Abstract
Microbiological screening of the target component of emericellipsin A of the Emericellopsis alkalina E101 strain has been carried out in various biotechnological systems at various pHs. The content of emericellipsin A was quantified under these conditions. It has been established that the new approved membrane-liquid cultivation method at pH 10 contributes to an increase in the yield of the main component of emericellipsin A. It has been shown that the new method of cultivating the strain E. alkalina E101 also promotes the synthesis of various isoforms of the main component of emericellipsin A. Some comparative analysis has been carried out.
Similar content being viewed by others
In recent years, antimicrobial peptides (AMPs) have attracted the attention of researchers as novel therapeutic agents with a number of advantages: high selectivity, low immunogenicity, a good possibility of penetrating into a target cell and a lower risk of resistance development due to the effect on the cell wall or membrane. In the past 2 decades, the total number of AMPs approved at the main pharmaceutical markets has increased by two times [1]. The major disadvantage of AMPs that prevents their application is cytotoxicity. At the same time, natural peptides are used both to develop pharmaceutical agents and as a model to create synthetic structural analogs on their basis [2]. At present, AMPs are recognized as a highly promising alternative class of new compounds for preventing antibiotic resistance [3, 4]. The yield of a natural compound can be increased due to extended screening, improved extraction techniques, and chemical synthesis, as well as synthesis in prokaryotic cells. These strategies open opportunities for obtaining structural analogs that can surpass the original natural compound in their pharmacodynamic or pharmacokinetic properties [5].
The discovery of peptaibols in micromycetes that inhabit cold and saline soils, sea depths, and other extreme environments expands opportunities for the search of new antibiotics, which can be prototypes of novel drugs. It is known that this group of nonribosomal peptides is synthesized solely by micromycetes. As a rule, a producer synthesizes a complex of several peptaibols, which are structurally homologous compounds that vary in their location in the peptide chain by one or several amino acids, which also determines the differences in their biological activity. Up to 11 isoforms of zervamicins were isolated from the cultures of different strains of the fungus Emericellopsis salmosynnemata, with zervamicin IIA (ZrvIIA) and zervamicin IIB (Zrv-IIB) being predominant among them [6]. Zrv-IIA and Zrv-IIB are structurally different from each other by only one amino-acid residue in the fourth position. In spite of their great structural homology, these isoforms were shown to be different in cytotoxicity, neurotoxicity, and antibacterial activity [7]. Albupeptins B and D are peptaibols that contain both stereoisomers of isovaline (Iva): the D- and L-configurations. They are active against Bacillus cinerea; the inhibitory effect depends on the structure and number of Iva residues (IC50: one Iva residue = 49.6 µg/mL; two Iva residues = 38.9 µg/mL; three Iva residues = 35.2 µg/mL; four Iva residues = 24.5 µg/mL). Albupeptin A is also inactive against Phytophthora infestans (IC50, >100 µg/mL), while compound D is active (IC50, four Iva residues = 16.3 µg/mL) [8]. Previously we isolated a complex of peptaibols with a marked antifungal activity from strains of the fungus Emericellopsis alkalina; the complex contains five isoforms of peptides with a single amino acid substitution: emercillipsins A–E. The dominant component of this complex, emericillipsin A (EmiA), had a considerable antifungal activity against the clinical isolates of pathogenic fungi with multiple drug resistance. The inhibitory activity of EmiA and its dehydroform against azole-resistant pathogenic isolates of Aspergillus spp. and Candida spp. manifests itself at the level of amphotericin B: 1 µg/mL, while for the clinical pathogenic isolates of Cryptococcus spp. it surpasses the reference agent by 2–4 times. At the same time, the activities of the isoforms B and C were significantly lower, while forms D and E proved to be inactive against pathogenic isolates of Aspergillus spp. and Candida spp. [9, 10]. Previously it was shown that the number of synthesized isoforms and the yield of EmiA varied between different strains during cultivation.
