Abstract
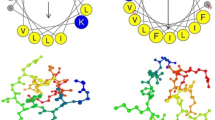
Pseudomonas aeruginosa easily obtains multidrug resistance and forms biofilm. These properties bring more challenges to its control. Antimicrobial peptides (AMPs) can inhibit the formation of biofilms. However the nature AMPs have many adverse factors which prevent its extensive application. Sacakin P has 43 amino acids and belongs to the IIa bacteriocins. In this study, the amino acid sequence with an α-helical structure at positions 18–33 of sakacin P was intercepted, and three AMPs were obtained by substituting amino acids at different positions with arginine (R) and lysine (K). The modified AMPs had good antibacterial activity against P. aeruginosa, and the minimum inhibitory concentration (MIC) was 2–8 µg/mL. P. aeruginosa biofilm formation was associated with nutrients and other environmental factors. Low pH and high NaCl and glucose concentrations effectively inhibited biofilm formation. Disinfectants such as 84 disinfectant, alcohol, and detergent did not effectively eliminate P. aeruginosa biofilms. Sakacin P16-R8 almost completely inhibited P. aeruginosa biofilm formation at 4 MIC and removed approximately 50% of a formed biofilm. The three AMPs were less toxic to eukaryotic cells, and the cell viability was >60% at 128 µg/mL of an AMP. The extracellular nucleic acid and protein content in the supernatant after treating P. aeruginosa with sakacin P16-R8 significantly increased with incubation time and peptide concentration. Flow cytometry showed that P. aeruginosa treated with 1 MIC and 2 MIC of sakacin P16-R8 showed 84.0 and 92.6% cell death, respectively. The results suggest that the modified AMPs exerted their antibacterial activity by increasing bacterial cell membrane permeability.









Similar content being viewed by others
REFERENCES
Rossolini, G.M. and Mantengoli, E., Clin. Microbiol. Infect., 2005, vol. 11, pp. 17–32. https://doi.org/10.1134/S1068162020070080
Bai, X., Chai, X.L., Yan, X.L., Gao, H., Zhang, H.L., and Zhao, J.J., Anim. Husb. Vet. Med., 2011, vol. 43, pp. 31–35.
Hancock R.E. and Speert D.P., Drug Resist. Update, 2000, vol. 3, pp. 247–255. https://doi.org/10.1054/drup.2000.0152
Lambert, P.A., J. R. Soc. Med., 2002, vol. 95, pp. 22–26.
Harmsen, M., Yang, L., Pamp, S.J., and Tolker-Nielsen, T., FEMS Immunol. Med. Microbiol., 2003, vol. 59, pp. 253–268. https://doi.org/10.1111/j.1574-695X.2010.00690.x
Mah, T.F. and O’Toole, G.A., Trends Microbiol., 2001, vol. 9, pp. 34–39. https://doi.org/10.1016/S0966-842X(00)01913-2
Irie, Y., Borlee, B.R., O’Connor, J.R., Hill, P.J., Harwood, C.S., and Wozniak, D.J., Proc. Natl. Acad. Sci. U. S. A., 2012, vol. 109, pp. 20632–20636. https://doi.org/10.1073/pnas.1217993109
Cady, N.C., McKean, K.A., Behnke, J., Kubec, R., Mosier, A.P., and Kasper, S.H., PLoS One, 2012, vol. 7, art. e38492. https://doi.org/10.1371/journal.pone.0038492
Borlee, B.R., Goldman, A.D., Murakami, K., Samudrala, R., Wozniak, D.J., and Parsek, M.R., Mol. Microbiol., 2010, vol. 75, pp. 827–842. https://doi.org/10.1111/j.1365-2958.2009.06991.x
Storz, M.P., Maurer, C.K., Zimmer, C., Wagner, N., Brengel, C., and de Jong, J.C., J. Am. Chem. Soc., 2012, vol. 134, pp. 16143–16146. https://doi.org/10.1021/ja3072397
Ganguly, K., Wu, R., Ollivault-Shiflett, M., Goodwin, P.M., Silks, L.A., and Iyer, R., J. Drug Target., 2011, vol. 19, pp.528–539. https://doi.org/10.3109/1061186X.2010.519032
da Costa, J.P., Cova, M., Ferreira, R., and Vitorino, R., Appl. Microbiol. Biotechnol., 2005, vol. 99, pp. 2023–2040. https://doi.org/10.1007/s00253-015-6375-x
de la Fuente-Núñez, C., Reffuveille, F., Fernández, L., and Hancock, R.E., Curr. Opin. Microbiol., 2013, vol. 16, pp. 580–589. https://doi.org/10.1016/j.mib.2013.06.013
Bahnsen, J.S., Franzyk, H., Sayers, E.J., Jones, A.T., and Nielsen, H.M., Pharm. Res., 2015, vol. 32, pp. 1546–1556.
