Abstract
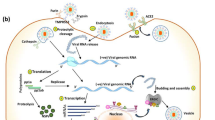
The Zinc finger antiviral proteins (ZAP) are expressed by the cells in response to different viral attacks. The ZAP marks the viral mRNA for degradation with the help of different cellular proteins for exosome. Beside the degradation of viral mRNA, the ZAP protein also inihibits translation of proteins from this mRNA. Although it is not a universal antiviral protein, but a number of important human pathogenic viruses are controlled by the ZAP protein. In this review article we have discussed the current progress made in understanding the structure, function, mechanism and therapeutic roles of the ZAP protein.


Similar content being viewed by others
REFERENCES
Berg, J.M., Zinc fingers and other metal-binding domains. Elements for interactions between macromolecules, J. Biol. Chem., 1990, vol. 265, pp. 6513–6516.
Liang, J., Song, W., Tromp, G., Kolattukudy, P.E., and Fu, M., Genome-wide survey and expression profiling of CCCH-zinc finger family reveals a functional module in macrophage activation, PLoS One, 2008, vol. 3, no. 8, p. e2880. https://doi.org/10.1371/journal.pone.0002880
Najafabadi, H.S., Mnaimneh, S., Schmitges, F.W., Garton, M., Lam, K.N., Yang, A., Albu, M., Weirauch, M.T., Radovani, E., Kim, P.M., Greenblatt, J., Frey, B.J., and Hughes, T.R., C2H2 zinc finger proteins greatly expand the human regulatory lexicon, Nat. Biotechnol., 2015, vol. 33, pp. 555–562. https://doi.org/10.1038/nbt.3128
Schmitges, F.W., Radovani, E., Najafabadi, H.S., Barazandeh, M., Campitelli, L.F., Yin, Y., Jolma, A., Zhong, G., Guo, H., Kanagalingam, T., Dai, W.F., Taipale, J., Emili, A., Greenblatt, J.F., and Hughes, T.R., Multiparameter functional diversity of human C2H2 zinc finger proteins, Genome Res., 2016, vol. 26, pp. 1742–1752. https://doi.org/10.1101/gr.209643.116
Klug, A., The discovery of zinc fingers and their applications in gene regulation and genome manipulation, Annu. Rev. Biochem., 2010, vol. 79, pp. 213–231. https://doi.org/10.1146/annurevbiochem-010909-095056
Mackay, J.P. and Crossley, M., Zinc fingers are sticking together, Trends Biochem. Sci., 1998, vol. 23, pp. 1–4.
Mao, R., Nie, H., Cai, D., Zhang, J., Liu, H., Yan, R., Cuconati, A., Block, T.M., Guo, J.T., and Guo, H., Inhibition of hepatitis B virus replication by the host zinc finger antiviral protein, PLoS Pathog., 2013, vol. 9, p. e1 003 494. https://doi.org/10.1371/journal.ppat.1003494
Fu, M. and Blackshear, P.J., RNA-binding proteins in immune regulation: A focus on CCCH zinc finger proteins, Nat. Rev. Immunol., 2016, vol. 17, pp. 130–143. https://doi.org/10.1038/nri.2016.129
Todorova, T., Bock, F.J., and Chang, P., PARP13 regulates cellular mRNA post-transcriptionally and functions as a pro-apoptotic factor by destabilizing TRAILR4 transcript, Nat. Commun., 2014, vol. 5, p. 5362. https://doi.org/10.1038/ncomms6362
Gao, G., Inhibition of tetroviral RNA production by ZAP, a CCCH-type zinc finger protein, Science, 2002, vol. 297, pp. 1703–1706. https://doi.org/10.1126/science.1074276
Kerns, J.A., Emerman, M., and Malik, H.S., Positive selection and increased antiviral activity associated with the PARP-containing isoform of human zinc-finger antiviral protein, PLoS Genet., 2008, vol. 4, pp. 0150–0158. https://doi.org/10.1371/journal.pgen.0040021
Vyas, S., Chesarone-Cataldo, M., Todorova, T., Huang, Y.-H., and Chang, P., A systematic analysis of the PARP protein family identifies new functions critical for cell physiology, Nat. Commun., 2013, vol. 4, p. 2240. https://doi.org/10.1038/ncomms3240
Welsby, I., Hutin, D., Gueydan, C., Kruys, V., Rongvaux, A., and Leo, O., PARP12, an interferon-stimulated gene involved in the control of protein translation and inflammation, J. Biol. Chem., 2014, vol. 289, pp 26642–26657. https://doi.org/10.1074/jbc.M114.589515
