Abstract
Background
Promoter methylation of colorectal cancer-related genes were examined with respect to phenotype and tumor progression.
Materials and Methods
We assayed promoter methylation of 11 genes including established CpG island methylator phenotype (CIMP) markers (MLH1, MINT1, MINT2, MINT31, p16 INK4a, p14 ARF, and CACNA1G) and four genes (COX2, DAPK, MGMT, and APC) frequently methylated in colorectal cancer in 285 patients with sporadic colorectal cancer.
Results
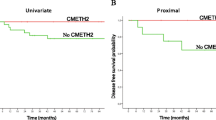
CIMP+ tumors were more than two times more frequent among high-frequency microsatellite instability tumors (MSI-H) than in tumors without MSI (P ≤ .0001–.002). COX2 and DAPK methylation were significantly associated with CIMP+ and MSI. KRAS showed tendency toward more frequent codon 12-13 mutations identified in tumors with APC and p16 INK4a methylation than in those with unmethylation (P = .033 and .05, respectively). Additionally, tumors with synchronous adenoma were associated with p16 INK4a methylation (P = .004). The p16 INK4a methylation was significantly associated with poor overall and disease-free survival in 131 rectal cancer patients who underwent curative operation, according to multivariate analyses (relative risk [RR] = 0.317 and 0.349; P = .033 and .024, respectively). Specifically, in 175 stage II and III patients receiving adjuvant-based fluoropyrimidine chemotherapy, p16 INK4a methylation and MINT31 unmethylation showed a significant or tendency toward an association with recurrence and DFS (P = .007–.032).
Conclusions
The study suggests that specific CIMP markers, such as p16 INK4a and MINT31, should be further verified as potential epigenetic targets for the design of efficient chemotherapy regimens. We also identified a subset of colorectal cancer, possibly comprising APC methylation-KRAS mutation-p16 INK4a methylation.



Similar content being viewed by others
References
Jones P, Laird P. Cancer epigenetics comes of age. Nat Genet. 1999;21:163–7.
Zitt M, Zitt M, Müller HM. DNA methylation in colorectal cancer—impact on screening and therapy monitoring modalities? Dis Markers. 2007;23:51–71.
Chan AO, Broaddus RR, Houlihan PS, Issa JP, Hamilton SR, Rashid A. CpG island methylation in aberrant crypt foci of the colorectum. Am J Pathol. 2002;160:1823–30.
Iacopetta B, Grieu F, Li W, Ruszkiewicz A, Caruso M, Moore J, et al. APC gene methylation is inversely correlated with features of the CpG island methylator phenotype in colorectal cancer. Int J Cancer. 2006;119:2272–8.
Sweeney C, Boucher KM, Samowitz WS, Wolff RK, Albertsen H, Curtin K, et al. Oncogenetic tree model of somatic mutations and DNA methylation in colon tumors. Genes Chromosomes Cancer. 2009;48:1–9.
Grady WM, Carethers JM. Genomic and epigenetic instability in colorectal cancer pathogenesis. Gastroenterology. 2008;135:1079–99.
Yamashita K, Dai T, Dai Y, Yamamoto F, Perucho M. Genetics supersedes epigenetics in colon cancer phenotype. Cancer Cell. 2003;4:121–31.
Goel A, Nagasaka T, Arnold CN, Inoue T, Hamilton C, Niedzwiecki D, et al. The CpG island methylator phenotype and chromosomal instability are inversely correlated in sporadic colorectal cancer. Gastroenterology. 2007;132:127–38.
Nosho K, Irahara N, Shima K, Kure S, Kirkner GJ, Schernhammer ES, et al. Comprehensive biostatistical analysis of CpG island methylator phenotype in colorectal cancer using a large population-based sample. PLoS One. 2008;3:e3698.
Ogino S, Meyerhardt JA, Kawasaki T, Clark JW, Ryan DP, Kulke MH, et al. CpG island methylation, response to combination chemotherapy, and patient survival in advanced microsatellite stable colorectal carcinoma. Virchows Arch. 2007;450:529–37.
