Abstract
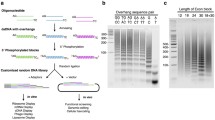
Random peptide libraries can be constructed either by in vitro synthesis of random peptides, or through translation of DNA sequences from synthetic random oligonucleotides. Here we describe an alternative way of making arbitrary peptide libraries with high diversity that can be used in screening as random peptide libraries. Genomic DNA digested with a frequent-cutting restriction enzyme recognizing four nucleotides will theoretically consist of small DNA pieces with average length of 256 nucleotides, and on average around 107 fragments can be generated from a genome of 3 × 109 bases. A peptide library translated from these fragments will have sufficient diversity for some protein interaction screening experiments. Moreover, the same genome digested with a different four-cutter enzyme or ligated into different reading frames will result in different nonoverlapping libraries. A series of such libraries could be generated with genomic DNAs from different species. In this study, human genomic DNA was digested with four-cutter restriction enzymes DpnII and Tsp509I, respectively, and cloned into yeast expression vector pGADT7 to generate arbitrary peptide libraries. These libraries were used in yeast two-hybrid assays to screen for binding motifs of the PDZ domain containing protein synectin. Our results showed that in addition to various native carboxy-terminal tails, synectin could also bind to many artificial ones, some of which contained a consensus sequence—(S/T)XC-COOH.
Similar content being viewed by others
References
Yang, M., Wu, Z., and Fields, S. (1995) Protein-peptide interactions analyzed with the yeast two-hybrid system. Nucleic Acids Res. 23, 1152–1156.
Fodor, S. P., Read, J. L., Pirrung, M. C., Stryer, L., Lu, A. T., and Solas, D. (1991) Light-directed, spatially addressable parallel chemical synthesis. Science 251, 767–773.
Houghten, R. A., Pinilla, C., Blondelle, S. E., Appel, J. R., Dooley, C. T., and Cuervo, J. H. (1991) Generation and use of synthetic peptide combinatorial libraries for basic research and drug discovery. Nature 354, 84–86.
Lam, K. S., Salmon, S. E., Hersh, E. M., Hruby, V. J., Kazmierski, W. M., and Knapp, R. J. (1991) A new type of synthetic peptide library for identifying ligand-binding activity. Nature 354, 82–84.
Parmley, S. F. and Smith, G. P. (1989) Filamentous fusion phage cloning vectors for the study of epitopes and design of vaccines. Adv. Exp. Med. Biol. 251, 215–218.
Cwirla, S. E., Peters, E. A., Barrett, R. W., and Dower, W. J. (1990) Peptides on phage: a vast library of peptides for identifying ligands. Proc. Natl. Acad. Sci. USA 87, 6378–6382.
Scott, J. K. and Smith, G. P. (1990) Searching for peptide ligands with an epitope library. Science 249, 386–390.
Venter, J. C., Adams, M. D., Myers, E. W., Li, P. W., Mural, R. J., Sutton, G. G., et al. (2001) The sequence of the human genome. Science 291, 1304–1351.
Hung, A. Y. and Sheng, M. (2002) PDZ domains: structural modules for protein complex assembly. J. Biol. Chem. 277, 5699–5702.
Songyang, Z., Fanning, A. S., Fu, C., Xu, J., Marfatia, S. M., Chishti, A. H., et al. (1997) Recognition of unique carboxyl-terminal motifs by distinct PDZ domains. Science 275, 73–77.
Vaccaro, P., Brannetti, B., Montecchi-Palazzi, L., Philipp, S., Helmer Citterich, M., Cesareni, G., and Dente, L. (2001) Distinct binding specificity of the multiple PDZ domains of INADL, a human protein with homology to INAD from Drosophila melanogaster. J. Biol. Chem. 276, 42122–42130.
Sambrook, J., Fritsch, E. F., and Maniatis, T., eds (1989) Molecular Cloning: A Laboratory Manual, 2nd Edition. Cold Spring Harbor Laboratory, Cold Spring Harbor, NY.
Gao, Y., Li, M., Chen, W., and Simons, M. (2000) Synectin, syndecan-4 cytoplasmic domain binding PDZ protein, inhibits cell migration. J. Cell Physiol. 184, 373–379.
Wang, L. H., Kalb, R. G., and Strittmatter, S. M. (1999) A PDZ protein regulates the distribution of the transmembrane semaphorin, M-SemF. J. Biol. Chem. 274, 14137–14146.
De Vries, L., Lou, X., Zhao, G., Zheng, B., and Farquhar, M. G. (1998) GIPC, a PDZ domain containing protein, interacts specifically with the C terminus of RGS-GAIP. Proc. Natl. Acad. Sci. USA 95, 12340–12345.
Bunn, R. C., Jensen, M. A., and Reed, B. C. (1999). Protein interactions with the glucose transporter binding protein GLUT1CBP that provide a link between GLUT1 and the cytoskeleton. Mol. Biol. Cell 10, 819–832.
Rousset, R., Fabre, S., Desbois, C., Bantignies, F., and Jalinot, P. (1998) The C-terminus of the HTLV-1 Tax oncoprotein mediates interaction with the PDZ domain of cellular proteins. Oncogene 16, 643–654.
Maximov, A., Sudhof, T. C., and Bezprozvanny, I. (1999) Association of neuronal calcium channels with modular adaptor proteins. J. Biol. Chem. 274, 24453–24456.
Murthy, K. K., Clark, K., Fortin, Y., Shen, S. H., and Banville, D. (1999) ZRP-1, a zyxin-related protein, interacts with the second PDZ domain of the cytosolic protein tyrosine phosphatase hPTP1E. J. Biol. Chem. 274, 20679–20687.
Ligensa, T., Krauss, S., Demuth, D., Schumacher, R., Camonis, J., Jaques, G., and Weidner, K. M. (2001). A PDZ domain protein interacts with the C-terminal tail of the insulin-like growth factor-1 receptor but not with the insulin receptor. J. Biol. Chem. 276, 33419–33427.
Huang, H., Zhang, L., Ma, S., and Gao, Y. (2004) Finding potential ligands for PDZ domains by tailfit. Chinese Med. Sci. J. 19, 97–100.
Booth, R. A., Cummings, C., Tiberi, M., and Liu, X. J. (2002) GIPC participates in G protein signaling downstream of insulin-like growth factor 1 receptor. J. Biol. Chem. 277, 6719–6725.
Hirakawa, T., Galet, C., Kishi, M., and Ascoli, M. (2003) GIPC binds to the human lutropin receptor (hLHR) through an unusual PDZ domain binding motif, and it regulates the sorting of the internalized human choriogonadotropin and the density of cell surface hLHR. J. Biol. Chem. 278, 49348–49357.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Huang, H., Gao, Y. A method for generation of arbitrary peptide libraries using genomic DNA. Mol Biotechnol 30, 135–142 (2005). https://doi.org/10.1385/MB:30:2:135
Issue Date:
DOI: https://doi.org/10.1385/MB:30:2:135