-
PDF
- Split View
-
Views
-
Cite
Cite
Nevedha Ravindran, Harshil Ramachandran, Nikhil Job, Arpita Yadav, K.P. Vaishak, Sourav Datta, B-box protein BBX32 integrates light and brassinosteroid signals to inhibit cotyledon opening, Plant Physiology, Volume 187, Issue 1, September 2021, Pages 446–461, https://doi.org/10.1093/plphys/kiab304
Close - Share Icon Share
Abstract
Cotyledon opening is a key morphological change that occurs in seedlings during de-etiolation. Brassinosteroids (BRs) inhibit the opening of cotyledons in darkness while light promotes cotyledon opening. The molecular regulation of the interplay between light and BR to regulate cotyledon opening is not well understood. Here, we show the B-box protein BBX32 negatively regulates light signaling and promotes BR signaling to inhibit cotyledon opening in Arabidopsis (Arabidopsis thaliana). BBX32 is highly expressed in the cotyledons of seedlings during de-etiolation. bbx32 and 35S:BBX32 seedlings exhibit enhanced and reduced cotyledon opening, respectively, in response to both light and brassinazole treatment in dark, suggesting that BBX32 mediates cotyledon opening through both light and BR signaling pathways. BBX32 expression is induced by exogenous BR and is upregulated in bzr1-1D (BRASSINAZOLE RESISTANT1-1D). Our in vitro and in vivo interaction studies suggest that BBX32 physically interacts with BZR1. Further, we found that PHYTOCHROME-INTERACTING FACTOR 3 (PIF3) interacts with BBX32 and promotes BR-mediated cotyledon closure. BBX32, BZR1, and PIF3 regulate the expression of common target genes that modulate the opening and closing of cotyledons. Our work suggests BBX32 integrates light and BR signals to regulate cotyledon opening during de-etiolation.
Introduction
After germination, a seed buried under the ground undergoes several morphological changes in its quest to reach light. Two organs that undergo dramatic variations during this transition are the hypocotyl and cotyledons. In dark, the seedling develops a long hypocotyl and cotyledons are closed forming an apical hook. Once the seedling reaches light, the elongation of the hypocotyl is restrained, the apical hook is released and cotyledons open up and begin to expand. While the inhibition of hypocotyl elongation in light has been extensively investigated, recent studies have started elucidating the spatio-temporal mechanistic regulation of each step in cotyledon development during de-etiolation (Sun et al., 2016; Dong et al., 2019; He et al., 2019). The kinetics of cotyledon opening and expansion in light differs a lot suggesting that they are distinct processes. While the opening of cotyledons initiates within 4 h after transfer to light and is completed within 2 d after the transition, the expansion of cotyledons can continue for 8 d after exposure to light (Dong et al., 2019). While protected by the closed cotyledons in darkness, light mediates the opening of cotyledons to expose the shoot apical meristem (SAM), initiating the development of the aerial parts of the plant. Although several studies indicate that light and the phytohormone brassinosteroid (BR) play antagonistic roles in the opening and closing of cotyledons, respectively, the molecular regulation of this process is not well defined (Wang et al., 2012). Recently, it was reported that light induces the expression of BR biosynthesis genes in the apical hook to promote hook opening in light, making the crosstalk between light and BR even more complex (Hamasaki et al., 2020).
The morphological and developmental changes that a plant undergoes in light or in dark are referred to as photomorphogenesis and skotomorphogenesis, respectively. Light signaling involves the perception of light by photoreceptors, followed by the transduction of the signal to ultimately modulate the expression of light-regulated genes (Yadav et al., 2020). The signal transducers can be grouped into positive or negative regulators of light signaling based on their photomorphogenesis promoting or inhibiting activities, respectively. The photoreceptors—phytochromes, cryptochromes, phototropins, and UV RESISTANCE LOCUS 8 (UVR8)—perceive light of different wavelengths to initiate light signal transduction (Yadav et al., 2020). Downstream of the photoreceptors, the bZIP transcription factor HY5 (ELONGATED HYPOCOTYL5) functions as a key positive regulator of light signaling. Furthermore, several members of the B-box containing protein family (BBX) have been shown to interact with HY5 (Datta et al., 2006, 2007, 2008; Gangappa et al., 2013; Xu et al., 2016; Zhang et al., 2017; Lin et al., 2018). Some BBX proteins act together with HY5 to promote photomorphogenesis, while some others inhibit its activity thereby acting as negative regulators of light signaling (Gangappa and Botto, 2014; Xu et al., 2016; Job et al., 2018; Yadukrishnan et al., 2018; Vaishak et al., 2019). The E3 ubiquitin ligase CONSTITUTIVE PHOTOMORPHOGENIC 1 (COP1) and the PHYTOCHROME-INTERACTING FACTOR (PIF) proteins that are bHLH (basic helix-loop-helix) transcription factors function as major inhibitors of light signaling. In light, photo-activated phytochromes target PIF3 for phosphorylation followed by degradation (Park et al., 2004; Ni et al., 2013). cop1 and pifq mutants exhibit fully expanded open cotyledons in dark indicating that COP1 and the PIF proteins inhibit cotyledon opening in dark. Among the PIFs, PIF1 and PIF3 play a major role in cotyledon closure as the pif1pif3 double mutant displays partially open cotyledons in dark (Shin et al., 2009). Additionally, PIF3 also negatively regulates cotyledon opening under red and far-red light (Kim et al., 2003; Xin et al., 2018). These findings suggest that several members of the light signaling pathway play important roles in regulating cotyledon opening or closure.