Nagasaka T, Sasamoto H, Notohara K, Cullings HM, Takeda M, Kimura K, et al. Colorectal cancer with mutation in BRAF, KRAS, and wild-type with respect to both oncogenes showing different patterns of DNA methylation. J Clin Oncol. 2004;22:4584–94.
Maier S, Dahlstroem C, Haefliger C, Plum A, Piepenbrock C. Identifying DNA methylation biomarkers of cancer drug response. Am J Pharmacogenomics. 2005;5:223–32.
Arnold CN, Goel A, Boland CR. Role of hMLH1 promoter hypermethylation in drug resistance to 5-fluorouracil in colorectal cancer cell lines. Int J Cancer. 2003;106:66–73.
Iacopetta B, Kawakami K, Watanabe T. Predicting clinical outcome of 5-fluorouracil-based chemotherapy for colon cancer patients: is the CpG island methylator phenotype the 5-fluorouracil-responsive subgroup? Int J Clin Oncol. 2008;13:498–503.
Tost J, Gut IG. DNA methylation analysis by pyrosequencing. Nat Protoc. 2007;2:2265–75.
Nosho K, Kure S, Irahara N, Shima K, Baba Y, Spiegelman D, et al. A prospective cohort study shows unique epigenetic, genetic, and prognostic features of synchronous colorectal cancers. Gastroenterology. 2009;137:1609–20.
Toyota M, Shen L, Ohe-Toyota M, Hamilton SR, Sinicrope FA, Issa JP. Aberrant methylation of the Cyclooxygenase 2 CpG island in colorectal tumors. Cancer Res. 2000;60:4044–8.
Mittag F, Kuester D, Vieth M, Peters B, Stolte B, Roessner A, Schneider-Stock R. DAPK promotor methylation is an early event in colorectal carcinogenesis. Cancer Lett. 2006;240:69–75.
Weisenberger DJ, Siegmund KD, Campan M, Young J, Long TI, Faasse MA, et al. CpG island methylator phenotype underlies sporadic microsatellite instability and is tightly associated with BRAF mutation in colorectal cancer. Nat Genet. 2006;38:787–93.
Ogino S, Nosho K, Kirkner GJ, Kawasaki T, Meyerhardt JA, Loda M, et al. CpG island methylator phenotype, microsatellite instability, BRAF mutation and clinical outcome in colon cancer. Gut. 2009;58:90–6.
Barault L, Charon-Barra C, Jooste V, de la Vega MF, Martin L, Roignot P, et al. Hypermethylator phenotype in sporadic colon cancer: study on a population-based series of 582 cases. Cancer Res. 2008;68:8541–6.
Lynch HT, de la Chapelle A. Genetic susceptibility to non-polyposis colorectal cancer. J Med Genet. 1999;36:801–18.
Suehiro Y, Wong CW, Chirieac LR, Kondo Y, Shen L, Webb CR, et al. Epigenetic-genetic interactions in the APC/WNT, RAS/RAF, and P53 pathways in colorectal carcinoma. Clin Cancer Res. 2008;14:2560–9.
Lind GE, Thorstensen L, Løvig T, Meling GI, Hamelin R, Rognum TO, et al. A CpG island hypermethylation profile of primary colorectal carcinomas and colon cancer cell lines. Mol Cancer. 2004;3:28.
Judson H, Stewart A, Leslie A, Pratt NR, Baty DU, Steele RJ, et al. Relationship between point gene mutation, chromosomal abnormality, and tumour suppressor gene methylation status in colorectal adenomas. J Pathol. 2006;210:344–50.
Ogino S, Kawasaki T, Kirkner GJ, Loda M, Fuchs CS. CpG island methylator phenotype-low (CIMP-low) in colorectal cancer: possible associations with male sex and KRAS mutations. J Mol Diagn. 2006;8:582–8.
Shen L, Toyota M, Kondo Y, Lin E, Zhang L, Guo Y, et al. Integrated genetic and epigenetic analysis identifies three different subclasses of colon cancer. Proc Natl Acad Sci USA. 2007;104:18654–9.
