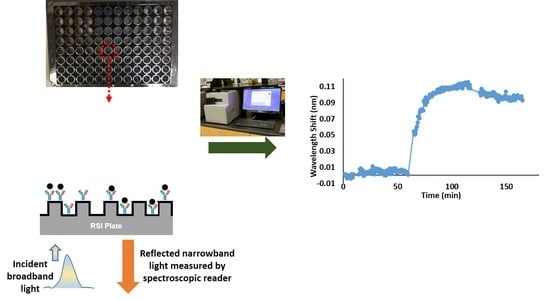
Quantification of Neuropeptide Y with Picomolar Sensitivity Enabled by Guided-Mode Resonance Biosensors
Abstract
:1. Introduction
2. Materials and Methods
2.1. Materials and Instruments
2.2. Sensor Plate Preparation
2.3. Surface Immobilization of Avidin-D
2.4. Surface Immobilization of Anti-NPY [Biotin]
2.5. Sandwich NPY Assay
3. Results and Discussion
3.1. Avidin-D Characterization
3.2. Anti-NPY Characterization
3.3. Sandwich NPY Assay
4. Conclusions
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References and Note
- Magnusson, R.; Wang, S.S. Optical Guided-Mode Resonance Filter. U.S. Patent 5,216,680, 1 June 1993. [Google Scholar]
- Magnusson, R.; Wang, S.S. New principle for optical filters. Appl. Phys. Lett. 1992, 61, 1022–1024. [Google Scholar] [CrossRef]
- Wawro, D.; Tibuleac, S.; Magnusson, R.; Liu, H. Optical fiber endface biosensor based on resonances in dielectric waveguide gratings. Proc. SPIE 2000, 3911, 86–94. [Google Scholar]
- Kikuta, H.; Maegawa, N.; Mizutani, A.; Iwata, K.; Toyota, H. Refractive index sensor with a guided-mode resonant grating filter. Proc. SPIE 2001, 4416, 219–222. [Google Scholar]
- Cunningham, B.; Li, P.; Lin, B.; Pepper, J. Colorimetric resonant reflection as a direct biochemical assay technique. Sens. Actuators B Chem. 2002, 81, 316–328. [Google Scholar] [CrossRef]
- Lin, S.; Ding, T.; Liu, J.; Lee, C.; Yang, T.; Chen, W.; Chang, J.A. A guided mode resonance aptasensor for thrombin detection. Sensors 2011, 11, 8953–8965. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Fang, Y.; Ferrie, A.M.; Li, G. Probing cytoskeleton modulation by optical biosensors. FEBS Lett. 2005, 579, 4175–4180. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Fang, Y.; Frutos, A.G.; Verklereen, R. Label-free cell-based assays for gpcr screening. Comb. Chem. High. Throughput Scr. 2008, 11, 357–369. [Google Scholar] [CrossRef] [Green Version]
- Erickson, J.C.; Clegg, K.E.; Palmiter, R.D. Sensitivity to leptin and susceptibility to seizures of mice lacking neuropeptide Y. Nature 1996, 381, 415–418. [Google Scholar] [CrossRef]
- Cizza, G.; Marques, A.H.; Eskandari, F.; Christie, I.C.; Torvik, S.; Silverman, M.N.; Sternberg, E.M. Elevated Neuroimmune Biomarkers in Sweat Patches and Plasma of Premenopausal Women with Major Depressive Disorder in Remission: The POWER Study. Biol. Psychiatry 2008, 64, 907–911. [Google Scholar] [CrossRef] [Green Version]
- Gruber, S.H.; Nomikos, G.G.; Mathé, A.A. Effects of acute and subchronic d-amphetamine on ventral striatal concentrations of neurotensin and neuropeptide Y in rats treated with antipsychotic drugs. Eur. Neuropsychopharmacol. 2006, 16, 592–600. [Google Scholar] [CrossRef]
- Kuo, L.E.; Zukowska, Z. Stress NPY and vascular remodeling: Implications for stress-related diseases. Peptides 2007, 28, 435–440. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Rasmusson, A.M.; Southwick, S.M.; Hauger, R.L.; Charney, D.S. Plasma neuropeptide Y (NPY) increases in humans in response to the α2 antagonist yohimbine. Neuropsychopharmacology 1998, 19, 95–98. [Google Scholar] [CrossRef]
- Morgan, C.A.; Rasmusson, A.M.; Wang, S.; Hoyt, G.; Hauger, R.L.; Hazlett, G. Neuropeptide-Y, cortisol, and subjective distress in humans exposed to acute stress: Replication and extension of previous report. Biol. Psychiatry 2002, 52, 136–142. [Google Scholar] [CrossRef]
- Tu, C.; Zhao, D.; Lin, X. Levels of neuropeptide-Y in the plasma and skin tissue fluids of patients with vitiligo. J. Derm. Sci. 2001, 27, 178–182. [Google Scholar] [CrossRef]