Several endogenous hormones interact with the light signaling pathway to regulate the cotyledon opening. The phytohormone BR inhibits cotyledon opening in dark (Wang et al., 2012). BR is synthesized in plants by the catalytic conversion of its sterol precursor campesterol (Gachotte et al., 1996; Kauschmann et al., 1996; Choe et al., 1999a, 1999b, 2000). DWARF4 (DWF4) and DE-ETIOLATED2 (DET2) are the two key BR biosynthetic genes that promote the conversion of campesterol to brassinolide (BL), the most active BR in plants (Li et al., 1996; Choe et al., 1998). DWF4 codes for an enzyme that catalyzes the rate-limiting C-22 hydroxylation step in the BR biosynthetic pathway (Choe et al., 1998; Asami et al., 2001). DET2 encodes a steroid reductase regulating BR biosynthesis downstream of DWF4 (Rozhon et al., 2014). The BR-deficient dwf4 and det2 mutants exhibit short hypocotyl and open cotyledons in dark that can be rescued by the addition of BL (Li et al., 1996; Fujioka and Sakurai, 1997; Azpiroz et al., 1998). Treatment with the BR biosynthetic inhibitor brassinazole (BRZ) that inhibits the activity of DWF4 induces cotyledon opening in Col-0 seedlings in dark. Post synthesis, BR signaling is mediated by several regulators including the BR receptor BRASSINOSTEROID INSENSITIVE 1 (BRI1), the kinase BIN2, and the transcription factors BRASSINAZOLE RESISTANT1 (BZR1) and BES1/BZR2 (BRI1 EMS SUPPRESSOR1; Wang et al., 2001, 2002; He et al., 2002; Yin et al., 2002; Peng et al., 2008; Zhu et al., 2013). While BRI1, BZR1, and BES1 promote BR signaling, BIN2 acts as a negative regulator of the pathway. BIN2 phosphorylates both BZR1 and BES1 and inactivates them in light or BR-deficient conditions (He et al., 2002, 2019; Gampala et al., 2007). The loss and gain of function mutants of these positive and negative regulators, respectively, exhibit BR insensitivity (Chory et al., 1991; Choe et al., 1998; Noguchi et al., 1999; Li et al., 2001). It is interesting to note that many BR-insensitive mutants like bri1-5, bin2-1 exhibit open cotyledons in dark (Li and He, 2016). The dominant mutant bzr1-1D shows closed cotyledons in dark similar to Col-0. The bzr1-1D seedlings are insensitive to BRZ (Li and He, 2016). This suggests that BZR1 promotes cotyledon closure in dark.
There are several evidences of a light signaling factor interacting with BR signaling component and modulating its activity. The photoreceptors PHYTOCHROME B (PHYB), CRYPTOCHROME 2 (CRY1), and UVR8 interact with BES1 and/or BZR1 to reduce their DNA-binding activity thereby repressing BR signaling (Liang et al., 2018; Wang et al., 2018; He et al., 2019; Wu et al., 2019). The B-box containing protein BBX20/BZS1 (B-BOX DOMAIN PROTEIN 20) identified as a suppressor of BZR1 is a positive regulator of photomorphogenesis and negatively regulates BR signaling (Fan et al., 2012). On the other hand, COP1, a key negative regulator of photomorphogenesis increases the ratio of active to inactive form of BZR1 by degrading the inactive form and thereby promotes BR signaling (Kim et al., 2014). COP1 also inhibits the activity of BIN2, the negative regulator of BR signaling to stabilize PIF3 and promote dark-dependent development (Ling et al., 2017). PIF4 interacts with BZR1 and BES1 to co-regulate downstream genes promoting elongation of the hypocotyl (Oh et al., 2012; Martinez et al., 2018). The hy5 mutant is insensitive to the application of BRZ in dark, suggesting that HY5 plays a role in BR-mediated cotyledon closure in dark (Li and He, 2016). HY5 interacts with the dephosphorylated active form of BZR1 and reduces the BZR1-mediated transcriptional regulation of cotyledon closure genes (Li and He, 2016).
