Manuscript accepted on : 17-03-2022
Published online on: 21-03-2022
Plagiarism Check: Yes
Reviewed by: Dr. Sadiya Parveen
Second Review by: Dr. Van-An Duong
Final Approval by: Dr. Fernando José Cebola Lidon
In Silico Analysis of Off-Target Effects of Ivermectin Drug
Simran Walia and Poonam Sharma
and Poonam Sharma
Department of Zoology, Gargi College, University of Delhi, Delhi, India
Corresponding Author E-mail: poonam.sharma@gargi.du.ac.in
DOI : http://dx.doi.org/10.13005/bbra/2970
ABSTRACT:
Researchers all across the world are repurposing medications to fight the coronavirus, but they come with a plethora of negative side effects. Ivermectin, a common repurposed medicine, binds to the importin protein in Covid-19 patients and suppresses viral replication. Ivermectin also binds to pentameric ligand-gated ion channels, increasing cell membrane ion permeability and triggering cell hyperpolarization. In this study, in silico analysis of non-target proteins of ivermectin and protein interactions was performed to better understand its off-target effects on other biological processes. Detailed information on the drug, its target and non-target proteins, their properties, protein-protein interactions, and pathways involved was analysed using databases such as DrugBank, NCBI Gene Database, BLAST, UCSC Gene Sorter, GeneMANIA, STRING, Kyoto Encyclopaedia of Genes and Genomes (KEGG) Pathway Database, and Reactome. Due to structural similarities and protein interactions with pentameric ligand-gated ion channels like α1β2γ2L GABA (A) receptors, GLRA-3 receptor, α7 nAChR, P2X4 cation channel, and IMPα/β1, ivermectin was found to bind to non-target protein families, GLR, GABA, nAChR, 5-HT receptors and, P2XR and ZACN, IPO5, RANBP6, TNPO1 and, TNPO2 proteins. These non-target proteins include neurotransmitter-gated ion channels, nuclear receptors, and transporter proteins, and they can interfere with signal transmission and neuroactive ligand-receptor interactions, as well as alter the function of proteins that interact with these target proteins indirectly. Off-target effects of ivermectin can be hypotension, visual hallucinations, loss of coordination and balance, depression, and neurological disorders. These findings highlight the need for a comprehensive evaluation of all repurposing drugs for their off-target effects before public use.
KEYWORDS: Covid-19; Ivermectin; Importin; Glycine receptors; GABA
Download this article as:| Copy the following to cite this article: Walia S, Sharma P. In Silico Analysis of Off-Target Effects of Ivermectin Drug . Biosci Biotech Res Asia 2022;19(1). |
| Copy the following to cite this URL: Walia S, Sharma P. In Silico Analysis of Off-Target Effects of Ivermectin Drug . Biosci Biotech Res Asia 2022;19(1). Available from: https://bit.ly/3L30GiS |
Introduction
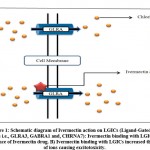
COVID-19 is the worldwide rapidly spreading illness caused by the severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2), first discovered in Wuhan, Hubei, China, in December 2019. As of February 20, 2022, WHO COVID-19 dashboard recorded 430,257,564 confirmed cases of COVID-19, with 5,922,049 deaths fatalities reported globally1. Scientists are constantly working on developing effective vaccines and medicines for the safe cure of COVID-19, but this is a very expensive and time-consuming procedure, thus drug repurposing gives an instant productive method for rapid and successful therapy to cope up with various strains of virus2. Ivermectin is a broad-spectrum antiparasitic drug authorized by the FDA for the treatment of onchocerciasis (river blindness), intestinal strongyloidiasis, head lice, and rosacea. It has a complex structure with a series of macrocyclic lactone isomers. It can be taken orally or directly administered to the skin, depending on the kind of illness. In invertebrates, it preferentially binds to muscle and nerve cells, glutamate-gated chloride channels, producing enhanced chloride ion membrane permeability and cell hyperpolarization and finally leading to parasite death3. Glycine receptors (GLRA3), γ-aminobutyric acid type A receptors (α1β2γ2LGABA A receptors), and α7 nicotinic acetylcholine receptors (nAChRs) are among the pentameric ligand-gated ion channels (pLGICs) that ivermectin and related drugs bind to and either directly activate or favourably regulate them resulting in increased permeability of the cell membrane to ions thus causing cell hyperpolarisation4,5,6 (Figure. 1).
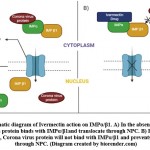
Ivermectin also stimulates the purinergic receptors (P2X4) cation channel involved in modulation of neurotransmission57. According to recent research, adding ivermectin to SARS-CoV-2-infected Vero-hSLAM cells blocks coronavirus entrance into the nucleus via the nuclear pore complex (NPC) because it binds to importin (IMPa/b1/ karyopherins), thus making it unstable to bind with coronavirus cargo protein8 (Figure. 2). Because most repurposing medications have a variety of mild to severe short- and long-term unfavourable side effects, this study was conducted to better understand ivermectin’s off-target effects by identifying and analysing various structurally comparable and interacting non-target proteins, and to open up the possibility of investigating in silico behaviour of ivermectin drug for effective pathological management and side effect management.