Zhu, Y. and Gao, G., ZAP-mediated mRNA degradation, RNA Biol., 2008, vol. 5, pp. 65–67. https://doi.org/10.4161/rna.5.2.6044
Gläsker, S., Töller, M., and Kümmerer, B.M., The alternate triad motif of the poly(ADP-ribose) polymerase-like domain of the human zinc finger antiviral protein is essential for its antiviral activity, J. Gen. Virol., 2014, vol. 95, pp. 816–822. https://doi.org/10.1099/vir.0.060988-0
Goodier, J.L., Pereira, G.C., Cheung, L.E., Rose, R.J., and Kazazian, H.H., The broad-spectrum antiviral protein ZAP restricts human retrotransposition, PLoS Genet., 2015, vol. 11, p. e1 005 252. https://doi.org/10.1371/journal.pgen.1005252
Lin, Y., Zhang, H., Liang, J., Li, K., Zhu, W., Fu, L., Wang, F., Zheng, X., Shi, H., Wu, S., Xiao, X., Chen, L., Tang, L., Yan, M., Yang, X., Tan, Y., Qiu, P., Huang, Y., Yin, W., Su, X., Hu, H., Hu, J., and Yan, G., Identification and characterization of alphavirus M1 as a selective oncolytic virus targeting ZAP-defective human cancers, Proc. Natl. Acad. Sci. U. S. A., 2014, vol. 111, pp. E4504–E4512. https://doi.org/10.1073/pnas.1408759111
Kaufman, H.L., Kohlhapp, F.J., and Zloza, A., Oncolytic viruses: a new class of immunotherapy drugs, Nat. Rev. Drug Discovery, 2015, vol. 14, pp. 642–662. https://doi.org/10.1038/nrd4663
Amé, J.C., Spenlehauer, C., and De Murcia, G., The PARP superfamily, BioEssays, 2004, vol. 26, pp. 882–893.
Guo, X., Carroll, J.-W.N., Macdonald, M.R., Goff, S.P., and Gao, G., The zinc finger antiviral protein directly binds to specific viral mRNAs through the CCCH zinc finger motifs, J. Virol., 2004, vol. 78, pp. 12 781–12 787. https://doi.org/10.1128/JVI.78.23.12781-12787.2004
Law, L.M.J., Albin, O.R., Carroll, J.W.N., Jones, C.T., Rice, C.M., and MacDonald, M.R., Identification of a dominant negative inhibitor of human zinc finger antiviral protein reveals a functional endogenous pool and critical homotypic interactions, J. Virol., 2010, vol. 84, pp. 4504–4512. https://doi.org/10.1128/JVI.02018-09
Chen, S., Xu, Y., Zhang, K., Wang, X., Sun, J., Gao, G., and Liu, Y., Structure of N-terminal domain of ZAP indicates how a zinc-finger protein recognizes complex RNA, Nat. Struct. Mol. Biol., 2012, vol. 19, pp. 430–435. https://doi.org/10.1038/nsmb.2243
Li, M.M.H., Lau, Z., Cheung, P., Aguilar, E.G., Schneider, W.M., Bozzacco, L., Molina, H., Buehler, E., Takaoka, A., Rice, C.M., Felsenfeld, D.P., and MacDonald, M.R., TRIM25 enhances the antiviral action of zinc-finger antiviral protein (ZAP), PLoS Pathog., 2017, vol. 13, p. e1 006 145. https://doi.org/10.1371/journal.ppat.1006145
Zheng, X., Wang, X., Tu, F., Wang, Q., Fan, Z., and Gao, G., TRIM25 is required for the antiviral activity of zinc-finger antiviral protein (ZAP), J. Virol., 2017, vol. 91, p. e00088-17. https://doi.org/10.1128/JVI.00088-17
Froshauer, S., Kartenbeck, J., and Helenius, A., Alphavirus RNA replicase is located on the cytoplasmic surface of endosomes and lysosomes, J. Cell Biol., 1988, vol. 107, pp. 2075–2086. https://doi.org/10.1371/journal.pbio.0050220
Frolova, E.I., Gorchakov, R., Pereboeva, L., Atasheva, S., and Frolov, I., Functional Sindbis virus replicative complexes are formed at the plasma membrane, J. Virol., 2010, vol. 84, pp. 11 679–11 695. https://doi.org/10.1128/jvi.01441-10
Guo, X., Ma, J., Sun, J., and Gao, G., The zinc-finger antiviral protein recruits the RNA processing exosome to degrade the target mRNA, Proc. Natl. Acad. Sci. U. S. A., 2007, vol. 104, pp. 151–156. https://doi.org/10.1073/pnas.0607063104
Chen, G., Guo, X., Lv, F., Xu, Y., and Gao, G., p72 DEAD box RNA helicase is required for optimal function of the zinc-finger antiviral protein, Proc. Natl. Acad. Sci. U. S. A., 2008, vol. 105, pp. 4352–4357. https://doi.org/10.1073/pnas.0712276105
Cordin, O., Banroques, J., Tanner, N.K., and Linder, P., The DEAD-box protein family of RNA helicases, Gene, 2006, vol. 367, pp. 17–37.