The key transcription factors involved in light and BR signaling pathways, HY5, PIFs, and BZR1 regulate several downstream genes to mediate cell expansion and organ movements. Several genes involved in auxin transport or signaling like LAX3 (LIKE AUX1 3), WAG2 (WAVY ROOT GROWTH2), PIN1 (PIN FORMED 1), and ACS5 (ACC SYNTHASE 5) are induced by BZR1 and inhibit the opening of cotyledons (Li and He, 2016). HY5 sequesters BZR1 and suppresses its binding to the promoters of LAX3, WAG2, PIN1, and ACS5 thereby promoting photomorphogenesis (Li and He, 2016). Organ-specific transcriptomic studies have identified some SMALL AUXIN UPREGULATED RNA (SAUR) genes that are induced by light specifically in cotyledons (Sun et al., 2016). Some of these genes like SAUR14, SAUR16, and SAUR50 promote cotyledon opening in light and are repressed by PIF3 in dark (Dong et al., 2019). In dark, PIF3 binds to the promoters of these genes and inhibits the binding of the activator TEOSINTE BRANCHED1/CYCLOIDEA/PROLIFERATING CELL FACTOR 4 (TCP4) to restrict cotyledon opening in darkness (Dong et al., 2019).
BBX32 is a member of the structural group V of the BBX family of proteins in Arabidopsis (Arabidopsis thaliana) containing a single B-box in its N-terminal end (Gangappa and Botto, 2014). BBX32 was initially characterized as a negative regulator of photomorphogenesis based on its role in promoting hypocotyl elongation (Holtan et al., 2011). The same study also indicated that BBX32 negatively regulates the accumulation of the pigment anthocyanin (Holtan et al., 2011). The overexpression of AtBBX32 in soybean resulted in an increase in seed yield probably owing to an increase in the duration of pod and seed development (Preuss et al., 2012). In Arabidopsis, BBX32 acts together with B-BOX DOMAIN PROTEIN 4 (BBX4) to delay flowering (Tripathi et al., 2017). A recent study indicated that BBX32 positively regulates primary root growth under phosphate-deficient conditions (Yeh et al., 2019).
Here, we report that BBX32 is expressed in cotyledons and integrates light and BR signaling pathways to inhibit cotyledon opening. BBX32 is induced by BR and physically interacts with BZR1. We further show that PIF3 interacts with BBX32 and participates in the BR signaling pathway to suppress cotyledon opening in dark. BBX32 integrates light and BR signaling pathways to regulate genes involved in cotyledon opening/closing during the transition from dark to light.
Results
BBX32 is expressed in cotyledons during early seedling development
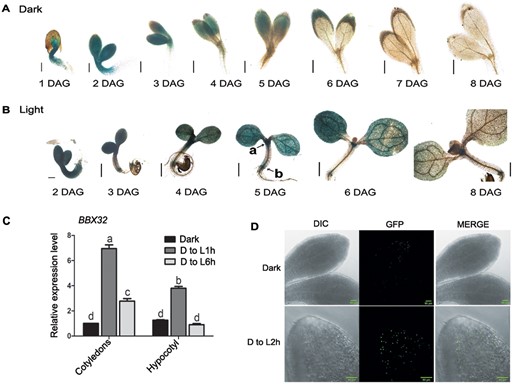
The B-box protein BBX32 is a negative regulator of photomorphogenesis (Holtan et al., 2011). The function of BBX32 during photomorphogenesis is limited to its role in promoting hypocotyl elongation (Holtan et al., 2011). However, organ-specific transcriptomic data have revealed that BBX32 is highly expressed in the cotyledons of seedlings both in dark and during early de-etiolation (Sun et al., 2016). We asked the question if BBX32 plays any role in cotyledon development during de-etiolation. We initially performed histochemical staining and examined the expression of BBX32 both in dark and white light (long day, 16 h/8 h) using the transgenic line ProBBX32:GUS (Figure 1, A and B). BBX32 promoter is transcriptionally active in cotyledons as indicated by enhanced GUS staining in the cotyledons of seedlings grown in dark and light (Figure 1, A and B). At 2 d after germination (DAG) stage, strong GUS expression is detectable in cotyledons and the hypocotyl (Figure 1, A and B). During the period 3–6 DAG, the BBX32 expression reduces in the hypocotyl, being increasingly restricted only to the SAM and the root–hypocotyl junction (Figure 1, A and B). At 6–8 DAG, the intensity of GUS expression reduces substantially even in cotyledons (Figure 1, A and B). When grown under dark or light for 5 d, GUS expression is clearly visible in the cotyledons (Supplemental Figure S1, A and B). GUS staining is also detected at the SAM region and near the root–hypocotyl junction of light-grown seedlings (Supplemental Figure S1B). We could not detect GUS staining in other parts of the hypocotyl or the root of dark or light-grown seedlings at 5 DAG (Supplemental Figure S1, A and B). Next, we quantified the relative expression level of BBX32 in cotyledons and hypocotyls of seedlings grown in dark for 3 d and then shifted to light for 1 h (D to L1h) and 6 h (D to L6h; Figure 1C). In comparison to levels in dark, the transcript levels of BBX32 enhance by seven folds in the cotyledons and four folds in the hypocotyl, after 1 h of light exposure (Figure 1C). The BBX32 expression level in cotyledons lowers down to three folds when seedlings are exposed to longer durations of light (6 h; Figure 1C). After 6 h of light exposure, BBX32 expression in the hypocotyl is comparable to the levels in dark (Figure 1C). We also examined the subcellular localization of BBX32 protein in the cotyledons of 35S:BBX32-GFP seedlings grown in dark and transferred to light (Figure 1D). In dark-grown seedlings, BBX32-GFP accumulates in the nuclei of the cells in the cotyledon (Figure 1D). Nuclear accumulation of BBX32-GFP increases when seedlings are exposed to 2 h of light after the dark period (Figure 1D). Taken together, our results indicate that BBX32 encodes a nuclear protein expressed in the cotyledons of seedlings during early seedling development, especially during the transition from dark to light.
