Abstract
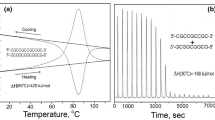
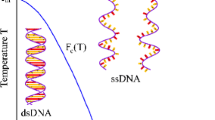
Loops are essential secondary structure elements in folded DNA and RNA molecules and proliferate close to the melting transition. Using a theory for nucleic acid secondary structures that accounts for the logarithmic entropy —c ln m for a loop of length m, we study homopolymeric single-stranded nucleic acid chains under external force and varying temperature. In the thermodynamic limit of a long strand, the chain displays a phase transition between a low-temperature/low-force compact (folded) structure and a high-temperature/high-force molten (unfolded) structure. The influence of c on phase diagrams, critical exponents, melting, and force extension curves is derived analytically. For vanishing pulling force, only for the limited range of loop exponents 2 < c ≲ 2.479 a melting transition is possible; for c ≤ 2 the chain is always in the folded phase and for 2.479 ≲ c always in the unfolded phase. A force-induced melting transition with singular behavior is possible for all loop exponents c < 2.479 and can be observed experimentally by single-molecule force spectroscopy. These findings have implications for the hybridization or denaturation of double-stranded nucleic acids. The Poland-Scheraga model for nucleic acid duplex melting does not allow base pairing between nucleotides on the same strand in denatured regions of the double strand. If the sequence allows these intra-strand base pairs, we show that for a realistic loop exponent c ≈ 2.1 pronounced secondary structures appear inside the single strands. This leads to a lower melting temperature of the duplex than predicted by the Poland-Scheraga model. Further, these secondary structures renormalize the effective loop exponent \( \hat{{c}}\), which characterizes the weight of a denatured region of the double strand, and thus affect universal aspects of the duplex melting transition.
Similar content being viewed by others
References
R.F. Gesteland, T.R. Cech, J.F. Atkins (Editors), The RNA World, 2nd edn. (Cold Spring Harbor Laboratory Press, Woodbury, 2005).
S.B. Smith, Y.J. Cui, C. Bustamante, Science 271, 795 (1996).
J. Liphardt, S. Dumont, S.B. Smith, I. Tinoco jr., C. Bustamante, Science 296, 1832 (2002).
M. Rief, H. Clausen-Schaumann, H.E. Gaub, Nat. Struct. Biol. 6, 346 (1999).
B. Maier, D. Bensimon, V. Croquette, Proc. Natl. Acad. Sci. U.S.A. 97, 12002 (2000).
U. Bockelmann, B. Essevaz-Roulet, F. Heslot, Phys. Rev. Lett. 79, 4489 (1997).
A. Mossa, M. Manosas, N. Forns, J.M. Huguet, F. Ritort, J. Stat. Mech. 2009, P02060 (2009).
I. Tinoco, O.C. Uhlenbeck, M.D. Levine, Nature 230, 362 (1971).
I. Tinoco jr., C. Bustamante, J. Mol. Biol. 293, 271 (1999).
A.V. Finkelstein, O.V. Galzitskaya, Phys. Life Rev. 1, 23 (2004).
P.G. de Gennes, Biopolymers 6, 715 (1968).
M.S. Waterman, T.F. Smith, Math. Biosci. 42, 257 (1978).
T.R. Einert, P. Näger, H. Orland, R.R. Netz, Phys. Rev. Lett. 101, 048103 (2008).
I.L. Hofacker, W. Fontana, P.F. Stadler, L.S. Bonhoeffer, M. Tacker, P. Schuster, Mon. Chem. 125, 167 (1994).
J.S. McCaskill, Biopolymers 29, 1105 (1990).
T.R. Einert, D.B. Staple, H. Kreuzer, R.R. Netz, Biophys. J. 99, 578 (2010).
A. Montanari, M. Mézard, Phys. Rev. Lett. 86, 2178 (2001).
U. Gerland, R. Bundschuh, T. Hwa, Biophys. J. 81, 1324 (2001).
M. Müller, F. Krzakala, M. Mézard, Eur. Phys. J. E 9, 67 (2002).
A. Hanke, M.G. Ochoa, R. Metzler, Phys. Rev. Lett. 100, 018106 (2008).
S. Cocco, J.F. Marko, R. Monasson, C. R. Phys. 3, 569 (2002).
D.K. Lubensky, D.R. Nelson, Phys. Rev. E 65, 031917 (2002).
R. Bundschuh, U. Gerland, Phys. Rev. Lett. 95, 208104 (2005).
T.R. Einert, R.R. Netz, Biophys. J., 100 (11), in print (2010).
Z.J. Tan, S.J. Chen, Biophys. J. 90, 1175 (2006).
Y.S. Mamasakhlisov, S. Hayryan, V.F. Morozov, C.K. Hu, Phys. Rev. E 75, 061907 (2007).
M. Baiesi, E. Orlandini, A.L. Stella, Phys. Rev. Lett. 91, 198102 (2003).
H. Orland, A. Zee, Nucl. Phys. B 620, 456 (2002).
R. Blossey, E. Carlon, Phys. Rev. E 68, 061911 (2003).
Y. Kafri, D. Mukamel, L. Peliti, Phys. Rev. Lett. 85, 4988 (2000).
B. Alberts, Molecular Biology of the Cell (Garland Science, 2002).
J.C.M. Gebhardt, T. Bornschlögl, M. Rief, Proc. Natl. Acad. Sci. U.S.A. 107, 2013 (2010).
T. Xia, J. SantaLucia jr., M.E. Burkard, R. Kierzek, S.J. Schroeder, X. Jiao, C. Cox, D.H. Turner, Biochemistry 37, 14719 (1998).
R. Bundschuh, T. Hwa, Europhys. Lett. 59, 903 (2002).
P.G. de Gennes, Scaling Concepts in Polymer Physics (Cornell University Press, 1979).
B. Duplantier, Phys. Rev. Lett. 57, 941 (1986).
M. Müller, Phys. Rev. E 67, 021914 (2003).
A. Erdélyi, Higher Transcendental Functions, Vol. 1 (McGraw-Hill, 1953).
P. Flajolet, A. Odlyzko, SIAM Discret. Math. 3, 216 (1990).
M. Abramowitz, I.A. Stegun (Editors), Handbook of Mathematical Functions, 10th edn. (U.S. Department of Commerce, 2002).
J.F. Léger, G. Romano, A. Sarkar, J. Robert, L. Bourdieu, D. Chatenay, J.F. Marko, Phys. Rev. Lett. 83, 1066 (1999).
D. Poland, H.A. Scheraga, J. Chem. Phys. 45, 1456 (1966).
D. Poland, H.A. Scheraga, J. Chem. Phys. 45, 1464 (1966).
T. Garel, H. Orland, Biopolymers 75, 453 (2004).
J. SantaLucia jr., Proc. Natl. Acad. Sci. U.S.A. 95, 1460 (1998).
I.L. Hofacker, Nucleic Acids Res. 31, 3429 (2003).
N.R. Markham, M. Zuker, Nucleic Acids Res. 33, W577 (2005).
P. Schuster, Rep. Prog. Phys. 69, 1419 (2006).
Author information
Authors and Affiliations
Corresponding author
Electronic supplementary material
Rights and permissions
About this article
Cite this article
Einert, T.R., Orland, H. & Netz, R.R. Secondary structure formation of homopolymeric single-stranded nucleic acids including force and loop entropy: Implications for DNA hybridization. Eur. Phys. J. E 34, 55 (2011). https://doi.org/10.1140/epje/i2011-11055-2
Received:
Accepted:
Published:
DOI: https://doi.org/10.1140/epje/i2011-11055-2