Abstract
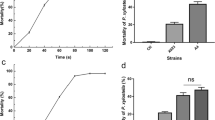
Chitinase is one of the most important enzymes due to its diversity and a variety of potential uses. This study is an attempt to enhance chitinase production for antifungal biocontrol by subjecting Bacillus thuringiensis NM101-19 strain grown on shrimp shell wastes to various doses of gamma irradiation. Six mutants (BM-4, BM-6, BM-8, BM-12, BM-15, and BM-17) obtained at gamma ray doses of 40, 60, 80, 100, 120 and 140 Gy, respectively produced higher levels of chitinolytic activities in comparision to the wild-type strain. The BM-15 mutant strain showed the highest chitinase production (65.41 U/mL) which was 2.60 times more than the wild type (25.11 U/mL). Biocontrol efficacy of the mutants was statistically superior to the wild-type strain against all tested phytopathogens. Random amplified polymorphic DNA (RAPD) and inter simple sequence repeat (ISSR) PCR techniques, with five primers for each, were used in order to detect the variation in DNA profile between the mutant and wild-type strains in response to gamma-radiation treatments. RAPD and ISSR analysis indicated the appearance and disappearance of DNA polymorphic bands at different gamma ray doses. The results confirmed that the mutagenesis technique is a potent strategy to enhance the chitinase activity for industrial and agricultural purposes.
Similar content being viewed by others
References
Abbasi, S., Safaie, N., Bakhsh, M.S., and Shahbazi, S., Biocontrol activities of gamma induced mutants of Trichoderma harzianum against some soilborne fungal pathogens and their DNA fingerprinting, Iran. J. Biotech., 2016, vol. 14, pp. 260–269.
Adsul, M.G., Bastawde, K.B., and Varma, A.J., Strain improvement of Penicillium janthinellum NCIM 1171 for increased cellulase production, Bioresour. Technol., 2007, vol. 98, pp. 1467–1473.
Bansode, V.B. and Bajekal, S.S., Characterization of chitinases from microorganisms isolated from Lonar Lake, Ind. J. Biotech., 2006, vol. 5, pp. 357–363.
Barboza-Corona, J.E., Reyes-Rios, D.M., Salcedo-Hernandez, R., and Bideshi, D.K., Molecular and biochemical characterization of an endochitinase, ChiA-HD73, from Bacillus thuringiensis subsp. kurstaki HD-73, Mol. Biotechnol., 2008, vol. 39, pp. 29–37.
Berini, F., Presti, I., Beltrametti, F., Pedroli, M., Varum, K.M., Pollegioni, L., Sjoling, S., and Marinelli, F., Production and characterization of a novel antifungal chitinase identified by functional screening of a suppressive-soil metagenome, Microb. Cell Fact., 2017, vol. 16, pp. 16–31.
Budi, S.W., Van Tuinen, D., Arnould, C., Dumas-Gaudut, E., Gianinazzi-Pearson, V., and Gianinazzi, S., Hydrolytic enzyme activity of Paenibacillus sp. strain B2 and effect of antagonistic bacterium on cell wall integrity of two soil-borne pathogenic fungi, Appl. Soil. Ecol., 2000, vol. 15, pp. 191–199.
Chandrasekaran, R., Revathi, K., Nisha, S., Kirubakaran, S.A., Sathish-Narayanan, S., and Senthil, N.S., Physiological effect of chitinase purified from Bacillus subtilis against the tobacco cutworm Spodoptera litura Fab., Pestic. Biochem. Physiol., 2012, vol.104, pp. 65–71.
Chang, W.T., Chen, M., and Wang, S.L., An antifungal chitinase produced by Bacillus subtilis using chitin waste as a carbon source, World J. Microbiol. Biotechnol., 2010, vol. 26, pp. 945–950.
De la Vega, L.M., Barboza-Corona, J.E., Aguilar-Uscanga, M.G., and Ramirez-lepe, M., Purification and characterization of an exochitinase from Bacillus thuringiensis subsp. aizawai and its action against phytopathogenic fungi, Can. J. Microbiol., 2006, vol. 52, pp. 651–657.