Materials and Methods
DrugBank was used to extract detailed information of ivermectin drug used for COVID-19 treatment. It provided information regarding DrugBank accession number, type, groups, weight, chemical formula, structure, and mode of action in the body, toxicity, absorption, and metabolism9. NCBI Gene Database was used to collect data on gene type, expression, function, interactions and for the amino acid sequences of α1β2γ2L GABA (A) receptors, GLRA-3 receptor, α7nAChR, P2X4 cation channel, and IMPα/β1, which are directly targeted by ivermectin. Next, Homo sapiens (human) Protein BLAST and UCSC Gene Sorter were used to identify proteins similar to these target proteins, and “Distance tree of results” was studied which gave the visual representation of structurally related proteins10. GeneMANIA and STRING were used to retrieve interactions in proteins, next detailed study was carried to gain their functional information11,12. For pathway analysis, Kyoto Encyclopaedia of Genes and Genomes (KEGG) Pathway Database and Reactome were used13.
Table 1: Important Non-target Proteins with a Similar Structure to GABRA1, CHRNA7, GLRA3, P2RX4, and KPNB1/IMB1
| S. No | Proteins | Blastp E-Value | Genome Position | ||
| Non-target proteins with a similar structure to GABRA1 | |||||
| 1 | GABRA2 | 0 | Chromosome 4, NC_000004.12 | ||
| 2 | GABRA3 | 6.00E-176 | Chromosome X, NC_000023.11 | ||
| 6 | GABRG1 | 2.00E-99 | Chromosome 4, NC_000004.12 | ||
| 7 | GABRG3 | 2.00E-92 | Chromosome 15, NC_000015.10 | ||
| 8 | GABRE | 9.00E-79 | Chromosome X, NC_000023.11 | ||
| 9 | GLRA2 | 4.00E-69 | Chromosome X, NC_000023.11 | ||
| 10 | GABRB1 | 2.00E-67 | Chromosome 4, NC_000004.12 | ||
| 12 | GLRA1 | 4.00E-65 | Chromosome 5, NC_000005.10 | ||
| 14 | GABRD | 2.00E-61 | Chromosome 1, NC_000001.11 | ||
| 15 | GABRQ | 2.00E-59 | Chromosome X, NC_000023.11 | ||
| 16 | GABRP | 1.00E-57 | Chromosome 5, NC_000005.10 | ||
| 19 | GLRB | 1.00E-48 | Chromosome 4, NC_000004.12 | ||
| 20 | CHRNB4 | 2.00E-16 | Chromosome 15, NC_000015.10 | ||
| 21 | HTR3B | 1.00E-15 | Chromosome 11, NC_000011.10 | ||
| Non-target proteins with a similar structure to CHRNA7 | |||||
| 1 | CHRFAM7A | 0 | Chromosome 15, NC_000015.10 | ||
| 2 | CHRNA2 | 4.00E-89 | Chromosome 8, NC_000008.11 | ||
| 3 | CHRNA3 | 7.00E-85 | Chromosome 15, NC_000015.10 | ||
| 4 | CHRNB4 | 1.00E-81 | Chromosome 15, NC_000015.10 | ||
| 5 | CHRNA10 | 9.00E-80 | Chromosome 11, NC_000011.10 | ||
| 6 | CHRNA6 | 2.00E-78 | Chromosome 8, NC_000008.11 | ||
| 7 | CHRNB2 | 3.00E-77 | Chromosome 1, NC_000001.11 | ||
| 8 | CHRNA4 | 6.00E-77 | Chromosome 20, NC_000020.11 | ||
| 10 | CHRNA1 | 5.00E-75 | Chromosome 2, NC_000002.12 | ||
| 11 | CHRNB3 | 2.00E-66 | Chromosome 8, NC_000008.11 | ||
| 12 | CHRNA5 | 7.00E-64 | Chromosome 15, NC_000015.10 | ||
| 13 | CHRNB1 | 8.00E-61 | Chromosome 17, NC_000017.11 | ||
| 14 | CHRND | 9.00E-56 | Chromosome 2, NC_000002.12 | ||
| 15 | CHRNG | 9.00E-51 | Chromosome 2, NC_000002.12 | ||
| 16 | CHRNE | 1.00E-46 | Chromosome 17, NC_000017.11 | ||
| 17 | HTR3A | 1.00E-41 | Chromosome 11, NC_000011.10 | ||
| 21 | GABRP | 2.00E-10 | Chromosome 5, NC_000005.10 | ||
| 22 | GABRG1 | 8.00E-10 | Chromosome 4, NC_000004.12 | ||
| 23 | ZACN | 1.00E-09 | Chromosome 17, NC_000017.11 | ||
| 24 | GLRA1 | 7.00E-09 | Chromosome 5, NC_000005.10 | ||
| Non-target proteins with a similar structure to P2RX4 | |||||
| 1 | P2RX1 | 6.00E-97 | Chromosome 17, NC_000017.11 | ||
| 2 | P2RX3 | 8.00E-84 | Chromosome 11, NC_000011.10 | ||
| 6 | P2RX7 | 1.00E-81 | Chromosome 12, NC_000012.12 | ||
| Non-target proteins with a similar structure to GLRA3 | |||||
| 1 | GLRA1 | 0 | Chromosome 5, NC_000005.10 | ||
| 3 | GLRA4 | 2.00E-163 | Chromosome X, NC_000023.11 | ||
| 4 | GLRB | 7.00E-97 | Chromosome 4, NC_000004.12 | ||
| 5 | GABRB3 | 3.00E-75 | Chromosome 15, NC_000015.10 | ||
| 6 | GABRB1 | 1.00E-73 | Chromosome 4, NC_000004.12 | ||
| 7 | GABRD | 6.00E-73 | Chromosome 1, NC_000001.11 | ||
| 8 | GABRA6 | 7.00E-71 | Chromosome 5, NC_000005.10 | ||