- Andrews, J.A.; Neises, K.D. Cells, biomarkers, and post-traumatic stress disorder: Evidence for peripheral involvement in a central disease, Peripheral cells, biomarkers, and PTSD. J. Neurochem. 2012, 120, 26–36. [Google Scholar] [CrossRef]
- Racaityte, K.; Lutz, E.; Unger, K.; Lubda, D.; Boos, K. Analysis of neuropeptide Y and its metabolites by high-performance liquid chromatography-electrospray ionization mass spectrometry and integrated sample clean-up with a novel restricted-access sulphonic acid cation exchanger. J. Chromatogr. A 2000, 890, 135–144. [Google Scholar] [CrossRef]
- Jia, M.; Belyavskaya, E.; Deuster, P.; Sternberg, E.M. Development of a Sensitive Microarray Immunoassay for the Quantitative Analysis of Neuropeptide, Y. Anal. Chem. 2012, 84, 6508–6514. [Google Scholar] [CrossRef]
- Fernandez, R.E.; Sanghavi, B.J.; Farmehini, V.; Chávez, J.L.; Hagen, J.; Kelley-Loughnane, N.; Swami, N.S. Aptamer-functionalized graphene-gold nanocomposites for label-free detection of dielectrophoretic-enriched neuropeptide Y. Electrochem. Commun. 2016, 72, 144–147. [Google Scholar] [CrossRef]
- Kim, S.; Xing, L.; Islam, A.E.; Hsiao, M.; Ngo, Y.; Pavlyuk, O.M.; Drummy, L.F. In Operando Observation of Neuropeptide Capture and Release on Graphene Field-Effect Transistor Biosensors with Picomolar Sensitivity. ACS Appl. Mater. Interfaces 2019, 11, 13927–13934. [Google Scholar] [CrossRef]
- Magnusson, R.; Abdallah, M.G.; Lee, K.J.; Buchanan-Vega, J.A.; Wenner, B.R.; Allen, J.W.; Allen, M.S.; Gimlin, S.; Weidanz, D.W. Guided-mode resonance sensing of neuropeptide-Y with a sandwich assay achieving pg/mL detection. Proc. SPIE 2019, 11020, 110200H-1–110200H-6. [Google Scholar]
- There is an unfortunate error in [21] in unit conversions. The detection level was stated in pg/mL; it should be ng/mL.
- Available online: http://www.resonantsensors.com (accessed on 12 December 2019).
- Moharam, M.G.; Gaylord, T.K.; Pommet, D.A.; Grann, E.B. Stable implementation of the rigorous coupled-wave analysis for surface-relief gratings: Enhanced transmittance matrix approach. J. Opt. Soc. Am. A 1995, 12, 1077. [Google Scholar] [CrossRef]
- Wang, J.; Karnaushenko, D.; Medina-Sánchez, M.; Yin, Y.; Ma, L.; Schmidt, O.G. Three-Dimensional Microtubular Devices for Lab-on-a-Chip Sensing Applications. ACS Sens. 2019, 4, 1476–1496. [Google Scholar] [CrossRef]
- Wan, Y.; Kim, Y.; Li, N.; Cho, S.K.; Bachoo, R.; Ellington, A.D.; Iqbal, S.M. Surface-Immobilized Aptamers for Cancer Cell Isolation and Microscopic Cytology. Cancer Res. 2010, 70, 9371–9380. [Google Scholar] [CrossRef] [Green Version]






© 2019 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Abdallah, M.G.; Buchanan-Vega, J.A.; Lee, K.J.; Wenner, B.R.; Allen, J.W.; Allen, M.S.; Gimlin, S.; Wawro Weidanz, D.; Magnusson, R. Quantification of Neuropeptide Y with Picomolar Sensitivity Enabled by Guided-Mode Resonance Biosensors. Sensors 2020, 20, 126. https://doi.org/10.3390/s20010126
Abdallah MG, Buchanan-Vega JA, Lee KJ, Wenner BR, Allen JW, Allen MS, Gimlin S, Wawro Weidanz D, Magnusson R. Quantification of Neuropeptide Y with Picomolar Sensitivity Enabled by Guided-Mode Resonance Biosensors. Sensors. 2020; 20(1):126. https://doi.org/10.3390/s20010126
Chicago/Turabian StyleAbdallah, Mohammad G., Joseph A. Buchanan-Vega, Kyu J. Lee, Brett R. Wenner, Jeffery W. Allen, Monica S. Allen, Susanne Gimlin, Debra Wawro Weidanz, and Robert Magnusson. 2020. "Quantification of Neuropeptide Y with Picomolar Sensitivity Enabled by Guided-Mode Resonance Biosensors" Sensors 20, no. 1: 126. https://doi.org/10.3390/s20010126