Abstract
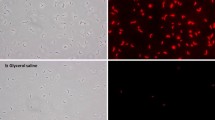
This study proposed a quenching protocol for metabolite analysis of Lactobacillus delbrueckii subsp. bulgaricus. Microbial cells were quenched with 60% methanol/water, 80% methanol/glycerol, or 80% methanol/water. The effect of the quenching process was assessed by the optical density (OD)-based method, flow cytometry, and gas chromatography-mass spectrometry (GC-MS). The principal component analysis (PCA) and orthogonal partial least squares-discriminant analysis (OPLS-DA) were employed for metabolite identification. The results indicated that quenching with 80% methanol/water solution led to less damage to the L. bulgaricus cells, characterized by the lower relative fraction of prodium iodide (PI)-labeled cells and the higher OD recovery ratio. Through GC-MS analysis, higher levels of intracellular metabolites (including focal glutamic acid, aspartic acid, alanine, and AMP) and a lower leakage rate were detected in the sample quenched with 80% methanol/water compared with the others. In conclusion, we suggested a higher concentration of cold methanol quenching for L. bulgaricus metabolomics due to its decreasing metabolite leakage.
概要
研究目的
为保加利亚乳杆菌的代谢组学研究提供一种简单有效的淬灭方案。
创新要点
采用适当的淬灭技术是获得微生物真实代谢物组数据的必要条件。 本研究首次建立了适用于保加利亚乳杆菌代谢组学研究的淬灭方法, 有助于完善乳酸菌代谢的生理学和遗传学概貌。
研究方法
分别应用 −40 °C 的60%甲醇/水、 80%甲醇/水和80%甲醇/甘油三种方式淬灭保加利亚乳杆菌。 利用光密度(OD)回收率实验、 流式细胞术和气质联用(GC-MS)分析菌体细胞完整性和细胞内外代谢物, 以及主成分分析法(PCA)和正交偏最小二乘法-判别分析法(OPLS-DA)分析代谢物泄露程度。
重要结论
80%冷甲醇/水更适用于淬灭保加利亚乳杆菌, 能有效减少代谢物泄露程度和增加胞内代谢物水平。
Article PDF
Similar content being viewed by others
References
Bolten, C.J., Kiefer, P., Letisse, F., et al., 2007. Sampling for metabolome analysis of microorganisms. Anal. Chem., 79(10):3843–3849. [doi:10.1021/ac0623888]
Canelas, A.B., Ras, C., ten Pierick, A., et al., 2008. Leakage-free rapid quenching technique for yeast metabolomics. Metabolomics, 4(3):226–239. [doi:10.1007/s11306-008-0116-4]
Carvalho, A.S., Silva, J., Ho, P., et al., 2004. Relevant factors for the preparation of freeze-dried lactic acid bacteria. Int. Dairy J., 14(10):835–847. [doi:10.1016/j.idairyj.2004.02. 001]
Castrillo, J.I., Hayes, A., Mohmmed, S., et al., 2003. An optimized protocol for metabolome analysis in yeast using direct infusion electrospray mass spectrometry. Phytochemistry, 62(6):929–937. [doi:10.1016/S0031-9422(02)00713-6]
Coulier, L., Bas, R., Jespersen, S., et al., 2006. Simultaneous quantitative analysis of metabolites using ion-pair liquid chromatography-electrospray ionization mass spectrometry. Anal. Biochem., 78(18):6573–6582. [doi:10.1021/ac0607616]
Díaz, M., Herrero, M., García, L.A., et al., 2010. Application of flow cytometry to industrial microbial bioprocesses. Biochem. Eng. J., 48(3):385–407. [doi:10.1016/j.bej.2009.07.013]
Duetz, W.A., Witholt, B., 2004. Oxygen transfer by orbital shaking of square vessels and deep well microtiter plates of various dimensions. Biochem. Eng. J., 17(3):181–185. [doi:10.1016/S1369-703X(03)00177-3]
Faijes, M., Mars, A.E., Smid, E.J., 2007. Comparison of quenching and extraction methodologies for metabolome analysis of Lactobacillus plantarum. Microb. Cell Fact., 6:27. [doi:10.1186/1475-2859-6-27]
Fonseca, F., Marin, M., Morris, G.J., 2006. Stabilization of frozen Lactobacillus delbrueckii subsp. bulgaricus in glycerol suspensions: freezing kinetics and storage temperature effects. Appl. Environ. Microbiol., 72(10): 6474–6482. [doi:10.1128/AEM.00998-06]
Garbayo, I., Vílchez, C., Vega, J.M., et al., 2004. Influence of immobilization parameters on growth and lactic acid production by Streptococcus thermophilus and Lactobacillus bulgaricus co-immobilized in calcium alginate gel beads. Biotechnol. Lett., 26(23):1825–1827. [doi:10.1007/s10529-004-5134-3]
Huang, L., Lu, Z., Yuan, Y., et al., 2006. Optimization of a protective medium for enhancing the viability of freezedried Lactobacillus delbrueckii subsp. bulgaricus based on response surface methodology. J. Ind. Microbiol. Biotechnol., 33(1):55–61. [doi:10.1007/s10295-005-0041-8]
Jana, S., Lorenz, C.R., Patricia, W., et al., 2009. A method for enzyme quenching in microbial metabolome analysis successfully applied to Gram-positive and Gram-negative bacteria and yeast. Anal. Biochem., 394(2):192–201. [doi:10.1016/j.ab.2009.07.016]
Koek, M.M., Muilwijk, B., van der Werf, M.J., et al., 2006. Microbial metabolomics with gas chromatography/mass spectrometry. Anal. Chem., 78(4):1272–1281. [doi:10. 1021/ac051683+]
Kumar, S., Wittmann, C., Heinzle, E., 2004. Minibioreactors. Biotechnol. Lett., 26(1):1–10. [doi:10.1023/B:BILE.0000009469.69116.03]
Lange, H.C., Eman, M., van Zuijlen, G., et al., 2001. Improved rapid sampling for in vivo kinetics of intracellular metabolites in Saccharomyces cerevisiae. Biotechnol. Bioeng., 75(4):406–415. [doi:10.1002/bit.10048]
Li, C., Zhao, J.L., Wang, Y.T., et al., 2009. Synthesis of cyclopropane fatty acid and its effect on freeze-drying survival of Lactobacillus bulgaricus L2 at different growth conditions. World J. Microbiol. Biotechnol., 25(9): 1659–1665. [doi:10.1007/s11274-009-0060-0]
Link, H., Anselment, B., Weuster-Botz, D., 2008. Leakage of adenylates during cold methanol/glycerol quenching of Escherichia coli. Metabolomics, 4(3):240–247. [doi:10. 1007/s11306-008-0114-6]
Meyer, H., Liebeke, M., Lalk, M., 2010. A protocol for the investigation of the intracellular Staphylococcus aureus metabolome. Anal. Biochem., 401(2):250–259. [doi:10. 1016/j.ab.2010.03.003]
Nielsen, J., 1997. Metabolic control analysis of biochemical pathways based on a thermokinetic description of reaction rates. Biochem. J., 321:133–138.