The structural diversity of synthesized peptaibols may also vary in the same producer strain, depending on cultivation techniques, the addition of precursors, and other physicochemical factors. Recently, X. Нао et al. [11] isolated new isoforms of acremopeptaibols from the strain Acremonium sp. IMB18-0 cultivated in the presence of bacterial biomass; the isoforms differed from those described in the literature in their antimicrobial activity and the absence of highly conserved threonine and hydroxyproline residues in the macromolecule. These isoforms demonstrated a marked antimicrobial activity against methicillin-resistant Staphylococcus aureus, Bacillus subtilis, and Candida albicans. The strain Trichoderma longibrachiatum Rifai DMG-3-1-1 synthesizes 23 peptaibols. The structures of 13 new peptaibols have been determined by NMR and MALDI-MS/MS. Thorough comparison of structures 1–23 has shown that only seven residues vary in the structures: 2 (Gln2/Asn2), 3 (Ile3/Val3), 4 (Ile4/Val4), 6 (Pro6/Hyp6), 8 (Leu8/Val8 ), 10 (Pro10/Hyp10) and 11 (Leuol11/Ileol11/Valol11). Peptaibols 2, 5, 9, 11, 21, and 22 exhibited moderate antibacterial activity against Staphylococcus aureus MRSA T144, as well as stronger cytotoxicity against BV2 and MCF-7 cells compared to other peptaibols of this strain. At the same time, it was shown that amino-acid residues 2, 3, and 4 had a strong effect on the cytotoxicity of the compound [12]. The discovery of novel unique AMP structures and the knowledge of the principles of dependence of activity on structure provide information for using chemical synthesis to create new compounds with higher antifungal activity and lower toxicity for the host compared to natural compounds [5].
This work was aimed at a comparative analysis of the diversity of produced isoforms of emericillipsins and assessment of the accumulation of the main component, emericillipsin A, during cultivation in different biotechnological systems and at different pH values.
MATERIALS AND METHODS
The accumulation and diversity of emericillipsin isoforms were assessed using the type strain of the mycelial fungus Emericellopsis alkalina Е101 (VKM F-4108; CBS 127350) from the collection Fungi from Extreme Environments, Department of Mycology and Algology, Faculty of Biology, Moscow State University (Russia). The strain was isolated from a soil sample taken on the coast of the soda lake Tanatar (Altai Krai, Russia) [13]. The screening of 64 strains of this species showed [14] that the strain E. alkalina Е101, together with the previously studied producer strain E. alkalina А118 (VKMР F-1428), was most productive with respect to the yield of the major component EmiА.
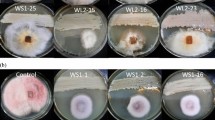
The producer strain E. alkalina Е101 was grown in a specialized liquid alkaline medium selected previously [15]. Media with different pH values were prepared using the following buffers: citrate-phosphate buffer for pH 7.00, phosphate-citrate buffer for pH 9.00, and carbonate-bicarbonate buffer for pH 10.00. Cultivation was carried out for 7, 14, and 21 days under stationary conditions in 750-mL Erlenmeyer flasks by the method of submerged culture in an Innova 40R shaker–incubator (Eppendorf New Brunswick, United States) and in a fermenter by the method of membrane-surface liquid culture on a bacterial cellulose matrix. Bacterial cellulose matrices were obtained by cultivating the strain Gluconacetobacter hansenii GH-1/2008 under stationary conditions at 27°С for 14 days in H-5 liquid medium. The resulting bacterial cellulose matrix was washed from producer cells with 0.1 n NaOH solution and distilled water, lyophilized, and sterilized. Stationary membrane-surface liquid cultivation in an alkaline medium was performed on a bacterial cellulose matrix as a substrate.
Culture liquid (CL) was separated by filtration through membrane filters in a Seitz filter funnel under vacuum. The CL was extracted 3 times with ethyl acetate or butanol at a ratio of 5 : 1. The resulting extracts were evaporated under vacuum in a Rotavapor-RBüchi (Switzerland) at 42°C to dryness and the remainder was dissolved in aqueous 50% ethanol to obtain alcohol concentrates.
The activity against conditionally pathogenic fungi was assessed in CL before and after the extraction, as well as in the extracts of mycelium. The antimicrobial activity was determined using sterile paper discs (Pasteur Institute of Epidemiology and Microbiology, Russia) moistened with antibiotic and dried under sterile conditions. The controls were standard discs with fluconazole for fungi (40 µg, Pasteur Institute of Epidemiology and Microbiology, Russia) and amoxicillin/clavulanic acid for bacteria (20/10 µg, Pasteur Institute of Epidemiology and Microbiology, Russia). The fungicidal activity was assessed using test strains: the mold fungus Aspergillus niger INA 00760 and the yeast Candida albicans АТСС 2091.
