Abstract
Main conclusion
S. plumbizincicola genetic transformation was optimized using a self-excision molecular-assisted transformation system by integrating the SpGRF4/SpGIF1 gene with XVE and Cre/loxP.
Abstract
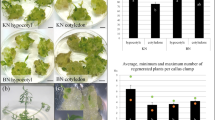
Sedum plumbizincicola, despite being an excellent hyperaccumulator of cadmium and zinc with significant potential for soil pollution phytoremediation on farmland, has nonetheless trailed behind other major model plants in genetic transformation technology. In this study, different explants and SpGRF4–SpGIF1 genes were used to optimize the genetic transformation of S. plumbizincicola. We found that petiole and stem segments had higher genetic transformation efficiency than cluster buds. Overexpression of SpGRF4–SpGIF1 could significantly improve the genetic transformation efficiency and shorten the period of obtaining regenerated buds. However, molecular assistance with overexpression of SpGRF4–SpGIF1 leads to abnormal morphology, resulting in plant tissue enlargement and abnormal growth. Therefore, we combined SpGRF4–SpGIF1 with XVE and Cre/loxP to obtain DNA autocleavage transgenic plants induced by estradiol, thereby ensuring normal growth in transgenic plants. This study optimized the S. plumbizincicola genetic transformation system, improved the efficiency of genetic transformation, and established a self-excision molecular-assisted transformation system. This work also established the basis for studying S. plumbizincicola gene function, and for S. plumbizincicola breeding and germplasm innovation.





Similar content being viewed by others
Data availability
The Gene and protein sequences of SpGRF4 and SpGIF1 have been uploaded to GenBank, and the submission ID were 2,777,890, 2,777,937. The datasets generated during and/or analyzed during the current study are available from the corresponding author on reasonable request.
Abbreviations
- GRF:
-
Growth-regulating factor
- GIF:
-
GRF-interacting factor
- XVE:
-
LexA-VP16-ER
References
Altpeter F, Springer NM, Bartley LE, Blechl AE, Brutnell TP, Citovsky V, Conrad LJ, Gelvin SB, Jackson DP, Kausch AP, Lemaux PG, Medford JI, Orozco-Cárdenas ML, Tricoli DM, Van Eck J, Voytas DF, Walbot V, Wang K, Zhang ZJ, Stewart CN Jr (2016) Advancing crop transformation in the era of genome editing. Plant Cell 28(7):1510–1520. https://doi.org/10.1105/tpc.16.00196
Bull T, Debernardi J, Reeves M, Hill T, Bertier L, Van Deynze A, Michelmore R (2023) GRF-GIF chimeric proteins enhance in vitro regeneration and Agrobacterium-mediated transformation efficiencies of lettuce (Lactuca spp.). Plant Cell Rep 42(3):629–643. https://doi.org/10.1007/s00299-023-02980-4
Chen Z, Cheng Q, Hu C, Guo X, Chen Z, Lin Y, Hu T, Bellizzi M, Lu G, Wang GL, Wang Z, Chen S, Wang F (2017) A chemical-induced, seed-soaking activation procedure for regulated gene expression in rice. Front Plant Sci 8:1447. https://doi.org/10.3389/fpls.2017.01447
Debernardi JM, Tricoli DM, Ercoli MF, Hayta S, Ronald P, Palatnik JF, Dubcovsky J (2020) A GRF-GIF chimeric protein improves the regeneration efficiency of transgenic plants. Nat Biotechnol 38(11):1274–1279. https://doi.org/10.1038/s41587-020-0703-0
Feng Q, Xiao L, He Y, Liu M, Wang J, Tian S, Zhang X, Yuan L (2021) Highly efficient, genotype-independent transformation and gene editing in watermelon (Citrullus lanatus) using a chimeric ClGRF4-GIF1 gene. J Integr Plant Biol 63(12):2038–2042. https://doi.org/10.1111/jipb.13199
Gelvin SB (2003) Agrobacterium-mediated plant transformation: the biology behind the “gene-jockeying” tool. Microbiol Mol Biol Rev 67(1):16–37. https://doi.org/10.1128/MMBR.67.1.16-37.2003
Kim JH (2019) Biological roles and an evolutionary sketch of the GRF-GIF transcriptional complex in plants. BMB Rep 52(4):227–238. https://doi.org/10.5483/BMBRep.2019.52.4.051
Kim JH, Kende H (2004) A transcriptional coactivator, AtGIF1, is involved in regulating leaf growth and morphology in Arabidopsis. Proc Natl Acad Sci USA 101(36):13374–13379. https://doi.org/10.1073/pnas.0405450101