Spatio-temporal expression of BBX32 in seedlings grown in dark, light, and upon transfer from dark to light. A, B, Tissue-specific transcriptional expression pattern of BBX32 in initial days of seedling development from Day 1 to Day 8 after germination using ProBBX32:GUS line in Col-0 background. The seedlings were grown in dark (A) and under white light at the fluence of 80 μmol m−2 s−1 (long day, 16 h/8 h) (B). a and b denote SAM and root–shoot junction, respectively. Scale bar indicates 200 μm. C, Relative expression level of BBX32 in cotyledons and hypocotyl of seedlings grown in dark (D) for 3 d and then shifted to light (L; 80 μmol m−2 s−1) for 1 h (D to L 1 h) and 6 h (D to L6h). n = 2, error bar indicates sem and letters denote statistical groups (P < 0.01) as determined by two-way ANOVA followed by Tukey’s post hoc test. UBQ10 was used for normalization. D, Sub-cellular localization of BBX32 in cotyledons of 35S:BBX32-GFP seedlings. The seedlings were grown for 4 d in dark and shifted to white light of 180 μmol m−2 s−1 fluence for 2 h (D to L 2 h). Scale bar denotes 50 μm.
BBX32 inhibits light-mediated cotyledon opening
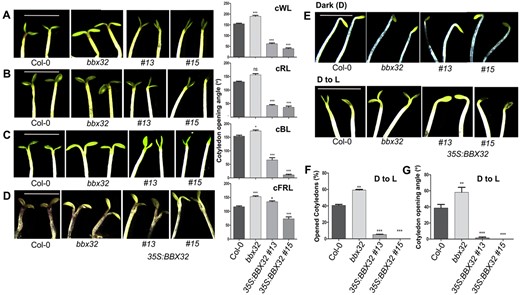
In order to investigate if the high expression levels of BBX32 in cotyledons have any functional significance, we decided to study cotyledon opening during de-etiolation in the loss-of and gain-of-function mutants of BBX32. We used the T-DNA insertion mutant bbx32 that has been characterized before and does not produce any detectable full-length BBX32 transcript (Holtan et al., 2011). However, some partial transcript may be produced in this mutant as reverse transcription quantitative polymerase chain reaction (RT-qPCR) primers amplifying the B-box region (BBX32 qFP and BBX32 qRP) showed 30% mRNA expression compared to the wild-type (WT; Supplemental Figure S2A; Table 1). We also generated several independent transgenic lines overexpressing BBX32. Of these, we chose two lines 35S:BBX32 #13 and 35S:BBX32 #15 that exhibit enhanced levels of BBX32 transcripts for further analysis (Supplemental Figure S2A). These lines also show elongated hypocotyl under white light as reported for BBX32 overexpressing lines in the previous study (Supplemental Figure S2, B and D; Holtan et al., 2011). We grew Col-0, bbx32, 35S:BBX32 #13, and 35S:BBX32 #15 seedlings under low fluence (4 μmol m−2 s−1) constant white, blue, red and 2 μmol m−2 s−1 far-red light and observed the cotyledon opening response. Cotyledons of bbx32 seedlings opened wider than Col-0 under constant white, blue, and far-red light (Figure 2, A, C, and D), while under red light the increase in the opening angle was non-significant (Figure 2B). 35S:BBX32 #13 and 35S:BBX32 #15 seedlings showed a severe reduction in the opening angle compared to the WT in all the light conditions tested except for 35S:BBX32 #13 under far-red light (Figure 2, A–D). When grown under 80 μmol m−2 s−1 cycling white light (16 h/8 h) for 4 d, bbx32 did not exhibit a significant difference in the cotyledon opening angle compared to Col-0 (Supplemental Figure S2, B and C). However, 35S:BBX32 #13 and 35S:BBX32 #15 seedlings showed a reduction in the opening angles (Supplemental Figure S2, B and C).