The antibacterial activity was assessed in relation to the Gram-negative bacterium Escherichia coli ATCC 25922 and the Gram-positive bacterium Bacillus subtilis АТСС 6633. The B. subtilis АТСС 6633 test culture was grown on a Gauze no. 2 medium of the following composition (g/L): tryptone, 2.5 (or Hottinger broth, 30 mL); peptone, 5; sodium chloride, 5; glucose, 10. E. coli ATCC 25922 was grown on the LB medium (tryptic soy agar). The fungal cultures of A. niger INA 00760 and C. albicans АТСС 2091 were grown on Czapek medium. The cultures were precultivated in agar slant tubes; the cells from the agar surface were suspended in the normal saline solution to a turbidity of 0.5 according to the McFarland standard (1.5 × 108 CFU/mL) and used within 15 min. The 24-h bacterial cultures and 5-day fungal cultures were used for inoculation. All test cultures were obtained from the culture collection of the Gauze Research Institute for New Antibiotics.
The resulting alcohol concentrates were combined, evaporated to dryness in a rotary evaporator (Labconco, United States), and then redissolved in 100 µL of 50% ethanol. The concentrates were analyzed by reversed-phase HPLC (RP HPLC). The analysis was carried out by Millichrome A-02 microcolumn chromatography (EcoNova Ltd., Novosibirsk) using Nucleosil-100-5-C18 stainless steel columns (L = 75.0 mm; D = 2.0 mm; d = 5 µm, Macherey-Nagel, Germany). Detection during the analysis was performed at 214 nm. The flow rate was 100 µL/min; the column thermostatting temperature was 35°С; the injection volume was 15 µL. The mobile phase composition was as follows: component A, H2O (MQ) + 0.02% TFA (HPLC, Sigma-Aldrich, Germany); component B, acetonitrile + 0.02% TFA. The mobile phase gradient was 0 to 100% B over 40 min, followed by an isocratic mode over 4 min at a 100% content of eluent C.
For the MALDI-TOF MS analysis, 0.3 µL of the acetonitrile–water fraction (collected during HPLC separation) of the sample and 0.5 µL of 2,5-dehydroxybenzoic acid (Sigma-Aldrich, Germany) solution in 20% acetonitrile + 79.5% water (МQ) + 0.5% TFA (HPLC, Sigma-Aldrich) at a concentration of 20 mg/mL were mixed on the spectrometer target. Spectrum recording and MS analysis were carried out with a MALDI-TOF MS spectrometer (Ultrafle Xtreme, Bruker Daltonics, Germany) with a UV laser (Nd) in the mode for recording positive ions using a reflectrone. The accuracy of mass determination was about 1 Da.
RESULTS AND DISCUSSION
For the strain E. alkalina Е101 [14], the formation of emericillipsins in the culture liquid and mycelium was originally studied at neutral and alkaline initial pH values (7.0, 9.0, and 10.0) under stationary and submerged cultivation conditions. It was shown that the EmiA content in mycelium always was much lower than in CL in the same variants. For mycelium, the higher content of EmiA was observed under submerged cultivation conditions, while the content of EmiA in CL was higher under stationary conditions in all variants under study, indicating that the antibiotic was excreted better to the culture liquid under these conditions.
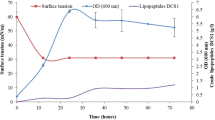
The content of EmiA in mycelium (0.25 mg/g) reached its maximum value on day 7 for submerged cultivation at pH 10.0 in the medium. The highest EmiA levels in CL were recorded on day 14 of growth at all initial pH values in the medium; however, the maximum EmiA content (6.0–6.5 mg/L) was observed at alkaline pH values (9.0 and 10.0) (Fig. 1). The experiment confirmed that EmiA is produced better under alkaline cultivation conditions, in spite of the fact that E. alkalina strains are able to grow and develop within a wide range of pH values in the medium (4.0–11.0) with the growth optimum at pH 10.0 [13].