Oldiges, M., Lütz, S., Pflug, S., et al., 2007. Metabolomics: current state and evolving methodologies and tools. Appl. Microbiol. Biotechnol., 76(3):495–511. [doi:10.1007/s00253-007-1029-2]
Otto, R., Brink, B., Veldkamp, H., et al., 1983. The relation between growth rate and electrochemical proton gradient of Streptococcus cremoris. FEMS Microbiol. Lett., 16(1): 69–74. [doi:10.1111/j.1574-6968.1983.tb00261.x]
Russell, J.B., Diez-Gonzalez, F., 1998. The effects of fermentation acids on bacterial growth. Adv. Microb. Physiol., 39:205–234. [doi:10.1016/S0065-2911(08)60017-X]
Schädel, F., David, F., Franco-Lara, E., 2011. Evaluation of cell damage caused by cold sampling and quenching for metabolome analysis. Appl. Microbiol. Biotechnol., 92(6): 1261–1274.
Schaefer, U., Boos, W., Takors, R., et al., 1999. Automated sampling device for monitoring intracellular metabolites dynamics. Anal. Biochem., 270(1):88–96. [doi:10.1006/abio.1999.4048]
Schiraldi, C., Valli, C., Molinaro, A., et al., 2006. Exopolysaccharides production in Lactobacillus bulgaricus and Lactobacillus casei exploiting microfiltration. J. Ind. Microbiol. Biotechnol., 33(5):384–390. [doi:10.1007/s10295-005-0068-x]
Siegumfeldt, H., Rechinger, K.B., Jakobsen, M., 2000. Dynamic changes of intracellular pH in individual lactic acid bacteria cells in response to a rapid drop in extracellular pH. Appl. Environ. Microbiol., 66(6):2330–2335. [doi:10.1128/AEM.66.6.2330-2335.2000]
Spura, J., Reimer, L.C., Wieloch, P., et al., 2009. A method for enzyme quenching in microbial metabolome analysis successfully applied to Gram-positive and Gram-negative bacteria and yeast. Anal. Biochem., 394(2):192–201. [doi:10.1016/j.ab.2009.07.016]
Tang, Y.J., Martin, H.G., Myers, S., et al., 2009. Advances in analysis of microbial metabolic fluxes via 13C isotopic labeling. Mass Spectrom. Rev., 28(2):362–375. [doi:10.1002/mas.20191]
van Dam, J.C., Eman, M.R., Frank, J., et al., 2002. Analysis of glycolytic intermediates in Saccharomyces cerevisiae using anion exchange chromatography and electrospray ionisation with tandem mass spectrometric detection. Anal. Chim. Acta, 460(2):209–218. [doi:10.1016/S0003-2670(02)00240-4]
Villas-Bôas, S.G., Bruheim, P., 2007. Cold glycerol-saline: the promising quenching solution for accurate intracellular metabolite analysis of microbial cells. Anal. Biochem., 370(1):87–97. [doi:10.1016/j.ab.2007.06.028]
Wittmann, C., Krömer, J.O., Kiefer, P., et al., 2004. Impact of the cold shock phenomenon on quantification of intracellular metabolites in bacteria. Anal. Biochem., 327(1): 135–139. [doi:10.1016/j.ab.2004.01.002]
Wu, H., Southam, A.D., Hines, A., et al., 2008. High-throughput tissue extraction protocol for NMR- and MS-based metabolomics. Anal. Biochem., 372(2):204–212. [doi:10.1016/j.ab.2007.10.002]
Author information
Authors and Affiliations
Corresponding author
Additional information
Project supported by the Open Research Fund for the MOE Key Laboratory of Dairy Science (No. 2012KLDSOF-07), the National Natural Science Foundation of China (No. 31000808), the Synergetic Innovation Center of Food Safety and Nutrition, and the Open Research Fund of Northeast Agricultural University, China
Rights and permissions
About this article
Cite this article
Chen, Mm., Li, Al., Sun, Mc. et al. Optimization of the quenching method for metabolomics analysis of Lactobacillus bulgaricus . J. Zhejiang Univ. Sci. B 15, 333–342 (2014). https://doi.org/10.1631/jzus.B1300149
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1631/jzus.B1300149