BBX32 inhibits light-mediated cotyledon opening. A–D, Representative images of cotyledon opening response and the corresponding cotyledon opening angles of Col-0, bbx32, 35S:BBX32 #13, and 35S:BBX32 #15 seedlings grown under low fluence continuous white (cWL), red (cRL), blue (cBL), far-red light (cFRL) respectively. The seedlings were grown in 4 μmol m−2 s−1 cWL for 4 d, 4 μmol m−2 s−1 cRL for 4 d, 4 μmol m−2 s−1 cBL for 6 d, and 2 μmol m−2 s−1 cFRL for 5 d. E, Representative images of cotyledon opening phenotype of the indicated genotypes grown in dark (D) and moved from dark to light (D to L) condition. The seedlings were grown in dark for 4 d and shifted to light (45 μmol m−2 s−1) for 8 h before capturing the images. A–E, Representative seedling images of each genotype were spliced together to prepare the composite image for comparison purpose. F, G, Graphical representation of percentage (%) of open cotyledons and the angle of cotyledon separation in indicated genotypes grown in dark for 4 d and shifted to light corresponding to (E). The scale bar represents 2.5 mm, n = 3. The error bars represent sem. Asterisks represent statistically significant differences (***P < 0.001, **P < 0.01, *P < 0.05) as determined by one-way ANOVA followed by Dunnett’s test; ns, non-significant.
Further, we investigated the cotyledon opening response in these lines in dark and during the transition from dark to light. In dark, seedlings of all the genotypes, Col-0, bbx32, 35S:BBX32 #13, and 35S:BBX32 #15, showed closed cotyledons (Figure 2E). Next, we shifted the seedlings grown in dark for 4 d to continuous white light for 8 h (45 μmol m−2 s−1). Seedlings in which the cotyledons started separating were scored as open and the percentage of seedlings with open cotyledons was determined (Figure 2F). Further, we calculated the cotyledon opening angle by taking an average of the angle of separation between the cotyledons of all the seedlings of a particular genotype after dark to light transition (Figure 2G). In our study, within 8 h 40% of Col-0 seedlings had open cotyledons, whereas 60% of the bbx32 seedlings showed open cotyledons (Figure 2F). While the angle between the cotyledons in the Col-0 seedlings was ∼40°, bbx32 seedlings showed an enhanced opening angle close to 60° (Figure 2G). 35S:BBX32 #13 and 35S:BBX32 #15 seedlings failed to open up the cotyledons (Figure 2, F and G). These results together suggest that BBX32 negatively regulates the light-mediated cotyledon opening.
BBX32 promotes BR-mediated cotyledon closure in dark
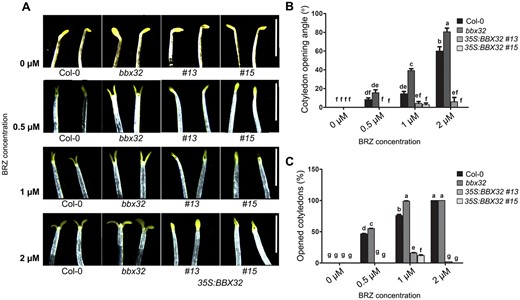
Light and the phytohormone BR antagonistically regulate photomorphogenesis (Wang et al., 2012). BR inhibits cotyledon opening in dark to promote skotomorphogenesis (Li and He, 2016). In a previous chromatin immunoprecipitation-based study, the transcription factor BZR1 was found to bind to the promoter of BBX32 suggesting that BBX32 might play a role in BR signaling (Sun et al., 2010). We, therefore, asked the question if BBX32 plays any role in BR-mediated cotyledon closure in dark. The addition of the BR biosynthetic inhibitor BRZ can induce cotyledon opening in dark in WT seedlings (Wang et al., 2012; Li and He, 2016). We hypothesized that if BBX32 is involved in the BR signaling downstream of BR synthesis, the plants overexpressing BBX32 might be insensitive to the BR deficiency induced by BRZ and exhibit closed cotyledons in dark. The bbx32 mutant might exhibit the opposite phenotype of enhanced cotyledon opening under similar conditions. In order to test our hypothesis, we examined the sensitivity of dark-grown Col-0, bbx32, 35S:BBX32 #13, and 35S:BBX32 #15 seedlings to different concentrations of BRZ (0.5 µM, 1 µM, 2 µM). We found that bbx32 is hypersensitive to BRZ and the overexpressor (OE) lines 35S:BBX32 #13 and 35S:BBX32 #15 are highly insensitive to BRZ-induced cotyledon opening response in darkness (Figure 3, A–C). In the absence of BRZ, seedlings of all genotypes showed closed cotyledons (Figure 3A). With the increase in concentrations of BRZ (0.5–2 µM), the number (%) of seedlings with open cotyledons and the angle of separation between the cotyledons increased in Col-0 (Figure 3, A–C). Under similar conditions, the percentage of seedlings with open cotyledons and the cotyledon opening angle were higher in bbx32 and lower in 35S:BBX32 #13 and 35S:BBX32 #15 compared to Col-0 (Figure 3, A–C), suggesting that BBX32 positively regulates BR-mediated cotyledon closure in dark. The hypocotyl lengths of the loss-of and gain-of-function mutants of BBX32 did not differ significantly from Col-0 after BRZ (1–2 µM) treatment but showed a small difference at lower concentration (Supplemental Figure S3).
