Abstract
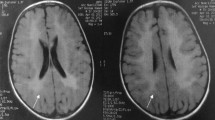
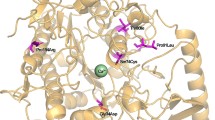
Metachromatic leukodystrophy (MLD) is a neurodegenerative disorder characterized by progressive demyelination resulting from impaired degradation and thus the accumulation of cerebroside-3-sulfate (sulfatide). It is caused by the deficiency of arylsulfatase A (ARSA) enzyme which is encoded by the ARSA gene. The present study reports the clinical, molecular, and bioinformatic investigation of three patients belonging to a consanguineous family with late-infantile MLD disorder. The results revealed a novel homozygous missense mutation c.699C>A (p.His231Gln) in exon 4 of ARSA gene in the three patients inherited from their heterozygous parents. Interestingly, this novel mutation is the second mutation identified in the substrate-binding site of ARSA protein and it was classified as damaging and deleterious by several bioinformatics tools. The c.699C>A (p.His231Gln) leads to changes in the pre-mRNA secondary structure and in the ARSA protein 3D structure with a significant root mean square deviation value which could probably affect its stability and function.

Similar content being viewed by others
References
Berná L, Gieselmann V, Poupetová H, Hrebícek M, Elleder M, Ledvinová J (2004) Novel mutations associated with metachromatic leukodystrophy: phenotype and expression studies in nine Czech and Slovak patients. Am J Med Genet 129:277–281
Cesani M, Capotondo A, Plati T, Sergi LS, Fumagalli F, Roncarolo MG, Naldini L, Comi G, Sessa M, Biffi A (2009) Characterization of new arylsulfatase A gene mutations reinforces genotype-phenotype correlation in metachromatic leukodystrophy. Hum Mutat 30:936–945
Cesani M, Lorioli L, Grossi S, Amico G, Fumagalli F, Spiga I, Filocamo M, and Biffi A (2015) Mutation Update of ARSA and PSAP Genes Causing Metachromatic Leukodystrophy. Hum mutat 37: 16–27
Choi Y, Chan AP (2015) PROVEAN web server: a tool to predict the functional effect of amino acid substitutions and indels. Bioinformatics 16:15–31
Deconinck N, Messaaoui A, Ziereisen F, Kadhim H, Sznajer Y, Pelc K, Nassogne MC, Vanier MT, Dan B (2008) Metachromatic leukodystrophy without arylsulfatase a deficiency: a new case of saposin-B deficiency. Eur J Paediatr Neurol 12:46–50
Gieselmann V (2008) Metachromatic leukodystrophy: genetics, pathogenesis and therapeutic options. Acta Paediatr 97:15–21
Gieselmann V, Zlotogora J, Harris A, Wenger DA, Morris CP (1994) Molecular genetics of metachromatic leukodystrophy. Hum Mutat 4:233–242
Grossi S, Regis S, Rosano C, Corsolini F, Uziel G, Sessa M, Di Rocco M, Parenti G, Deodato F, Leuzzi V et al (2008) Molecular analysis of ARSA and PSAP genes in twenty-one Italian patients with metachromatic leukodystrophy: identification and functional characterization of 11 novel ARSA alleles. Hum Mutat 29:E220–E230
Halsal DJ, Eugene Halligan P, Terence Elsey S, and Timothy M. Cox (1999) Metachromatic leucodystrophy: a newly identified mutation in arylsulphatase A, D281Y, found as a compound heterozygote with I179L in an adult onset case. HUMAN MUTATION Mutation in Brief #273
Kreysing A, Von Figura K, Gieselmann V (1990) Structure of the arylsulfatase a gene. Eur J Biochem 19:1627–1631
Lewin HA, Stewart-Haynes JA (1992) A simple method for DNA extraction from leukocytes for use in PCR. BioTechniques 13:522–524
Li B, Krishnan VG, Mort ME, Xin F, Kamati KK, Cooper DNSD, Mooney RP (2009) Automated inference of molecular mechanisms of disease from amino acid substitutions. Bioinformatics 25:2744–2750
Lugowska A, Berger J, Tylki-Szyman’ska A (2005) Molecular and phenotypic characteristics of metachromatic leukodystrophy patients from Poland. Clin Genet 68:48–54