| 9 | GABRP | 7.00E-71 | Chromosome 5, NC_000005.10 | ||
| 10 | GABRR2 | 1.00E-70 | Chromosome 6, NC_000006.12 | ||
| 11 | GABRA2 | 7.00E-70 | Chromosome 4, NC_000004.12 | ||
| 14 | GABRR1 | 2.00E-68 | Chromosome 6, NC_000006.12 | ||
| 15 | GABRG1 | 1.00E-67 | Chromosome 4, NC_000004.12 | ||
| 16 | GABRG3 | 1.00E-65 | Chromosome 15, NC_000015.10 | ||
| 18 | GABRQ | 6.00E-64 | Chromosome X, NC_000023.11 | ||
| 19 | GABRE | 1.00E-58 | Chromosome X, NC_000023.11 | ||
| 20 | CHRNA6 | 2.00E-14 | Chromosome 8, NC_000008.11 | ||
| 21 | CHRNB1 | 8.00E-13 | Chromosome 17, NC_000017.11 | ||
| Non-target proteins with similar structure to KPNB1/IMB1 | |||||
| 1 | TNPO1 | 9.00E-18 | Chromosome 5, NC_000005.10 | ||
| 2 | TNPO2 | 6.00E-24 | Chromosome 19, NC_000019.10 | ||
| 3 | IPO5 | 8.00E-08 | Chromosome 13 – NC_000013.11 | ||
| 4 | RANBP6 | 0.00005 | Chromosome 9, NC_000009.12 | ||
Table 2: Protein-Protein Interaction developed using STRING Database
| Proteins interacting with GABRA1, GABRB2 and, GABRG2 | |
| GABRA4 | Gamma-amino butyric acid receptor subunit alpha-4 |
| GABRA5 | Gamma-aminobutyric acid receptor subunit alpha-5 |
| GABRA6 | Gamma-aminobutyric acid receptor subunit alpha-6 |
| GABRB3 | Gamma-aminobutyric acid receptor subunit beta-3 |
| GPHN | Gephyrin |
| DNM1 | Dynamin-1 |
| GABRB1 | Gamma-aminobutyric acid receptor subunit beta-1 |
| UBQLN1 | Ubiquilin-1 |
| Proteins interacting with CHRNA7 | |
| CHRNA2 | Neuronal acetylcholine receptor subunit alpha-2 |
| CHRNA3 | Neuronal acetylcholine receptor subunit alpha-3 |
| CHRNA4 | Neuronal acetylcholine receptor subunit alpha-4 |
| CHRNA9 | Neuronal acetylcholine receptor subunit alpha-9 |
| CHRNB2 | Neuronal acetylcholine receptor subunit beta-2 |
| CHRNB4 | Neuronal acetylcholine receptor subunit beta-4 |
| APP | Amyloid-beta A4 protein |
| MTMR10 | Myotubularin-related protein 10 |
| LYNX1 | Ly-6/neurotoxin-like protein 1 |
| SLURP1 | Secreted Ly-6/uPAR-related protein 1 |
| Proteins interacting with P2RX4 | |
| P2RX1 | P2X purinoceptor 1 |
| P2RX7 | P2X purinoceptor 7 |
| P2RX6 | P2X purinoceptor 6 |
| P2RY1 | P2Y purinoceptor 1 |
| P2RY2 | P2Y purinoceptor 2 |
| P2RY4 | P2Y purinoceptor 4 |
| P2RY6 | P2Y purinoceptor 6 |
| P2RY12 | P2Y purinoceptor 12 |
| PANX1 | Pannexin-1 |
| GRIN2A | Glutamate receptor ionotropic, NMDA 2A |
| GRIN1 | Glutamate receptor ionotropic, NMDA 1 |
| Proteins interacting with GLRA3 | |
| GARS | Glycine–tRNA ligase |
| GLRB | Glycine receptor subunit beta |
| GABRB3 | Gamma-amino butyric acid receptor subunit beta-3 |
| BEST1 | Bestrophin-1 |
| ANO1 | Anoctamin-1 |
| ANO6 | Anoctamin-6 |
| GPHIN | Gephyrin |
| NBEA | Neurobeachin |
| CABLES1 | CDK5 and ABL1 enzyme substrate 1 |
| NDUFAF6 | NADH dehydrogenase [ubiquinone] 1 alpha sub complex assembly factor 6 |
| Proteins interacting with KPNB1 AND KPNA1 | |
| KPNA2 | Karyopherin subunit alpha 2 |
| SNUPN | Snurportin-1 |
| NUP98 | Nuclear pore complex protein Nup98-Nup96 |
| NUP62 | Nuclear pore glycoprotein p62 |
| NUP153 | Nuclear pore complex protein Nup153 |
| RCC1 | Regulator of chromosome condensation |
| RAN | GTP-binding nuclear protein Ran |
| RANBP1 | Ran-specific GTPase-activating protein |
| RANBP2 | E3 SUMO-protein ligase RanBP2 |
Result
Structurally similar non-target protein families, GLR, GABA, nAChR, 5-HT receptors, and, P2XR as well as five proteins: ZACN, IPO5, RANBP6, TNPO1 and, TNPO2 were analyzed. Using databases, for GABRA1- 21 non-target proteins, CHRNA7- 24 non-target proteins, P2RX4-6 non-target, GLRA3-21 non-target and, KPNB1/IMB1-4 non-target proteins were found (Table 1). Using STRING, protein interactions were generated with target proteins: α1β2γ2L GABA (A) receptors, GLRA-3 receptor, α7 nAChR, P2X4 cation channel, and IMPα/β1 that may cause disturbances in the functioning of these genes indirectly (Table 2). For GABRA1, GABRB2, and GABRG2, the proteins generated by STRING were; GABRA4, GABRA5, GABRA6, GABRB3, GPHN, DNM1, GABRB1, and UBQLN1. In case of CHRNA7, STRING produced 10 proteins; CHRNA2, CHRNA3, CHRNA4, CHRNA9, CHRNB2, CHRNB4, APP, MTMR10, LYNX1 and, SLURP1 and eleven proteins were generated for P2RX4; P2RX1, P2RX7, P2RX6, P2RY1, P2RY2, P2RY4, P2RY6, P2RY12, PANX1, GRIN2A and, GRIN1. For GLRA3, the outputs of ten proteins; GLRB, ANO1, ANO6, GABRB3, BEST1, GARS, GPHIN, NBEA, CABLES1, and NUDFAF6 were generated which interact with each other. For KPNB1/ IMPB1 and KPNA1/ IMPA1, STRING generated nine interacting proteins; KPNA2, SNUPN, NUP98, NUP62, NUP153, RCC1, RAN, RANBP1, and RANBP2 (Figure. 3).