Alkaliphilic fungi E. alkalina that inhabit the coasts of saline lakes often coexist with numerous and diverse prokaryotes forming biofilms at the phase separation interface. It is considered that the modeling of natural conditions for the producer can increase the accumulation of target antibiotics in the medium [16]. The yields of EmiA were compared in stationary and membrane-surface liquid cultures, with bacterial cellulose substrates as a model of a natural bacterial film. The maximum EmiA levels in CL and mycelium were observed in the stationary membrane-surface liquid culture (Table 1). The yield of EmiА in the strain E. alkalina Е101 on bacterial cellulose as a substrate increased by 1.7 times compared to the surface cultivation: 11.05 mg/L. In addition to the main target peptaibol EmiА, the synthesis of other, minor isoforms that had not been found previously under stationary conditions also occurred. At the same time, the extract from CL and mycelium obtained under membrane-surface liquid cultivation conditions exhibited activity against Gram-negative bacteria, while the extract obtained under stationary cultivation conditions was not active against these bacteria, indicating that the producer began to synthesize antibacterial compounds under these conditions in addition to the major antifungal component.
The diversity of various isoforms in the peptide complex synthesized in the membrane-surface liquid culture on a bacterial cellulose membrane was assessed later. After the analytical RP HPLC, the fractions of peaks with different retention times at elution with the acetonitrile concentration gradient from 70 to 100% were collected (Fig. 2). At the same time, close to the area of localization of EmiA peaks on the chromatogram, there were other peaks with longer and shorter retention times. Fractions from 1 to 17 (Fig. 2) were collected and analyzed by the MALD-TOF-MS technique. The resulting masses of molecular ions [M + H]+ of fraction components are presented in Table 2.
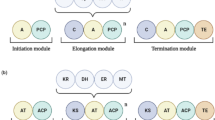
The fragmentation of individual peptaibols with molecular masses of 1022.8, 994.5, and 1036.7 suggests several possible (no less than two) peptide structures for these molecules, which correspond to the same molecular mass; however, this needs to be confirmed by NMR spectroscopy (Fig. 3).
The study of the difference between the masses of isolated components in analytical RP HPLC (Table 2) has shown that their difference (in mass) corresponds either to the “–CH2–” fragment (M = 14 Da) or to the H2O molecule (M = 18 Da), as is represented graphically in the scheme (Fig. 4). The dependence shows the molecular bonds of individual previously detected forms of peptaibols, as well as their expected molecular masses (in red). In view of the similar retention times of the components (Table 2, Fig. 2), as well as the existence of a chain of relationships between the molecular masses of these components (Fig. 3b), it can be supposed that the masses of the components correspond to compounds of the same chemical nature. Some peptaibol masses presented on the scheme were not only detected previously but also identified. For example, the mass of 1050 Da corresponds to EmiА per se [10], while the masses of 1036.7 and 1032.9 Da correspond to isoform EmiB [10] and dehydroform [14], respectively. Thus, the constructed scheme links different forms of previously detected emericillipsins A–E and previously undetected masses, 1004.7, 1018.7, 1046.8, and 1064.8 [10], which fit in a common homologous series. The presented scheme of masses is tentative in view of the fact that the similarity of chemical forms has not yet been confirmed by more precise spectral methods. Antifungal activity has been shown previously for the isolated isoforms of emericillipsins with masses of 1032.9 and 1036.7 Da [10].
The scheme of relationship between the masses of different molecular ions detected during the analysis (in black). The arrows show the relationship between the forms; the numbers above the arrows denote the difference of molecular masses between the connected units. Yellow rectangles are the forms EmiA and dEmiA (dehydroform); the tentative molecular masses are in red; red arrows show (supposedly) the relationships between the forms; the differences between the masses of associated molecular forms are indicated above the arrows.
Thus, it has been established that the new method of membrane-surface liquid cultivation of emericillipsin peptaibol producers leads to an increase in the diversity of synthesized natural forms of peptides and a 1.7 times increase in the yield of the major compound. The study of the potential diversity of AMPs synthesized by the producer at the level of the primary or spatial structure can be a stimulus for the detection of other types of their biological activity and the creation of novel synthetic peptides on their basis.