35S:BBX32 seedlings are insensitive to BR-deficiency-induced cotyledon opening in dark. A, Representative images depicting the cotyledon opening response of Col-0, bbx32, 35S:BBX32 #13, and 35S:BBX32 #15 to BRZ treatment in dark. The seedlings were grown on MS plates with 1.5% sucrose containing different concentrations (0.5 µM, 1 µM, 2 µM) of BRZ in dark for 7 d. The plate without BRZ was used as control. The scale bar represents 2.5 mm. Representative seedling images of each genotype were spliced together to prepare the composite image for comparison purpose. B, C, Cotyledon opening angle and percentage of open cotyledons of indicated genotypes treated with BRZ at various concentrations, respectively. The data represent mean of 30 seedlings. Error bar = sem. The statistical groups denoted by letters were determined by using two-way ANOVA followed by Tukey’s post hoc test (P < 0.05).
BR induces BBX32 expression and BBX32 interacts with BZR1
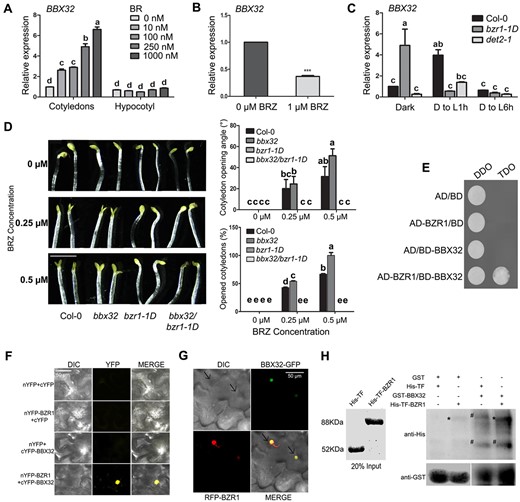
Since BBX32 plays a positive role in BR-mediated response, we investigated its role in BR synthesis and/or signaling. We initially examined if exogenous application of BR regulates the transcript levels of BBX32. Dark-grown Col-0 seedlings treated with increasing concentrations of the BR BL (Brassinolide; 0.1–1 μM) showed a consistent increase in the relative expression levels of BBX32 in cotyledons but not in the hypocotyl (Figure 4A). This suggests that BR positively regulates the BBX32 expression specifically in cotyledons. We also detected a reduction in the transcript levels of BBX32 in the cotyledons of BRZ-treated dark-grown Col-0 seedlings (Figure 4B). Additionally, transgenic lines expressing ProBBX32:GUS grown in dark in the presence of BR or BRZ showed enhanced and reduced GUS staining respectively further validating the induction of the BBX32 expression by BR (Supplemental Figure S4, A and B). Additionally, we found that the expression of BBX32 is significantly reduced in the BR biosynthesis mutant det2-1 both in dark and after transfer to light, suggesting BR-mediated regulation of the BBX32 expression in vivo (Figure 4C).