Lv, Y.F., Wang, J.J., Gao, H., Wang, Z.Y., Dong, N., and Ma, Q.Q., PLoS One, 2014, vol. 9, art. e86364. https://doi.org/10.1371/journal.pone.0086364
Hasper, H.E., Kramer, N.E., Smith, J.L., Hillman, J.D., Zachariah, C., and Kuipers, O.P., Science, 2006, vol. 313, pp. 1636–1637. https://doi.org/10.1126/science.1129818
Hao, G., Shi, Y.H., Tang, Y.L., and Le, G.W., J. Microbiol., 2013, vol. 51, pp. 200–206. https://doi.org/10.1007/s12275-013-2441-1
Brogden, N.K. and Brogden, K.A., Int. J. Antimicrob. Agents, 2011, vol. 38, pp. 217–225. https://doi.org/10.1016/j.ijantimicag.2011.05.004
Chen, Y., Shapira, R., Eisenstein, M., and Montville, T.J., Appl. Environ. Microb., 1997, vol. 63, pp. 524. https://doi.org/10.1128/AEM.63.2.524-531.1997
Nilsson, L., Huss, H.H., and Gram, L., Int. J. Food Mocrobiol., 1997, vol. 38, pp. 217–227. https://doi.org/10.1016/S0168-1605(97)00111-6
Katla, T., Møretrø, T., Sveen, I., Aasen, I.M., Axelsson, L., and Rørvik, L.M., J. Appl. Microbiol., 2010, vol, 93, pp. 191–196. https://doi.org/10.1046/j.1365-2672.2002.01675.x
Ma, Z., Wei, D.D., Yan, P., Zhu, X., Shan, A.S., and Bi, Z.P., Biomaterials, 2015, vol. 52, pp. 517–530. https://doi.org/10.1016/j.biomaterials.2015.02.063
Lyu, Y.F., Yang, Y., Lyu, X.T., Dong, N., and Shan, A.S., Sci. Rep. (UK), 2016, vol. 6, p. 27258. https://doi.org/10.1038/srep27258
Sun, J.B., Xia, Y.Q., Li, D., Du, Q., and Liang, D.H., Biochim. Biophys. Acta., 2014, vol. 1838, pp. 2985–2993. https://doi.org/10.1038/srep27258
Ong, Z.Y., Wiradharma, N., and Yang, Y.Y., Adv. Drug Deliver. Rev., 2014, vol. 78, pp. 28–45. https://doi.org/10.1016/j.addr.2014.10.013
Ong, Z.Y., Gao, S.J., and Yang, Y.Y., Adv. Funct. Mater., 2013, vol. 23, pp. 3682–3692. https://doi.org/10.1002/adfm.201202850
Liu, X.L., Cao, R., Wang, S., Jia, J.L., and Fei, H., J. Med. Chem., 2016, vol. 59, pp. 5238. https://doi.org/10.1021/acs.jmedchem.5b02016
Ma, Z., Yang, J., Han, J.Z., Gao, L., Liu, H.X., and Lu, Z.X., J. Med. Chem., 2016, vol. 59, pp. 10946–10962. https://doi.org/10.1021/acs.jmedchem.6b00922
Funding
This study was supported by grants from the National Key R&D Program of China (Grant no. 2018YFC1602500), Agricultural Independent Innovation Program in Jiangsu Province (Grant no. CX (18)3053), NSFC (Grant no. 31972174).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
COMPLIANCE WITH ETHICAL STANDARDS
This article does not contain any studies involving animals or human participants performed by any of the authors. This article does not contain any studies involving human participants performed by any of the authors.
Conflict of Interests
The authors declare that they have no conflicts of interest.
Additional information
The authors contributed equally to the work.
Rights and permissions
About this article
Cite this article
Bingxue Chang, Ma, W., Lu, Z. et al. Design and Antibacterial Mechanism of Peptides Derived from Sakacin P. Russ J Bioorg Chem 48, 399–410 (2022). https://doi.org/10.1134/S1068162022020054
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1134/S1068162022020054