Linder, P. and Jankowsky, E., From unwinding to clamping: The DEAD box RNA helicase family, Nat. Rev. Mol. Cell Biol., 2011, vol. 12, pp. 505–516. https://doi.org/10.1038/nrm3154
Janknecht, R., Multi-talented dead-box proteins and potential tumor promoters: P68 RNA helicase (DDx5) and its paralog, p72 RNA helicase (DDx17), Am. J. Transl. Res., 2010, vol. 2, pp. 223–234.
Erazo, A. and Goff, S.P., Nuclear matrix protein Matrin 3 is a regulator of ZAP-mediated retroviral restriction, Retrovirology, 2015, vol. 12, p. 57. https://doi.org/10.1186/s12977-015-0182-4
Huang, Z., Wang, X., and Gao, G., Analyses of SELEX-derived ZAP-binding RNA aptamers suggest that the binding specificity is determined by both structure and sequence of the RNA, Protein Cell, 2010, vol. 1, pp. 752–759. https://doi.org/10.1007/s13238-010-0096-9
Wandtke, T., Woźniak, J., and Kopiński, P., Aptamers in diagnostics and treatment of viral infections, Viruses, 2015, vol. 7, pp. 751–780.
González, V.M., Martín, M.E., Fernández, G., and García-Sacristán, A., Use of aptamers as diagnostics tools and antiviral agents for human viruses, Pharmaceuticals, 2016, vol. 9, no. 4, p. 78.
Takata, M.A., Gonçalves-Carneiro, D., Zang, T.M., Soll, S.J., York, A., Blanco-Melo, D., and Bieniasz, P.D., CG dinucleotide suppression enables antiviral defense targeting non-self RNA, Nature, 2017, vol. 550, p. 7674. https://doi.org/10.1038/nature24039
Vacca, I., Viral infection: Adapt or get zapped, Nat. Rev. Microbiol., 2017, vol. 15, p. 641. https://doi.org/10.1038/nrmicro.2017.129
Zhu, Y., Chen, G., Lv, F., Wang, X., Ji, X., Xu, Y., Sun, J., Wu, L., Zheng, Y.-T., and Gao, G., Zinc-finger antiviral protein inhibits HIV-1 infection by selectively targeting multiply spliced viral mRNAs for degradation, Proc. Natl. Acad. Sci. U. S. A., 2011, vol. 108, pp. 15 834–15 839. https://doi.org/10.1073/pnas.1101676108
Goff, S.P., Evolution: Zapping viral RNAs, Nature, 2017, vol. 550, pp. 46–47. https://doi.org/10.1038/nature24140
Wang, L.F., Yu, M., Hansson, E., Pritchard, L.I., Shiell, B., Michalski, W.P., and Eaton, B.T., The exceptionally large genome of Hendra virus: Support for creation of a new genus within the family Paramyxoviridae, J. Virol., 2000, vol. 74, pp. 9972–9979. https://doi.org/10.1128/JVI.74.21.9972-9979.2000
Salladini, E., Delauzun, V., and Longhi, S., The Henipavirus V protein is a prevalently unfolded protein with a zinc-finger domain involved in binding to DDB1, Mol. BioSyst., 2017, vol. 13, pp. 2254–2267. https://doi.org/10.1039/C7MB00488E
Li, T., Chen, X., Garbutt, K.C., Zhou, P., and Zheng, N., Structure of DDB1 in complex with a paramyxovirus V protein: Viral Hijack of a propeller cluster in ubiquitin ligase, Cell, 2006, vol. 124, pp. 105–117. https://doi.org/10.1016/j.cell.2005.10.033
Lin, G.Y., Paterson, R.G., Richardson, C.D., and Lamb, R.A., The V protein of the paramyxovirus SV5 interacts with damage-specific DNA binding protein, Virology, 1998, vol. 249, pp. 189–200. https://doi.org/10.1006/viro.1998.9317
Bick, M.J., Carroll, J.-W.N., Gao, G., Goff, S.P., Rice, C.M., and MacDonald, M.R., Expression of the zinc-finger antiviral protein inhibits alphavirus replication, J. Virol., 2003, vol. 77, pp. 11 555–11 562. https://doi.org/10.1128/JVI.77.21.11555
Wang, X., Tu, F., Zhu, Y., and Gao, G., Zinc-finger antiviral protein inhibits XMRV infection, PLoS One, 2012, vol. 7, no. 6, p. e39 159. https://doi.org/10.1371/journal.pone.0039159
Switzer, W.M., Jia, H., Zheng, H.Q., Tang, S., and Heneine, W., No association of xenotropic murine leukemia virus-related viruses with prostate cancer, PLoS One, 2011, vol. 6, no. 5, p. e19 065. https://doi.org/10.1371/journal.pone.0019065
Hohn, O., Krause, H., Barbarotto, P., Niederstadt, L., Beimforde, N., Denner, J., Miller, K., Kurth, R., and Bannert, N., Lack of evidence for xenotropic murine leukemia virus-related virus (XMRV) in German prostate cancer patients, Retrovirology, 2009, vol. 6, p. 92. https://doi.org/10.1186/1742-4690-6-92
Furuta, R.A., Miyazawa, T., Sugiyama, T., Kuratsune, H., Ikeda, Y., Sato, E., Misawa, N., Nakatomi, Y., Sakuma, R., Yasui, K., Yamaguti, K., and Hirayama, F., No association of xenotropic murine leukemia virus-related virus with prostate cancer or chronic fatigue syndrome in Japan, Retrovirology, 2011, vol. 8, p. 20. https://doi.org/10.1186/1742-4690-8-20
Baraniuk, J.N., Xenotropic murine leukemia virus-related virus in chronic fatigue syndrome and prostate cancer, Curr. Allergy Asthma Rep., 2010, vol. 10, pp. 210–214.