Discussion
During COVID-19 pandemic, a 24-fold spike in ivermectin retail was observed in the United Statesin in August 2021. A recent study reported, potentially severe side effects such as disorientation, ataxia, seizures, visual impairment, hypotension, and dizziness in patients receiving ivermectin to treat COVID 1914, 15. The work focuses on potential non-target proteins that ivermectin drug may bind owing to structural similarities with target proteins, as well as proteins interacting with these target proteins that may cause indirect abnormalities, potentially leading to severe adverse effects in patients.
GLR (Glycine receptor family): Glycine receptors are made up of five subunits that create a pentameric protein structure with four alphas (GLRA1, GLRA2, GLRA3, GLRA4), and one beta chain (GLRB). They belong to the ligand-gated ion channel protein family and serve as a primary inhibitory ion channel neurotransmitter in the central nervous system (CNS) of mammals16. When glycine binds to receptors, it activates the ion channel, causing an influx of chloride ions into the cytoplasm and hyperpolarization of the postsynaptic membrane. Ivermectin binding further increases the inflow of chloride ions, causing overexcitation, and disturbance in the cell, making them valuable targets for neuroactive medicines. GLRA2 gene codes for a protein that binds to alkaloid strychnine and serves as a competitive antagonist, causing excitotoxicity in the body, which causes pain, muscular cramping, and excessive startle reactions. According to studies on oscillator mutations, substantial loss of function alleles in GLRA1 may induce prenatal or neonatal death in humans17.
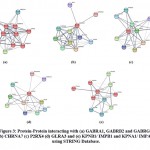 |
Figure 3: Protein-Protein interacting with (a) GABRA1, GABRB2 and GABRG2 (b) CHRNA7 (c) P2RX4 (d) GLRA3 and (e) KPNB1/ IMPB1 and KPNA1/ IMPA1 using STRING Database. |
GABA (Gamma-aminobutyric acid receptor family): GABA-A receptors are a pentameric ligand-gated chloride channels that include subunits from the alpha, beta, gamma, delta, and rho classes. Non-target proteins generated were GABRB3, GABRA6, GABRB1, GABRR2, GABRR1, GABRA4, GABRG1, GABRA3, GABRR3, GABRA5, GABRA2, GABRG3, GABRD, GABRQ and, GABRE. GABA is a key inhibitory neurotransmitter that acts as a ligand for receptors, allowing chloride ions to flow through the cell membrane. Neuronal damage is caused by GABA receptor excitotoxicity in a range of neurodegenerative disorders, like Parkinson’s disease, Alzheimer’s disease (AD), and Lou Gehrig’s disease (ALS)18. The dysfunction of these channels causes anxiety disorders, epilepsy, and neurodevelopmental illnesses including autism19. These are also involved in pathways like retrograde endocannabinoid signaling and taste transduction.
nAChRs(Nicotinic Acetylcholine Receptors): The non-target proteins obtained for all the target proteins are CHRNA1, CHRNA2, CHRNA3, CHRNA4, CHRNA5, CHRNA6, CHRNA9, CHRNA10, CHRNB1, CHRNB2, CHRNB3, CHRNB4, CHRND, CHRNE, and CHRNG. These receptors carry five subunits and act as ion channels in the nervous system. nAChR superfamily of ligand-gated ion channels contains a central pore that serves as a cation channel and is made up of pentameric structures with homomeric or heteromeric alpha and beta subunits. Nicotine is the agonist that binds to these receptors, and each subunit has four transmembrane domains and an external amino terminus. Mutations in the CHRNB3/A6 genes have been linked to nicotine and alcohol addiction, whereas CHRNA3, CHRNA5 has been linked to lung cancer 20, 21.