Lugowska A, Mierzewska H, Bekiesinska-Figatowska M, Szczepanik E, Goszczanska-Ciuchta A, Bednarska-Makaruk M (2014) A homozygote for the c.459+1G>a mutation in the ARSA gene presents with cerebellar ataxia as the only first clinical sign of metachromatic leukodystrophy. J Neurol Sci 338:214–217
Lukatela G, Krauss N, Theis K, Selmer T, Gieselmann V, von Figura K, Saenger W (1998) Crystal structure of human arylsulfatase a: the aldehyde function and the metal ion at the active site suggest a novel mechanismfor sulfate ester hydrolysis. Biochemistry 37:3654–3664
Pauline CN, Henikoff S (2001) Predicting deleterious amino acid substitutions. Genome Res 11:863–874
Sandhoff K, Kolter H, Harzer K (2001) Sphingolipid activator proteins. In: Scriver CR, Beaudet AL, Sly WS, Valle D (eds) The metabolic and molecular bases of inherited disease, 8th edn. McGraw-Hill, New York, pp 3371–3388
Schestag F, Yaghootfam A, Habetha M, Poeppel P, Dietz F, Klein RA, Zlotogora J, Gieselmann V (2002) The functional consequences of mis-sense mutations affecting an intra-molecular salt bridge in arylsulphatase A. Biochem J 367:499–504
Schwarz JM, Rödelsperger C, Schuelke M, Seelow D (2010) Mutation Taster evaluates disease-causing potential of sequence alterations. Nat Methods 7:575–576
Stein C, Gieselmann V, Kreysing J, Schmidt B, Pohlmann R, Waheed A, Meyer HE, O’Brien JS, Von Figura K (1989) Cloning and expression of human arylsulfatase A. J Biol Chem 264:1252–1259
Sunyaev S, Ramensky V, Koch I, Lathe W 3rd, Kondrashov AS, Bork P (2001) Prediction of deleterious human alleles. Hum Mol Genet 10:591–597
Von Figura K, Gieselmann V, Jaeken J (2001) Metachromatic leukodystrophy. In: Scriver CR, Beaudet AL, Sly WS, Valle D (eds) The metabolic and molecular bases of inherited disease. McGrawHill, New York, pp 3695–3724
Wang J, Zhang W, Pan H, Bao X, Wu Y, Wu X, Jiang Y (2007) ARSA gene mutations in five Chinese metachromatic leukodystrophy patients. Pediatr Neurol 36:397–401
Zuker M, Jacobson AB (1998) Using reliability information to annotate RNA secondary structures. RNA 4:669–679
Acknowledgments
We thank the patients and their families for their cooperation in the present study. This work was supported by The Ministry of the Higher Education and the Scientific Research in Tunisia. We also extend our thanks to the doctors of NeuroPediatrics Service in the CHU Hedi Chaker of Sfax, for their help and availability. We owe special thanks to Mr. Wassim Hariz, founder and former head of the English Unit and Editorial Board at the Sfax Faculty of Science, for having proofread this paper.
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Conflict of Interest
The authors declare that they have no conflict of interest.
Additional information
Highlights
- Novel homozygous mutation p.His231Gln among the substrate binding site of catalytic domain of ARSA protein was identified in three patients with metachromatic leukodystrophy.
- The bioinformatics investigations supported the pathogenicity of the novel p.His231Gln mutation and classify it as damaging and deleterious.
- The three patients belonging to a consanguineous family and shearing the same novel deleterious homozygous mutation p.His231Gln presented late-infantile form, but with clinical heterogeneity.
- To our knowledge, p.His231Gln is the second mutation identified in the substrate-binding site and it is described for the first time in patients with metachromatic leukodystrophy.
Rights and permissions
About this article
Cite this article
Issa, A.B., Feki, F.K., Jdila, M.B. et al. Clinical, Molecular, and Computational Analysis Showed a Novel Homozygous Mutation Among the Substrate-Binding Site of ARSA Protein in Consanguineous Family with Late-Infantile MLD. J Mol Neurosci 66, 17–25 (2018). https://doi.org/10.1007/s12031-018-1141-z
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12031-018-1141-z