Dhananasekaran, S., Palanivel, R., and Pappu, S., Adsorption of methylene blue, bromophenol blue, and coomassie brilliant blue by α-chitin nanoparticles, J. Adv. Res., 2016, vol. 7, pp. 113–124.
Gomaa, E.Z., Chitinase production by Bacillus thuringiensis and Bacillus licheniformis, their potential in antifungal biocontrol, J. Microbiol., 2012, vol. 50, pp. 103–111.
Gundry, M., and Vijg, J., Direct mutation analysis by highthroughput sequencing, from germline to low-abundant, somatic variants, Mut. Res., 2012, vol. 729, pp. 1–15.
Hart, L., Zach, S., and Seidl-Seiboth, V., Fungal chitinases, diversity, mechanistic properties and biotechnological potential, Appl. Microbiol. Biotechnol., 2012, vol. 93, pp. 533–543.
Hayes, M., Carney, B., Slater, J., and Bruck, W., Mining marine shellfish wastes for bioactive molecules, chitin and chitosan—Part A, extraction methods, Biotechnol. J., 2008, vol. 3, pp. 871–877.
Islam, R., Jeong, Y.T., Ryu, Y.J., Song, C.H., and Lee, Y.S., Isolation, identification and optimal culture conditions of Streptomyces albidoflavus C247 producing antifungal agents against Rhizoctonia solani AG2-2, Mycobiology, 2009, vol. 37, pp. 114–120.
Kamil, Z., Rizk, M., Saleh, M., and Moustafa, S., Isolation and identification of rhizosphere soil chitinolytic bacteria and their potential in antifungal biocontrol, Global J. Mol. Sci., 2007, vol. 2, pp. 57–66.
Kavil, S.P., Pallavesam, A., Sumesh, K.M., Bhairappanavar, S.B., Sadashiv, S.O., Chandrashekhar, U., and Catherine, R.P., Improvement of chitinase producing capacity of Shigella sp. by random mutation and altering the nutrient component of production media, World J. Pharm. Sci., 2014, vol. 3, pp. 1717–1726.
Khoushab, F. and Yamabhai, M., Chitin research revisited, Mar. Drugs., 2010, vol. 8, pp. 1988–2012.
Kishore, G.K., Pande, S., and Podile, A.R., Biological control of late leaf spot of peanut, Arachis hypogaea L., with chitinolytic bacteria, Phytopathol., 2005, vol. 95, pp. 1157–1165.
Liu, D., Cai, J., Xie, C., Liu, C., and Chen, Y.H., Purification and partial characterization of a 36-kDa chitinase from Bacillus thuringiensis spp. colmeri, and its biocontrol potential, Enz. Microb. Technol., 2010, vol. 46, pp. 252–256.
Mallakuntla, M.K., Vaikuntapu, P.R., Bhuvanachandra, B., Das, S.N., and Podile, A.R., Transglycosylation by a chitinase from Enterobacter cloacae subsp. cloacae generates longer chitin oligosaccharides, Sci. Rep., 2017, vol. 7, pp. 5113–5125.
Marsjan, P.A. and Oldenbroek, J.K., Molecular markers, a tool for exploring gene diversity, in The State of the World’s Animal Genetic Resources for Food and Agriculture. FAO Research Report, Rome: FAO, 2007, pp. 359–379.
Mohamadi, A.S., Shahbazi, S., and Askari, H., Investigation of γ-radiation on morphological characteristics and antagonist potential of Trichoderma viride against Rhizoctonia solani, Int. Res. J. Appl. Basic Sci., 2014, vol. 8, pp. 329–336.
Najafi, M.B. and Pezeshki, P., Bacterial mutation; types, mechanisms and mutant detection methods. A review, Euro. Sci. J., 2013, vol. 4, pp. 1857–7881.
