicting associations between miRNA and disease based on molecular association network. Sci Rep 11, 12640 (2021). https://doi.org/10.1038/s41598-021-91991-w
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-021-91991-w
This article is cited by
-
RETRACTED ARTICLE: Graph Neural Network on Psychological Prediction of College Students Special Education
Journal of Autism and Developmental Disorders (2024)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.