Abstract
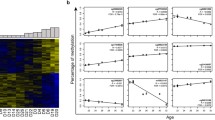
On the basis of bioinformatics and statistical analysis of GEO projects to determine the genome-wide profile of human DNA methylation, a list of 41 CpG dinucleotides with high predictive potential was formed to create models for predicting human age from biological samples. The methylation level was determined for 1208 samples of individuals from the Republic of Belarus (275—blood, 466—buccal epithelium, 467—semen). The correlation coefficients R were calculated, and mathematical models for determining the age of individual were constructed. The mean value of the accuracy of age prediction from blood samples using 12 CpG dinucleotides was 3.4 years (3.3 for men, 3.5 for women); from buccal epithelium samples using 6 CpG dinucleotides, it was 4.6 years (4.5 for men, 4.7 for women); from semen samples using 5 CpG dinucleotides, it was 3.0 years. The results obtained will be used as a basis for the development of calculators for predicting the age of an individual based on traces of biological character for forensic experts.

Similar content being viewed by others
REFERENCES
Zolotarenko, A.D., Chekalin, E.V., and Bruskin, S.A., Modern molecular genetic methods for age estimation in forensics, Russ. J. Genet., 2019, vol. 55, no. 12, pp. 1460–1471. https://doi.org/10.1134/S1022795419120147
Kil’chevskii, A., Mosse, I., Shapturenko, M., and Burakova, A., Contribution of genetics to forensic science in Belarus, Nauka Innovatsii, 2020, no. 10(212), pp. 22–28.
Hannum, G., Guinney, J., Zhao, L., et al., Genome-wide methylation profiles reveal quantitative views of human aging rates, Mol. Cell, 2013, vol. 24, pp. 359–367. https://doi.org/10.1016/j.molcel.2012.10.016
Kurdyukov, S. and Bullock, M., DNA methylation analysis: choosing the right method, Biology (Basel), 2016, vol. 5, no. 1, pp. e1–e21. https://doi.org/10.3390/biology5010003
Bocklandt, S., Lin, W., Sehl, M.E., et al., Epigenetic predictor of age, PLoS One, 2011, vol. 6, pp. 1–6. https://doi.org/10.1371/journal.pone.0014821
Naue, J., Hoefsloot, H.C.J., Mook, O.R.F., et al., Chronological age prediction based on DNA methylation: massive parallel sequencing and random forest regression, Forensic Sci. Int. Genet., 2017, vol. 31, pp. 19–28. https://doi.org/10.1016/j.fsigen.2017.07.015
Park, J.L., Kim, J.H., Seo, E., et al., Identification and evaluation of age-correlated DNA methylation markers for forensic use, Forensic Sci. Int. Genet., 2016, vol. 23, pp. 64–70. https://doi.org/10.1016/j.fsigen.2016.03.005
Vidaki, A., Daniel, B., and Syndercombe Court, D., DNA methylation-based forensic age prediction using artificial neural networks and next generation sequencing, Forensic Sci. Int. Genet., 2017, vol. 28, pp. 225–236. https://doi.org/10.1016/j.fsigen.2017.02.009
Zbiec-Piekarska, R., Spolnicka, M., Kupiec, T., et al., Development of a forensically useful age prediction method based on DNA methylation analysis, Forensic Sci. Int. Genet., 2015, vol. 17, pp. 173–179. https://doi.org/10.1016/j.fsigen.2015.05.001
Eipel, M., Mayer, F., Arent, T., et al., Epigenetic age predictions based on buccal swabs are more precise in combination with cell type-specific DNA methylation signatures, Aging (Albany, NY), 2016, vol. 8, no. 5, pp. 1034–1048. https://doi.org/10.18632/aging.100972
McEwen, L.M., O’Donnell, K.J., McGill, M.G., et al., The PedBE clock accurately estimates DNA methylation age in pediatric buccal cells, Proc. Natl. Akad. Sci, U.S.A., 2020, vol. 117, no. 38, pp. 23329–23335. https://doi.org/10.1073/pnas.1820843116
Koop, B.E., Mayer, F., Gündüz, T., et al., Postmortem age estimation via DNA methylation analysis in buccal swabs from corpses in different stages of decomposition—a “proof of principle” study, Int. J. Legal Med., 2021, vol. 135, no. 1, pp. 167–173. https://doi.org/10.1007/s00414-020-02360-7
Dongen, J., Ehli, E.A., Jansen, R., et al., Genome-wide analysis of DNA methylation in buccal cells: a study of monozygotic twins and mQTLs, Epigenet. Chromatin, 2018, vol. 11, no. 1, pp. e1–e14. https://doi.org/10.1186/s13072-018-0225-x