5-HT Receptors (5-hydroxytryptamine receptors): The 5-HT family includes HTR3B, HTR3A, HTR3C, and HTR3E, which encode for ligand-gated ion channel proteins that operate act as a transmitter, regulator, and mitogen. 5-HT3 receptors permit fast, depolarizing responses in neurons by going through a sequence of conformational changes that are triggered by ligand binding and result in the rapid opening of an intrinsic cation-selective channel. They are cation-specific ion channels that are otherwise nonselective and mainly expressed in the cerebral cortex, amygdala, hippocampus, and testis. HTR3A is involved in colon carcinogenesis and its malfunction has neurological consequences, whereas HTR3B is linked to serious major depression in women21, 22. These receptors are involved in pathways like serotonergic synapse and taste transduction.
ZACN (zinc activated ion channel):In mammals, the “Cysteine Loop” superfamily, LGICs is composed about 40 homologous subunits divided into four families that represent acetylcholine, serotonin, gamma-aminobutyric acid, or glycine-controlled channels. The ligand-gated ion channel ZACN belongs to a novel subgroup of the cysteine-loop superfamily. ZAC subunit transcripts have been discovered from different regions in human body like stomach, spinal cord, trachea, and also from foetal brain. Human and dog genomes have genes from this subgroup, but in case of mouse and rat genomes, they appear to have been lost 23.
P2XR:Sensory afferent neurons express P2XR receptors, a class of ionotropic purinergic receptors which are involved in sensory transduction. The non-target proteins found were P2RX1, P2RX2, P2RX3, P2RX5, P2RX6 and, P2RX7. When P2X receptors bind ATP, their cation-permeable ligand-gated ion channels open. P2X receptors are found throughout the nervous system because they are involved in synaptic transmission (e.g., on glial cells and nerve terminals). P2X7R receptors affect the release of growth factors and hormones and also modulates the immune response like inflammation. In a variety of clinical conditions, P2X7R activation or upregulation by favourably regulating vascular endothelial growth factor (VEGF) production and accumulation, promotes neo-angiogenesis24. Spiral ganglion neurons (SGNs) and hair cells express P2RX320 throughout cochlear development25.
IPO5, RANBP6, TNPO1 and, TNPO2:Nucleocytoplasmic transport is carried out by nuclear pore complexes present in the nuclear membranes. Nuclear localization signal (NLS) import receptor, also known as karyopherins, an alpha and beta subunits importin heterodimer is required for the proteins transportation. The importin beta family of proteins encoded by IPO5, TNPO1, TNPO2, and RANBP6 genes, guide nuclear proteins by interacting with nuclear localization signals. Mutations in the Transportin-dependent NLS of FUS (fused in sarcoma) mislocalizes this protein and result in amyotrophic lateral sclerosis, demonstrating the Transportin-dependent import significance in human health24. It also helps in YB-1 (Y-box binding protein 1) nuclear translocation and mediated the release of the HIV-1 genome from the capsid protein shell and the transport of the viral nucleus into the nucleus of the host cell27,28. IPO5 gene’s aberrant expression and alternative splicing may have a role in the pathophysiology of schizophrenia29.
