Abstract
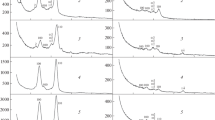
Bacterial nanocellulose, which was produced by Medusomyces gisevii Sa-12 cultured in enzymatic hydrolysates of Miscanthus, was studied by X-ray diffraction. The characteristics of the supramolecular structure of the crystalline component of cellulose samples, such as the degree of crystallinity and the size and shape of elementary fibrils, were determined. The atomic structure was compared with the known structural models of cellulose, and the synthesized bacterial nanocellulose was shown to be cellulose Iα. The lengths and angles of the triclinic unit cell were determined. The composition of the culture medium made from enzymatic hydrolysates of Miscanthus was found to have an effect on the shape and size of elementary fibrils and have no effect on the degree of crystallinity and the fraction of cellulose allomorph Іα. The use of Medusomyces gisevii Sa-12 as the producing organism allows the preparation of bacterial nanocellulose with a high degree of crystallinity in the range of 86–93% containing cellulose allomorph Іα as the major component.




Similar content being viewed by others
REFERENCES
T. Heinze, O. A. El Seoud, and A. Koschella, Cellulose Derivatives. Synthesis, Structure, and Properties (Springer, Switzerland, 2018).
Y. G. Baklagina, V. V. Klechkovskaya, S. V. Kononova, et al., Crystallogr. Rep. 63 (3), 303 (2018).
L. A. Aleshina, E. K. Gladysheva, V. V. Budaeva, et al., Crystallogr. Rep. 63 (6), 955 (2018).
A. Kuila and V. Sharma, Principles and Applications of Fermentation Technology (Scrivener, Beverly, 2018).
H. Barud, R. R. Silva, H. Barud, et al., Carbohyd. Polym. 153, 406 (2016).
U. M. Islam, M. W. Ullah, S. Khana, et al., Int. J. Biol. Macromol. 102, 1166 (2017).
M. Gama, F. Dourado, and S. Bielecki, Bacterial Nanocellulose from Biotechnology to Bio-Economy (Elsevier, Amsterdam, 2016).
M. T. Luo, C. Zhao, C. Huang, et al., Indian J. Microbiol. 57 (4), 393 (2017).
M. Velasquez-Riano and V. Bojaca, Cellulose 24, 2677 (2017).
A. Vazquez, M. L. Foresti, P. Cerrutti, and M. Galvagno, J. Polym. Environ. 21 (2), 545 (2013).
C. Molina-Ramírez, C. Castro, R. Zuluaga, and P. Gañán, J. Polym. Environ. 26 (2), 830 (2018).
F. Mohammadkazemi, Am. J. Appl. Indr. Chem. 1 (1), 10 (2017).
V. V. Budaeva, E. A. Skiba, O. V. Baibakova, et al., Catal. Ind. 8 (1), 81 (2016).
E. A. Skiba, V. V. Budaeva, O. V. Baibakova, et al., Catal. Ind. 8 (2), 168 (2016).
Y. A. Gismatulina and V. V. Budaeva, Ind. Crop. Prod. 109, 227 (2017).
O. V. Baibakova, E. A. Skiba, V. V. Budaeva, and V. N. Zolotukhin, Polzunovskii Vestn. 1 (4), 147 (2016).
V. V. Budaeva, E. I. Makarova, and Yu. A. Gismatulina, Key Eng. Mater. 670, 202 (2016).
M. N. Denisova, E. I. Makarova, I. N. Pavlov, et al., Biotechnol. Appl. Biochem. 178 (6), 1196 (2016).
E. K. Gladysheva, E. A. Skiba, V. N. Zolotukhin, and G. V. Sakovich, Appl. Biochem. Microbiol. 54 (2), 179 (2018).
M. A. Torlopov, V. I. Mikhaylov, E. V. Udoratina, et al., Cellulose 25 (2), 1031 (2018).
P. T. Chandrasekaran, N. K. Bari, and S. Sinha, Cellulose 24, 4367 (2017).
M. Khandelwal, A. H. Windle, and N. Hessler, J. Mater. Sci. 5, 4839 (2016).
P. C. S. Faria-Tischer, C. A. Tischer, L. Heux, et al., Mat. Sci. Eng. C-Bio. S. 51 (1), 167 (2015).
J. Sugiyama, R. Vuong, and H. Chanzy, Macromolecules 24, 4168 (1991).
A. D. French, Cellulose 21, 885 (2014).
Y. Nishiyama, J. Sugiyama, H. Chanzy, and P. Langan, J. Am. Chem. Soc. 125, 14300 (2003).
Y. Nishiyama, P. Langan, and H. Chanzy, J. Am. Chem. Soc. 124, 9074 (2002).
A. B. Poma, M. Chwastyk, and M. Cieplak, Cellulose 23 (3), 1573 (2016).
L. A. Aleshina, S. V. Glazkova, L. A. Lugovskaya, et al., Khim. Rastit. Syr’ya, No. 1, 5 (2001).
K. Cheng, J. Catchmark, and A. Demirci, J. Biol. Eng. 3 (12), 1 (2009).
R. J. Moon, A. Martini, J. Nairn, et al., Chem. Soc. Rev. 40, 3941 (2011).
L. H. Thomas, V. T. Forsyth, A. Šturcová, et al., Plant Physiol. 161, 465 (2013).
A. A. Baker, W. Helbert, J. Sugiyama, et al., Biophys. J. 79, 1139 (2000).
S.-Y. Ding, S. Shuai, and Y. Yining, Cellulose 2, 863 (2014).
V. G. Savchenko, G. G. Belozerskaya, V. A. Makarov, et al., RF Patent No. 2624242 (August 10, 2016), Byull. Izobret., 2017, no. 19, p. 19.
Funding
The study was supported by the Russian Science Foundation (project no. 17-19-01054).
Author information
Authors and Affiliations
Corresponding authors
Additional information
Translated by T. Safonova
Rights and permissions
About this article
Cite this article
Aleshina, L.A., Gladysheva, E.K., Budaeva, V.V. et al. X-ray Diffraction Study of Bacterial Nanocellulose Produced by Medusomyces Gisevii Sa-12 Cultured in Enzymatic Hydrolysates of Miscanthus. Crystallogr. Rep. 64, 914–919 (2019). https://doi.org/10.1134/S1063774519060026
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1134/S1063774519060026