Wozniak, A., Heidegger, A., Piniewska-Rog, D., et al., Development of the VISAGE enhanced tool and statistical models for epigenetic age estimation in blood, buccal cells and bones, Aging (Albany, NY), 2021, vol. 13, no. 5, pp. 6459–6484. https://doi.org/10.18632/aging.202783
Lee, H.Y., Jung, S.E., Oh, Y.N., et al., Epigenetic age signatures in the forensically relevant body fluid of semen: a preliminary study, Forensic Sci. Int. Genet., 2015, vol. 19, pp. 28–34. https://doi.org/10.1016/j.fsigen.2015.05.014
Alsaleh, H., McCallum, N.A., Halligan, D.L., and Haddrill, P.R., A multi-tissue age prediction model based on DNA methylation analysis, Forensic Sci. Int. Genet., Suppl. Ser., 2017, vol. 6, pp. 62–64. https://doi.org/10.1016/j.fsigss.2017.09.056
Li, L., Song, F., Huang, Y., Zhu, H., and Hou, Y., Age-associated DNA methylation determination of semen by pyrosequencing in Chinese Han population, Forensic Sci. Int. Genet., 2017, vol. 6, pp. e99–e100. https://doi.org/10.1016/j.fsigss.2017.09.042
Jenkins, T.G., James, E.R., Alonso, D.F., et al., Cigarette smoking significantly alters sperm DNA methylation patterns, Andrology, 2017, vol. 5, no. 6, pp. 1089–1099. https://doi.org/10.1111/andr.12416
Lee, J.W., Choung, C.M., Jung, J.Y., et al., A validation study of DNA methylation-based age prediction using semen in forensic casework samples, Leg. Med. (Tokyo), 2018, vol. 31, pp. 74–77. https://doi.org/10.1016/j.legalmed.2018.01.005
Fleckhaus, J., Freire-Aradas, A., Rothschild, M.A., and Schneider, P.M., Impact of genetic ancestry on chronological age prediction using DNA methylation analysis, Forensic Sci. Int. Genet., 2017, vol. 6, pp. e399–e400. https://doi.org/10.1016/j.fsigss.2017.09.162
Donkin, I. and Barres, R., Sperm epigenetics and influence of environmental factors, Mol. Metab., 2018, vol. 14, pp. 1–11. https://doi.org/10.1016/j.molmet.2018.02.00
Kipen, V.N., Bogdanova, M.V., and Burakova, A.A., Minimum sample size justification for prediction of human chronological age, Mol. Prikl. Genet., 2021, vol. 30, pp. 39–48. https://doi.org/10.47612/1999-9127-2021-30-39-48
Kipen, V.N., Bogdanova, M.V., Burakova, A.A., et al., Predictive capacity of CpG markers for determination of human chronological age, Mol. Prikl. Genet., 2020, vol. 28, pp. 70–80.
Hong, S.R., Jung, S., Lee, E.H., et al., DNA methylation-based age prediction from saliva: high age predictability by combination of 7 CpG markers, Forensic Sci. Int. Genet., 2017, vol. 29, pp. 118–125. https://doi.org/10.1016/j.fsigen.2017.04.006
Funding
The present study was performed within the framework of activity 2 “Development of a Methodology for Determining the Probable Age of an Individual on the Basis of DNA Characteristics” of the scientific and technical program of the Union State “Development of Innovative Gene-Geographic and Genomic Technologies for Identification of an Individual and Human Traits on the Basis of the Study of the Gene Pools of the Union State Regions” (DNA identification).
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Conflict of interest. The authors declare no conflict of interest.
Statement of compliance with standards of research involving humans as subjects. All procedures performed in studies involving human participants were in accordance with the ethical standards of the institutional and/or national research committee and with the 1964 Helsinki Declaration and its later amendments or comparable ethical standards. Informed consent was obtained from all individual participants involved in the study.
Additional information
Translated by A. Kazantseva
Rights and permissions
About this article
Cite this article
Lemesh, V.A., Kipen, V.N., Bahdanava, M.V. et al. Determination of Human Chronological Age from Biological Samples Based on the Analysis of Methylation of CpG Dinucleotides. Russ J Genet 57, 1389–1397 (2021). https://doi.org/10.1134/S1022795421120097
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1134/S1022795421120097