Kawakami K, Ruszkiewicz A, Bennett G, Moore J, Grieu F, Watanabe G, et al. DNA hypermethylation in the normal colonic mucosa of patients with colorectal cancer. Br J Cancer. 2006;94:593–8.
Wettergren Y, Odin E, Nilsson S, Carlsson G, Gustavsson B. p16INK4a gene promoter hypermethylation in mucosa as a prognostic factor for patients with colorectal cancer. Mol Med. 2008;14:412–21.
Issa JP. Aging, DNA methylation and cancer. Crit Rev Oncol Hematol. 1999;32:31–43.
Chirieac LR, Shen L, Catalano PJ, Issa JP, Hamilton SR. Phenotype of microsatellite-stable colorectal carcinomas with CpG island methylation. Am J Surg Pathol. 2005;29:429–36.
Koopman M, Kortman GA, Mekenkamp L, Ligtenberg MJ, Hoogerbrugge N, Antonini NF, et al. Deficient mismatch repair system in patients with sporadic advanced colorectal cancer. Br J Cancer. 2009;100:266–73.
Alhaja E, Adan J, Pagan R, Mitjans F, Cascallo M, Rodriguez M, et al. Anti-migratory and anti-angiogenic effect of p16: a novel localization at membrane ruffles and lamellipodia in endothelial cells. Angiogenesis. 2004;7:323–33.
Trzeciak L, Hennig E, Kolodziejski J, Nowacki M, Ostrowski J. Mutations, methylation and expression of CDKN2a/p16 gene in colorectal cancer and normal colonic mucosa. Cancer Lett. 2001;163:17–23.
Nakayama G, Hibi K, Kodera Y, Koike M, Fujiwara M, Nakao A. p16 methylation in serum as a potential marker for the malignancy of colorectal carcinoma. Anticancer Res. 2007;27:3367–70.
Wang CH, Chang HC, Hung WC. p16 inhibits matrix metalloproteinase-2 expression via suppression of Sp1-mediated gene transcription. J Cell Physiol. 2006;208:246–52.
Morita S, Iida S, Kato K, Takagi Y, Uetake H, Sugihara K. The synergistic effect of 5-aza-2′-deoxycytidine and 5-fluorouracil on drug-resistant tumors. Oncology. 2006;71:437–45.
Nagasaka T, Sharp GB, Notohara K, Kambara T, Sasamoto H, Isozaki H, et al. Hypermethylation of O6-methylguanine-DNA methyltransferase promoter may predict nonrecurrence after chemotherapy in colorectal cancer cases. Clin Cancer Res. 2003;9:5306–12.
Skorpen F, Krokan HE. The methylation status of the gene for O6-methylguanine-DNA methyltransferase in human Mer+ and Mer- cells. Carcinogenesis. 1995;16:1857–63.
Hawkins NJ, Lee JH, Wong JJ, Kwok CT, Ward RL, Hitchins MP. MGMT methylation is associated primarily with the germline C>T SNP (rs16906252) in colorectal cancer and normal colonic mucosa. Mod Pathol. 2009;22:1588–99.
Ogino S, Kawasaki T, Kirkner GJ, Suemoto Y, Meyerhardt JA, Fuchs CS. Molecular correlates with MGMT promoter methylation and silencing support CpG island methylator phenotype-low (CIMP-low) in colorectal cancer. Gut. 2007;56:1564–71.
Acknowledgment
This work was supported by the Korea Health 21 R&D Project, Ministry of Health, Welfare, and Family Affairs (grant A062254).
Author information
Authors and Affiliations
Corresponding author
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Kim, J.C., Choi, J.S., Roh, S.A. et al. Promoter Methylation of Specific Genes is Associated with the Phenotype and Progression of Colorectal Adenocarcinomas. Ann Surg Oncol 17, 1767–1776 (2010). https://doi.org/10.1245/s10434-009-0901-y
Received:
Published:
Issue Date:
DOI: https://doi.org/10.1245/s10434-009-0901-y