BR induces BBX32 expression and BBX32 genetically and physically interacts with BZR1. A, B, Relative expression level of BBX32 in the cotyledons of Col-0 seedlings treated with BR and BRZ, respectively. A, RT-qPCR data show the expression of BBX32 in cotyledons and hypocotyls of 5-d-old dark-grown Col-0 seedlings grown in the absence or the presence of BL (0 nM, 10 nM, 100 nM, 250 nM, and 1000 nM). B, RT-qPCR data of the BBX32 expression in the cotyledons of 5-d-old dark-grown Col-0 seedlings treated with BRZ (1 µM) compared to untreated Col-0 seedlings, n = 2. Error bar represents sem, UBQ10 was used for normalization. Student’s t test was performed to calculate the significance denoted by asterisks (***P <0.0001). C, Relative expression level of BBX32 in the cotyledons of bzr1-1D and det2-1 compared to Col-0. The seedlings were grown in dark for 5 d and shifted to 1-h and 6-h light (D to L1h and D to L6h; 80 μmol m−2 s−1) before harvesting the cotyledons. In (A) and (C), n = 2, error bar represents sem, UBQ10 was used for normalization. Two-way ANOVA followed by Tukey’s post hoc test was performed to determine the statistical groups designated by letters (P <0.01). D, Representative seedling images and graphical representation of the cotyledon opening angle and opened cotyledon percentage of indicated genotypes grown in different concentrations of BRZ. The seedlings were grown for 7 d in dark with (0.25 µM and 0.5 µM) or without BRZ. The scale bar measures 2.5 mm. The data in the graphs are the mean of 30 seedlings. Error bar = SEM, two-way ANOVA followed by Tukey’s post hoc test was performed to determine the statistical groups (P <0.05) denoted by letters. Representative seedling images of each genotype were spliced together to prepare the composite image for comparison purpose. E, Yeast two-hybrid assay showing BBX32–BZR1 interaction, AD and BD refer to the activation domain and binding domain of GAL4 containing pGADT7 and pGBKT7 vectors, respectively. DDO indicates the double-dropout medium lacking leucine and tryptophan. TDO indicates the triple-dropout medium lacking leucine, tryptophan, and histidine. F, BiFC analysis showing the interaction of BBX32 with BZR1 in planta, nYFP, and cYFP refer to the N-terminal and C-terminal of yellow fluorescent protein, respectively. Vectors in the indicated combinations were transformed into Agrobacterium and co-infiltrated into 3-week-old N. benthamiana leaves and fluorescent images were captured after 36 h. Scale bar represents 50 µm. G, Colocalization of BBX32 and BZR1 in the nucleus of N. benthamiana leaf epidermal cells. BBX32 is fused with green fluorescent protein (GFP) at C terminus and BZR1 is fused with red fluorescent protein (RFP) at N-terminus and are driven by 35S promoter. Both the constructs were transformed into Agrobacterium and co-infiltrated into 3-week-old leaves and images were captured after 2 d. The scale bar measures 50 µm. H, In vitro pull-down assay showing the interaction between BBX32 and BZR1. BBX32 fused to GST and BZR1 fused to His-TF were purified along with empty tags. His-TF or His-TF-BZR1 were used as prey proteins and incubated with GST (29 kDa) or GST-BBX32 (50 kDa). Anti-GST and anti-His antibodies were used to detect the respective proteins. Input is 20% of the purified proteins used and is shown with CBB stain. Asterik denotes His-TF-BZR1 band, hash indicates non-specific bands.
BZR1, the key transcription factor in the BR signaling pathway, plays an important role in BR-mediated inhibition of the cotyledon opening in dark (Li and He, 2016). We asked the question if BZR1 and BBX32 regulate the dynamics of the cotyledon opening in dark through a common pathway. Since BZR1 is known to bind to the promoter of BBX32, we first examined if BZR1 regulates the transcription of BBX32 by comparing the mRNA levels of BBX32 between Col-0 and the dominant mutant bzr1-1D grown in dark (Sun et al., 2010). Our results indicate that BBX32 transcript levels are highly upregulated in the cotyledons of dark-grown bzr1-1D seedlings compared to the WT (Figure 4C; Supplemental Figure S4C). This induction was, however, absent when the seedlings were transferred to light for 1 h or 6 h (Figure 4C). Also, we could not detect a significant difference in the BBX32 expression levels in the hypocotyls of Col-0 and bzr1-1D seedlings grown in dark (Supplemental Figure S4C). These results suggest that BZR1 regulates the BBX32 expression in the cotyledons of seedlings grown in dark. In order to understand the physiological significance of this regulation we examined the cotyledon opening response of dark-grown Col-0, bbx32, bzr1-1D, and bbx32bzr1-1D seedlings upon the application of BRZ (Figure 4D). While bbx32 exhibits enhanced cotyledon opening upon BRZ treatment, bzr1-1D and bbx32bzr1-1D are resistant to BRZ perhaps due to the stabilization of the BRZ-resistant BZR1 protein by the dominant bzr1-1D mutation (Figure 4D). This suggests that BBX32 and BZR1 genetically interact to regulate BR-mediated inhibition of cotyledon opening in dark.
We further examined the possibility of a physical interaction between BZR1 and BBX32. Our yeast two-hybrid assay indicated the presence of interaction between BZR1 and BBX32 (Figure 4E). We could not detect the interaction between BES1 and BBX32 in yeast (Supplemental Figure S4D). In order to the check the interaction between BZR1 and BBX32 in planta, we co-transformed Nicotiana benthamiana leaves with nYFP-BZR1 and cYFP-BBX32 along with the controls. YFP signal was detected only when nYFP-BZR1 and cYFP-BBX32 were transfected together indicating that the two proteins physically interact in planta (Figure 4F). We could also detect the colocalization of BBX32-GFP and RFP-BZR1 in the nucleus of N. benthamiana leaf epidermal cells (Figure 4G). Our in vitro pull-down assay showed an interaction between GST-BBX32 and His-TF-BZR1 (Figure 4H). All these in vitro and in vivo interaction studies suggest that BBX32 physically interacts with BZR1.