Change history
12 August 2023
An Erratum to this paper has been published: https://doi.org/10.1134/S0003683823320017
REFERENCES
Lau, J.L. and Dunn, M.K., Bioorg.Med. Chem., 2018, vol. 26, no. 10, pp. 2700–2707. https://doi.org/10.1016/j.bmc.2017.06.052
Casagrande, N., Borghese, C., Gabbatore, L., Morbiato, L., De Zotti, M., and Aldinucci, D., Int. J. Mol. Sci., 2021, vol. 22, no. 16, pp. 1–19. https://doi.org/10.3390/ijms22168362
Chen, C.H. and Lu, T.K., Antibiotics, 2020, vol. 9, no. 24, pp. 1–20. https://doi.org/10.3390/antibiotics9010024
Bin, H.A., Jiang, X., Bergen, P.J., and Zhu, Y., Int. J. Mol. Sci., 2021, vol. 22, no. 11691, pp. 1–56. https://doi.org/10.3390/ijms222111691
Aldholmi, M., Marchand, P., Ourliac-Garnier, I., Le Pape, P., and Ganesan, A., Pharmaceuticals, 2019, vol. 12, no. 4, pp. 1–21. https://doi.org/10.3390/ph12040182
Egorova-Zachernyuk, T.A., Shvets, V., Versluis, K., Heerma, W., Creemers, A.F.L., Nieuwenhuis, S.A.M., Lugtenburg, J., and Raap, J., J. Pept. Sci., 1996, vol. 2, pp. 341–350. https://doi.org/10.1002/psc.72
D'yachenko, I.A., Murashev, A.N., and Ovchinnikova, T.V., Toksikol. Vestn., 2008, no. 3, pp. 35–38.
Otto, A., Laub, A., Porzel, A., Schmidt, J., Wessjohann, L., Westermann, B., and Arnold, N., Eur. J. Org. Chem., 2015, vol. 34, pp. 7449–7459. https://doi.org/10.1002/ejoc.201501124
Sadykova, V.S., Gavryushina, I.A., Kuvarina, A.E., Markelova, N.N., Sedykh, N.G., Georgieva, M.L., Barashkova, A.C., and Rogozhin, E.A., Appl. Biochem. Microbiol., 2020, vol. 56, no. 3, pp. 292–297. https://doi.org/10.1134/S000368382003010
Kuvarina, A.E., Gavryushina, I.A., Kulko, A.B., Ivanov, I.A., Rogozhin, E.A., Georgieva, M.L., and Sadykova, V.S., J. Fungi, 2021, vol. 7, no. 153, pp. 1–19. https://doi.org/10.3390/jof7020153
Hao, X., Li, Sh., Ni, J., Wang, G., Li, F., Li, Q., Chen, Sh., Shu, J., and Gan, M., J. Nat. Prod., 2021, vol. 84, no. 11, pp. 2990–3000. https://doi.org/10.1021/acs.jnatprod.1c00834
Zhang, S.-H., Zhao, X., Xu, R., Yang, Y., Tang, J., Yue, X.-L., Wang, Y.-T., Tan, X.-Y., Zhang, G.-G., and Li, C.-W., Chem. Biodiversity, 2022, vol. 19, p. e202200627. https://doi.org/10.1002/cbdv.202200627
Grum-Grzhimaylo, A.A., Georgieva, M.L., Debets, A.J.M., and Bilanenko, E.N., IMA Fungus, 2013, vol. 4, no. 2, pp. 213–228. https://doi.org/10.5598/imafungus.2013.04.02.07
Kuvarina, A.E., Gavryushina, I.A., Sykonnikov, M.A., Efimenko, T.A., Markelova, N.N., Bilanenko, E.N., et al., Molecules, 2022, vol. 27, no. 1736, pp. 1–16. https://doi.org/10.3390/molecules27051736
Baranova, A.A., Georgieva, M.L., Bilanenko, E.N., Andreev, Y.A., Rogozhin, E.A., and Sadykova, V.S., Appl. Biochem. Microbiol., 2017, vol. 53, no. 6, pp. 703–710. https://doi.org/10.1134/S0003683817060035
Galloway, W.R.J.D., Bender, A., Welch, M., and Spring, D.R., Chem. Commun., 2009, pp. 2446–2462. https://doi.org/10.1039/b816852k
Funding
The work was supported by the Russian Science Foundation, project no. 20-75-00062 (Kuvarina A.E. and Sukonnikov M.A.)
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
The authors declare that they have no conflicts of interest. This article does not contain any studies involving animals or human participants performed by any of the authors.
Additional information
Translated by E. Makeeva
The original online version of this article was revised: Due to a retrospective Open Access order.
Rights and permissions
Open Access. This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Kuvarina, A.E., Sukonnikov, M.A., Rogozhin, E.A. et al. Formation of Various Antimicrobial Peptide Emericellipsin Isoforms in Emericellopsos alkalina under Different Cultivation Conditions. Appl Biochem Microbiol 59, 160–167 (2023). https://doi.org/10.1134/S0003683823020060
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1134/S0003683823020060