Chen, E.Q., Dai, J., Bai, L., and Tang, H., The efficacy of zinc finger antiviral protein against hepatitis B virus transcription and replication in transgenic mouse model, Virol. J., 2015, vol. 12, p. 25. https://doi.org/10.1186/s12985-015-0245-0
Zhu, Y., Wang, X., Goff, S.P., and Gao, G., Translational repression precedes and is required for ZAP-mediated mRNA decay, EMBO J., 2012, vol. 31, pp. 4236–4246.
Schmid, M. and Jensen, T.H., The exosome: A multipurpose RNA-decay machine, Trends Biochem. Sci., 2008, vol. 33, pp. 501–510.
Lykke-Andersen, S., Brodersen, D.E., and Jensen, T.H., Origins and activities of the eukaryotic exosome, J. Cell Sci., 2009, vol. 122, pp. 1487–1494. https://doi.org/10.1242/jcs.047399
Chlebowski, A., Lubas, M., Jensen, T.H., and Dziembowski, A., RNA decay machines: The exosome, Biochim. Biophys. Acta, Gene Regul. Mech., 2013, vol. 1829, pp. 552–560.
Garzia, A., Jafarnejad, S.M., Meyer, C., Chapat, C., Gogakos, T., Morozov, P., Amiri, M., Shapiro, M., Molina, H., Tuschl, T., and Sonenberg, N., The E3 ubiquitin ligase and RNA-binding protein ZNF598 orchestrates ribosome quality control of premature polyadenylated mRNAs, Nat. Commun., 2017, vol. 8, p. 16 056. https://doi.org/10.1038/ncomms16056
Sundaramoorthy, E., Leonard, M., Mak, R., Liao, J., Fulzele, A., and Bennett, E.J., ZNF598 and RACK1 regulate mammalian ribosome-associated quality control function by mediating regulatory 40S ribosomal ubiquitylation, Mol. Cell, 2017, vol. 65, pp. 751–760. https://doi.org/10.1016/j.molcel.2016.12.026
Juszkiewicz, S. and Hegde, R.S., Initiation of quality control during poly(A) translation requires site-specific ribosome ubiquitination, Mol. Cell, 2017, vol. 65, pp. 743–750. https://doi.org/10.1016/j.molcel.2016.11.039
Cano, F., Rapiteanu, R., Sebastiaan Winkler, G., and Lehner, P.J., A non-proteolytic role for ubiquitin in deadenylation of MHC-I mRNA by the RNA-binding E3-ligase MEX-3C, Nat. Commun., 2015, vol. 6, p. 8670. https://doi.org/10.1038/ncomms9670
Oh, Y. and Chung, K.C., UHRF2, a ubiquitin E3 ligase, acts as a small ubiquitin-like modifier E3 ligase for zinc finger protein 131, J. Biol. Chem., 2013, vol. 288, pp. 9102–9111. https://doi.org/10.1074/jbc.M112.438234
Feng, Q., Jagannathan, S., and Bradley, R.K., The RNA surveillance factor UPF1 represses myogenesis via its E3 ubiquitin ligase activity, Mol. Cell, 2017, vol. 67, pp. 239–251. e6. https://doi.org/10.1016/j.molcel.2017.05.034
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
The authors declare that they have no conflict of interest. This article does not contain any studies involving animals or human participants performed by any of the authors.
About this article
Cite this article
Syed Lal Badshah, Ullah, A. & Syed, S. The Role of Zinc-Finger Antiviral Proteins in Immunity against Viruses. Mol. Genet. Microbiol. Virol. 35, 78–84 (2020). https://doi.org/10.3103/S0891416820020020
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.3103/S0891416820020020