Neeraja, C., Anil, K., Purushotham, P., Suma, K., Sarma, P., and Moerschbacher, B.M., Biotechnological approaches to develop bacterial chitinases as a bioshield against fungal diseases of plants, Crit. Rev. Biotechnol., 2010, vol. 30, pp. 231–241.
Nurdebyandaru, N., Mubarik, N.R., and Prawasti, T.S., Chitinolytic bacteria isolated from Chili rhizosphere, Chitinase characterization and application as biocontrol for Aphis gossypii, Microb. Ind., 2010, vol. 4, pp. 13–107.
Ozgen, A., Sezen, K., Demir, I., Demirbag, Z., and Nalcacioglu, R., Molecular characterization of chitinase genes from a local isolate of Serratia marcescens and their contribution to the insecticidal activity of Bacillus thuringiensis strains, Curr. Microbiol., 2013, vol. 67, pp. 499–504.
Raghuwanshi, S., Deswal, D., and Karp M., Bioprocessing of enhanced cellulase production from a mutant of Trichoderma asperellum RCK2011 and its application in hydrolysis of cellulose, Fuel, 2014, vol. 124, pp. 183–189.
Roopavathi, A.S., Vigneshwari, R., and Jayapradha, R., Chitinase, production and applications, J. Chem. Pharm. Res., 2015, vol. 7, pp. 924–931.
Sharma, N., Sharma, K.P., Gaur, R.K., and Gupta, V.K., Role of chitinase in plant defense, Asian J. Biochem., 2011, vol. 6, pp. 29–37.
Shehata, A.I., Phylogenetic diversity of Staphylococcus aureus by random amplification of polymorphic DNA, Aust J. Basic Appl. Sci., 2008, vol. 2, pp. 858–863.
Spadaro, D. and Lodovica, M., Improving the efficacy of biocontrol agents against soil borne pathogens, Crop. Prot., 2005, vol. 24, pp. 601–613.
Strange, R.N. and Scott, P.R., Plant disease, a threat to global food security, Ann. Rev. Phytopathol., 2005, vol. 43, pp. 83–116.
Suganthi, M., Arvinth, S., Kumar, R.R., and Chandrashekara, K.N., Detection of chitinase activity and its characterization from Pseudomonas fluorescens of tea rhizosphere, J. Plant Crop., 2015, vol. 43, pp. 236–239.
Sun, Y., Liu, W., Han, B., Zhang, J., and Liu, B., Purification and characterization of two types of chitosanase from Microbacterium sp., Biotechnol. Lett., 2006, vol. 28, pp. 1393–1399.
Wang, S.L., Chao, C.H., Liang, T.W., and Chen, C.C., Purification and characterization of protease and chitinase from Bacillus cereus TKU006 and conversion of marine wastes by these enzymes, Mar. Biotechnol., 2009, vol. 11, pp. 334–344.
Xie, C., Chen, Y., Cai, J., and Liu, C., Essential expression and inducible synthesis polymorphism of chitinase in Bacillus thuringiensis, Sheng. Wu. Gong. Cheng. Xue. Bao., 2010, vol. 26, pp. 1532–1538.
Yadav, E., Pathak, D.V., Sharma, S.K., Kumar, M., and Sharma, P.K., Isolation and characterization of mutants of Pseudomonas maltophila PM-4 altered in chitinolytic activity and antagonistic activity against root rot pathogens of clusterbean, Cyamopsis tetragonoloba, Ind. J. Microbiol., 2007, vol. 47, pp. 64–71.
Author information
Authors and Affiliations
Corresponding author
Additional information
The article is published in the original.
Rights and permissions
About this article
Cite this article
Gomaa, E.Z., El-Mahdy, O.M. Improvement of Chitinase Production by Bacillus thuringiensis NM101-19 for Antifungal Biocontrol through Physical Mutation. Microbiology 87, 472–485 (2018). https://doi.org/10.1134/S0026261718040094
Received:
Published:
Issue Date:
DOI: https://doi.org/10.1134/S0026261718040094