Protein-Protein Interaction
Protein interactions with target proteins (α1β2γ2L GABA (A) receptors, GLRA-3 receptor, 7 nAChR, P2X4 cation channel, and IMPα/β1) were investigated in order to gain insight and understanding into how they may induce indirect disruptions in the function of these genes. These interactions between non-target and target proteins also aid in determining the molecular basis of disruptions and the cell’s response to them. For GABRA1, GABRB2, and GABRG2 most of the proteins are of the GABA family and play an important function in the neurotransmission of chloride ions through the cell membrane. DNM1 is a microtubule bundle producer and is associated with developmental and epileptic encephalopathy and Lennox-Gastaut syndrome-like diseases30. UBQLN1 regulates the protein degradation processes and routes. In case of CHRNA7, out of the ten interacting proteins, seven proteins are nicotinic acetylcholine receptors (nAChRs) and plays role in ethanol and nicotine addiction, whereas LYNX1 functions as a modulator of these receptors31. Autosomal dominant Alzheimer’s disease and cerebro-arterial amyloidosis have both been linked to mutations in the APP gene. APP gene mutations can result into Autosomal dominant Alzheimer disease and cerebro-arterial amyloidosis, on the other hand, SLURP1 mutations leads to a rare autosomal recessive skin condition, Mal de Meleda32. Majority of the eleven proteins generated by STRING for P2RX4 are P2 receptors, which are overexpressed in various cancer cell types i.e., P2Y2R activation by ATP causes tumor invasion and angiogenesis33. The glutamate-gated ion channel proteins, GRIN2A and GRIN1 are members of the glutamate-gated ion channel protein family, and mutations in these genes induce cognitive impairment34. For GLRA3, the outputs of ten proteins generated are mainly involved in ion transmissions in the nervous system of humans. Disturbances in BEST1 protein cause retinal degenerative diseases, whereas ANO1 and ANO6 function as a calcium-activated chloride channel and have been implicated in hypertension35. GARS catalyze the ligation of glycine to the 3′-end of its cognate tRNA and NBEA disturbance generates neurodevelopmental disease (NDD)36. CABLES1 is critical for neuronal development; GABRB3 and GLRB are ligand-gated chloride channels and play a key role in the down-regulation of neuronal excitability. KPNB1/ IMPB1 and KPNA1/ IMPA1, STRING generated nine interacting proteins; seven proteins function as nucleocytoplasmic transporters. As adapter proteins for KPNB1 receptor, KPNA1 and KPNA2 proteins play a vital role in nuclear protein import, and their dysregulation causes a variety of diseases, including cancers37. NUP98 and NUP96 proteins, on the other hand, are involved in bidirectional trafficking through the nuclear pore complex (NPC) and have been linked to leukemia38. RAN participates in both the import and export of proteins and RNA along with the nucleus. When the ivermectin drug binds to these non-target proteins, it disrupts the central nervous system (CNS) signal transmission and neuroactive ligand-receptor interactions in the human biological system. Glycine signaling disruption in the hippocampus has been linked to neuropsychiatric diseases, whereas changes in gamma-aminobutyric acid (GABA) levels leads to dis-balance between excitatory and inhibitory signals, and are implicated in the development of a variety of neuropsychiatric disorders39,40. Gene expression changes in CHRNA6, HTR3A, and HTR3B have also been linked to motor impairments, adult psychopathology, and major depression in humans41,42,43. TNPO1 is involved in nucleocytoplasmic transport and may be significant in the progression of atherosclerotic coronary artery disease (CAD), TNPO2 and IPO5 has been reported to induce gastric cancer cells growth, oesophageal cancer and cough reflex sensitization, which can further lead to chronic cough,44,45,46,47. Protein-protein interactions studies also revealed that the majority of the proteins interacting were members of the “Cys Loop” superfamily and LGICs, primarily involved in rapid synaptic neurotransmission in the brain, and disruption in target proteins due to ivermectin overdose may cause indirect disturbances in the functioning of these proteins, eventually leading to neural disorders and the development of gastric, oesophageal, or lung cancer.
Conclusion
The findings of this study provided insight on the significance of extensive investigation of the various non-target proteins of all the approved repurposing drugs’ against any chronic and complex diseases to fully comprehend their off-target effects before public use. The current investigation found that while ivermectin, a repurposing drug can treat COVID-19 patients, it can also accidentally disrupt important biological processes by binding with non-target proteins or may lead to indirect disturbances in the proteins interacting with these target proteins. This can result in severe off-target effects such as impaired vision, hypotension, visual hallucinations, loss of coordination and balance, depression, seizures and neurodegenerative disorders, like Parkinson’s disease, Alzheimer’s disease in patients and can eventually also lead to cancer in the consumer’s body.
Acknowledgment
This work was supported by DBT STAR College grant and Gargi College, University of Delhi.
Conflict of Interest
The authors have no conflict of interest to declare.
Funding Sources
There is no funding sources
References
- https://covid19.who.int/
- Singh TU, Parida S, Lingaraju MC, Kesavan M, Kumar D, Singh RK. Drug repurposing approach to fight COVID-19. Pharmacol Rep. 2020;72(6):1479-1508.
CrossRef - https://go.drugbank.com/drugs/DB00602
- Huang X, Chen H, Shaffer PL. Crystal Structures of Human GlyRα3 Bound to Ivermectin. Structure. 2017;25(6):945-950.e2.
CrossRef - Estrada-Mondragon A, Lynch JW. Functional characterization of ivermectin binding sites in α1β2γ2L GABA(A) receptors. Front Mol Neurosci. 2015; 8:55. Published 2015 Sep 25.
CrossRef - Krause RM, Buisson B, Bertrand S, et al. Ivermectin: a positive allosteric effector of the alpha7 neuronal nicotinic acetylcholine receptor. Mol Pharmacol. 1998;53(2):283-294.
CrossRef - Priel A, Silberberg SD. Mechanism of ivermectin facilitation of human P2X4 receptor channels. J Gen Physiol. 2004;123(3):281-293.
CrossRef - Caly L, Druce JD, Catton MG, Jans DA, Wagstaff KM. The FDA-approved drug ivermectin inhibits the replication of SARS-CoV-2 in vitro. Antiviral Res. 2020; 178:104787.
CrossRef - Wishart DS, Knox C, Guo AC, Shrivastava S, Hassanali M, Stothard P, Chang Z, Woolsey J. DrugBank: a comprehensive resource for in silico drug discovery and exploration. Nucleic Acids Res. 2006 Jan 1;34(Database issue):D668-72. doi: 10.1093/nar/gkj067. PMID: 16381955; PMCID: PMC1347430.