Liu H, Zhao H, Wu L, Liu A, Zhao F-J, Xu W (2017a) Heavy metal ATPase 3 (HMA3) confers cadmium hypertolerance on the cadmium/zinc hyperaccumulator Sedum plumbizincicola. New Phytol 215(2):687–698. https://doi.org/10.1111/nph.14622
Liu H, Zhao H, Wu L, Xu W (2017b) A genetic transformation method for cadmium hyperaccumulator Sedum plumbizincicola and non-hyperaccumulating ecotype of Sedum alfredii. Front Plant Sci 16(8):1047. https://doi.org/10.3389/fpls.2017.01047
Luo G, Palmgren M (2021) GRF-GIF chimeras boost plant regeneration. Trends Plant Sci 26(3):201–204. https://doi.org/10.1016/j.tplants.2020.12.001
Siligato R, Wang X, Yadav SR, Lehesranta S, Ma G, Ursache R, Sevilem I, Zhang J, Gorte M, Prasad K, Wrzaczek M, Heidstra R, Murphy A, Scheres B, Mähönen AP (2016) MultiSite gateway-compatible cell type-specific gene-inducible system for plants. Plant Physiol 170(2):627–641. https://doi.org/10.1104/pp.15.01246
Sun P, Zhang W, Wang Y, He Q, Shu F, Liu H, Wang J, Wang J, Yuan L, Deng H (2016) OsGRF4 controls grain shape, panicle length and seed shattering in rice. J Integr Plant Biol 58(10):836–847. https://doi.org/10.1111/jipb.12473
Wu LH, Liu YJ, Zhou SB, Guo FG, Bi D, Guo XH, Baker AJM, Smith JAC, Luo YM (2013) Sedum plumbizincicola X.H. Guo et S.B. Zhou ex L.H. Wu (Crassulaceae): a new species from Zhejiang Province, China. Plant Syst Evol 299(3):487–498. https://doi.org/10.1007/s00606-012-0738-x
Ye X, Vaghchhipawala Z, Williams EJ, Fu C, Liu J, Lu F, Hall EL, Guo SX, Frank L, Gilbertson LA (2023) Cre-mediated autoexcision of selectable marker genes in soybean, cotton, canola and maize transgenic plants. Plant Cell Rep 42(1):45–55. https://doi.org/10.1007/s00299-022-02935-1
Zhang Y, Li H, Ouyang B, Lu Y, Ye Z (2006) Chemical-induced autoexcision of selectable markers in elite tomato plants transformed with a gene conferring resistance to lepidopteran insects. Biotechnol Lett 28(16):1247–1253. https://doi.org/10.1007/s10529-006-9081-z
Zhang D, Sun W, Singh R, Zheng Y, Cao Z, Li M, Lunde C, Hake S, Zhang Z (2018) GRF-interacting factor1 regulates shoot architecture and meristem determinacy in maize. Plant Cell 30(2):360–374. https://doi.org/10.1105/tpc.17.00791
Zhang Y, Zhang Q, Chen QJ (2020) Agrobacterium-mediated delivery of CRISPR/Cas reagents for genome editing in plants enters an era of ternary vector systems. Sci China Life Sci 63(10):1491–1498. https://doi.org/10.1007/s11427-020-1685-9
Zhang X, Xu G, Cheng C, Lei L, Sun J, Xu Y, Deng C, Dai Z, Yang Z, Chen X, Liu C, Tang Q, Su J (2021) Establishment of an Agrobacterium-mediated genetic transformation and CRISPR/Cas9-mediated targeted mutagenesis in Hemp (Cannabis Sativa L.). Plant Biotechnol J 19(10):1979–1987. https://doi.org/10.1111/pbi.13611
Zhao H, Wang L, Zhao FJ, Wu L, Liu A, Xu W (2019) SpHMA1 is a chloroplast cadmium exporter protecting photochemical reactions in the Cd hyperaccumulator Sedum plumbizincicola. Plant Cell Environ 42(5):1112–1124. https://doi.org/10.1111/pce.13456
Zuo J, Niu QW, Chua NH (2000) Technical advance: an estrogen receptor-based transactivator XVE mediates highly inducible gene expression in transgenic plants. Plant J 24(2):265–273. https://doi.org/10.1046/j.1365-313x.2000.00868.x
Zuo J, Niu QW, Møller SG, Chua NH (2001) Chemical-regulated, site-specific DNA excision in transgenic plants. Nat Biotechnol 19(2):157–161. https://doi.org/10.1038/84428
Funding
This research was funded by the National Natural Science Foundation of China (32172666).
Author information
Authors and Affiliations
Contributions
WX and ZS conceived and designed research. YZ, YM, XW, and LH conducted experiments. YZ, HR and WX analyzed data. WX, ZS, and HR supervised the study. YZ wrote the draft of the manuscript. All authors read and approved the manuscript.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Additional information
Communicated by Dorothea Bartels.
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Zhang, Y., Mo, Y., Ren, H. et al. Improving Sedum plumbizincicola genetic transformation with the SpGRF4–SpGIF1 gene and the self-excision CRE/LoxP system. Planta 259, 119 (2024). https://doi.org/10.1007/s00425-024-04393-3
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s00425-024-04393-3