CrossRef - Kent WJ, Hsu F, Karolchik D, Kuhn RM, Clawson H, Trumbower H, Haussler D. Exploring relationships and mining data with the UCSC Gene Sorter. Genome Res. 2005 May;15(5):737-41. doi: 10.1101/gr.3694705. PMID: 15867434; PMCID: PMC1088302.
CrossRef - Warde-Farley D, Donaldson SL, Comes O, Zuberi K, Badrawi R, Chao P, Franz M, Grouios C, Kazi F, Lopes CT, Maitland A, Mostafavi S, Montojo J, Shao Q, Wright G, Bader GD, Morris Q. The GeneMANIA prediction server: biological network integration for gene prioritization and predicting gene function. Nucleic Acids Res. 2010 Jul;38(Web Server issue):W214-20. doi: 10.1093/nar/gkq537. PMID: 20576703; PMCID: PMC2896186.
CrossRef - Szklarczyk D, Gable AL, Nastou KC, Lyon D, Kirsch R, Pyysalo S, Doncheva NT, Legeay M, Fang T, Bork P, Jensen LJ, von Mering C. The STRING database in 2021: customizable protein-protein networks, and functional characterization of user-uploaded gene/measurement sets. Nucleic Acids Res. 2021 Jan 8;49(D1): D605-D612. doi: 10.1093/nar/gkaa1074. Erratum in: Nucleic Acids Res. 2021 Oct 11;49(18):10800. PMID: 33237311; PMCID: PMC7779004.
CrossRef - Kanehisa M, Furumichi M, Tanabe M, Sato Y, Morishima K. KEGG: new perspectives on genomes, pathways, diseases and drugs. Nucleic Acids Res. 2017 Jan 4;45(D1):D353-D361. doi: 10.1093/nar/gkw1092. Epub 2016 Nov 28. PMID: 27899662; PMCID: PMC52Betz H, Laube B. Glycine receptors: recent insights into their structural organization and functional diversity. J Neurochem. 2006;97(6):1600-1610.
CrossRef - Centers for Disease Control and Prevention. “Rapid increase in ivermectin prescriptions and reports of severe illness associated with use of products containing ivermectin to prevent or treat COVID-19.” CDC Health Alert Network no. CDCHAN-004498 (2021): 26.
CrossRef - Lind JN, Lovegrove MC, Geller AI, Uyeki TM, Datta SD, Budnitz DS. Increase in outpatient ivermectin dispensing in the US during the COVID-19 pandemic: a cross-sectional analysis. J Gen Intern Med2021;36:2909-2911.
CrossRef - Buckwalter MS, Cook SA, Davisson MT, White WF, Camper SA. A frameshift mutation in the mouse alpha 1 glycine receptor gene (Glra1) results in progressive neurological symptoms and juvenile death. Hum Mol Genet. 1994;3(11):2025-2030.
CrossRef - Louzada PR, Paula Lima AC, Mendonca-Silva DL, Noël F, De Mello FG, Ferreira ST. Taurine prevents the neurotoxicity of beta-amyloid and glutamate receptor agonists: activation of GABA receptors and possible implications for Alzheimer’s disease and other neurological disorders. FASEB J. 2004;18(3):511-518.
CrossRef - Zhu S, Noviello CM, Teng J, Walsh RM Jr, Kim JJ, Hibbs RE. Structure of a human synaptic GABAANature. 2018;559(7712):67-72. doi:10.1038/s41586-018-0255-3
CrossRef - Hoft NR, Corley RP, McQueen MB, Huizinga D, Menard S, Ehringer MA. SNPs in CHRNA6 and CHRNB3 are associated with alcohol consumption in a nationally representative sample. Genes Brain Behav. 2009;8(6):631-637.
CrossRef - Carcereny E, Ramirez JL, Sanchez-Ronco M, Isla D, Cobo M, Moran T, de Aguirre I, Okamoto T, Wei J, Provencio M, Lopez-Vivanco G, Camps C, Domine M, Alberola V, Sanchez JM, Massuti B, Mendez P, Taron M, Rosell R. Blood-based CHRNA3 single nucleotide polymorphism and outcome in advanced non-small-cell lung cancer patients. Lung Cancer. 2010;68(3):491-7.
CrossRef - Tang J, Wang Z, Liu J, Zhou C, Chen J. Downregulation of 5-hydroxytryptamine receptor 3A expression exerts an anticancer activity against cell growth in colorectal carcinoma cells in vitro. Oncol Lett. 2018;16(5):6100-6108.
CrossRef - Yamada K, Hattori E, Iwayama Y, et al. Distinguishable haplotype blocks in the HTR3A and HTR3B region in the Japanese reveal evidence of association of HTR3B with female major depression. Biol Psychiatry. 2006;60(2):192-201.
CrossRef - Davies PA, Wang W, Hales TG, Kirkness EF. A novel class of ligand-gated ion channel is activated by Zn2+. J Biol Chem. 2003;278(2):712-717.
CrossRef - Tassetto M, Scialdone A, Solini A, Di Virgilio F. The P2X7 Receptor: A Promising Pharmacological Target in Diabetic Retinopathy. Int J Mol Sci. 2021;22(13):7110. Published 2021 Jul 1.
CrossRef - Wang Z, Jung JS, Inbar TC, Rangoussis KM, Faaborg-Andersen C, Coate TM. The Purinergic Receptor P2rx3 is Required for Spiral Ganglion Neuron Branch Refinement during Development. eNeuro. 2020;7(4): ENEURO.0179-20.2020. Published 2020 Aug 10.
CrossRef - Twyffels L, Gueydan C, Kruys V. Transportin-1 and Transportin-2: protein nuclear import and beyond. FEBS Lett. 2014;588(10):1857-68. doi: 10.1016/j.febslet.2014.04.023. Epub 2014 Apr 26. PMID: 24780099
CrossRef - Mordovkina DA, Kim ER, Buldakov IA, et al. Transportin-1-dependent YB-1 nuclear import. Biochem Biophys Res Commun. 2016;480(4):629-634.
CrossRef - Fernandez J, Machado AK, Lyonnais S, et al. Transportin-1 binds to the HIV-1 capsid via a nuclear localization signal and triggers uncoating. Nat Microbiol. 2019;4(11):1840-1850.
CrossRef - Wang ZQ, Liu Y, Wu N, et al. Genetic and functional study of the IPO5 gene in schizophrenia. Psychiatry Res. 2011;187(3):460-461.
CrossRef - https://www.genecards.org/cgi-bin/carddisp.pl?gene=DNM1
- Feduccia AA, Chatterjee S, Bartlett SE. Neuronal nicotinic acetylcholine receptors: neuroplastic changes underlying alcohol and nicotine addictions. Front Mol Neurosci. 2012; 5:83. Published 2012 Aug 3.
CrossRef - Bchetnia M, Laroussi N, Youssef M, et al. Particular Mal de Meleda phenotypes in Tunisia and mutations founder effect in the Mediterranean region. Biomed Res Int. 2013;2013:206803. doi:10.1155/2013/206803
CrossRef - Jin H, Kim HJ. NLRC4, ASC and Caspase-1 Are Inflammasome Components that Are Mediated by P2Y2R Activation in Breast Cancer Cells. Int J Mol Sci. 2020;21(9):3337. Published 2020 May 8.
CrossRef - https://www.genecards.org/cgi-bin/carddisp.pl?gene=GRIN2A
- Wang B, Li C, Huai R, Qu Z. Overexpression of ANO1/TMEM16A, an arterial Ca2+-activated Cl- channel, contributes to spontaneous hypertension. J Mol Cell Cardiol. 2015; 82:22-32.
CrossRef - Mulhern MS, Stumpel C, Stong N, et al. NBEA: Developmental disease gene with early generalized epilepsy phenotypes. Ann Neurol. 2018 Nov;84(5):788-795.
CrossRef - Yasuhara N, Kumar PK. Aptamers that bind specifically to human KPNA2 (importin-α1) and efficiently interfere with nuclear transport. J Biochem. 2016;160(5):259-268.
CrossRef - Terlecki-Zaniewicz S, Humer T, Eder T, et al. Biomolecular condensation of NUP98 fusion proteins drives leukemogenic gene expression. Nat Struct Mol Biol. 2021;28(2):190-201.
CrossRef - Xu TL, Gong N. Glycine and glycine receptor signaling in hippocampal neurons: diversity, function and regulation. Prog Neurobiol. 2010;91(4):349-361.
CrossRef - Jembrek MJ, Vlainic J. GABA Receptors: Pharmacological Potential and Pitfalls. Curr Pharm Des. 2015;21(34):4943-4959.
CrossRef - Glavan D, Gheorman V, Gresita A, Hermann DM, Udristoiu I, Popa-Wagner A. Identification of transcriptome alterations in the prefrontal cortex, hippocampus, amygdala and hippocampus of suicide victims. Sci Rep. 2021;11(1):18853.
CrossRef - Schechter DS, Moser DA, Aue T, et al. Maternal PTSD and corresponding neural activity mediate effects of child exposure to violence on child PTSD symptoms. PLoS One. 2017;12(8):e0181066.
CrossRef - Krzywkowski K, Davies PA, Feinberg-Zadek PL, Bräuner-Osborne H, Jensen AA. High-frequency HTR3B variant associated with major depression dramatically augments the signaling of the human 5-HT3AB receptor. Proc Natl Acad Sci U S A. 2008;105(2):722-727.
CrossRef - Zhang X, Sun R, Liu L. Potentially critical roles of TNPO1, RAP1B, ZDHHC17, and PPM1B in the progression of coronary atherosclerosis through microarray data analysis. J Cell Biochem. 2019;120(3):4301-4311.
CrossRef - Gong L, Wen T, Li Z, et al. TNPO2 operates downstream of DYNC1I1 and promotes gastric cancer cell proliferation and inhibits apoptosis. Cancer Med. 2019;8(17):7299-7312.
CrossRef - Li XF, Aierken AL, Shen L. IPO5 promotes malignant progression of esophageal cancer through activating MMP7. Eur Rev Med Pharmacol Sci. 2020 Apr;24(8):4246-4254.
- Abdulqawi R, Dockry R, Holt K, et al. P2X3 receptor antagonist (AF-219) in refractory chronic cough: a randomised, double-blind, placebo-controlled phase 2 study. Lancet. 2015;385(9974):1198-1205.
CrossRef

This work is licensed under a Creative Commons Attribution 4.0 International License.