PIF3 interacts with BBX32 and promotes BR-mediated cotyledon closure in dark
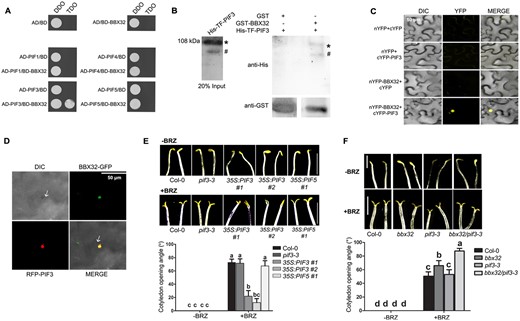
Cotyledon opening is also regulated by the PIF proteins. The quadruple mutant pifq exhibits expanded open cotyledons in dark (Shin et al., 2009). To determine if BBX32 interacts with the PIF proteins (PIF1, PIF3, PIF4, and PIF5) we performed a yeast-two hybrid assay. We found that BBX32 physically interacts with PIF3 but not with any of the other PIF proteins (Figure 5A). In our in vitro pull-down experiment, BBX32 showed an interaction with PIF3 (Figure 5B). To check the interaction between BBX32 and PIF3 in planta, we performed bimolecular fluorescence complementation (BiFC) and colocalization experiments. YFP signal was detected in leaves infiltrated with nYFP-BBX32 and cYFP-PIF3 together suggesting interaction between BBX32 and PIF3 (Figure 5C). We also observed that BBX32-GFP and RFP-PIF3 colocalize together in the nucleus of N. benthamiana leaf epidermal cells (Figure 5D). The role of PIF3 in negatively regulating light-mediated cotyledon opening is well established (Kim et al., 2003; Shin et al., 2009; Xin et al., 2018). We generated two PIF3 OE lines—35S:PIF3 #1, 35S:PIF3 #2—with high levels of PIF3 expression and confirmed their photomorphogenic phenotypes (hypocotyl length and cotyledon opening) together with pif3-3 and Col-0 (Supplemental Figure S5, A–D). However, whether PIF3 plays any role in the BR-mediated regulation of cotyledon closure in dark is not known. To investigate this, we grew Col-0, pif3-3, 35S:PIF3 #1, 35S:PIF3 #2 seedlings in dark in the presence of BRZ (2 µM). While BRZ induces cotyledon opening in Col-0 and pif3-3, the PIF3 overexpression lines were partially insensitive to BRZ-mediated cotyledon opening in dark (Figure 5E). We did not find a significant difference between the opening angle of Col-0 and pif3-3 (Figure 5E). Upon treating with BRZ, 35S:PIF5 seedlings showed open cotyledons similar to Col-0 suggesting that different PIF proteins exhibit varied responses to BRZ (Figure 5E). Our results indicate that BBX32 and PIF3 physically interact with each other and play roles in the BR-mediated cotyledon closure in dark. In order to understand the physiological significance of the interaction in inhibiting the cotyledon opening in dark, we generated the double mutant and examined the cotyledon opening response of dark-grown Col-0, bbx32, pif3-3, and bbx32pif3-3 seedlings upon the application of BRZ at various concentrations (Figure 5F; Supplemental Figure S5E). Although pif3-3 did not show much difference from Col-0 in the cotyledon opening angle, the loss of PIF3 enhances the cotyledon opening angle of bbx32 suggesting that the two proteins might together regulate some common targets involved in the cotyledon opening (Figure 5F; Supplemental Figure S5E). Taken together our results suggest that PIF3 interacts with BBX32 and is involved in the BR-mediated cotyledon closure in dark.
PIF3 interacts with BBX32 and 35S:PIF3 is hyposensitive to BRZ-induced cotyledon opening in dark. A, Yeast two-hybrid assay showing the interaction of BBX32 with PIF3. The PIF genes PIF1, PIF3, PIF4, and PIF5 are in AD—activation domain containing vector, and BBX32 is in BD—binding domain containing vector. DDO indicates double-dropout medium without leucine and tryptophan. TDO indicates triple-dropout medium without leucine, tryptophan, and histidine. B, In vitro pull-down assay showing the interaction between BBX32 and PIF3. BBX32 fused to GST and PIF3 fused to His-TF were purified along with empty GST. His-TF-PIF3 was used as prey protein and incubated with GST (29 kDa) or GST-BBX32 (50 kDa). Input is 20% of the purified prey protein used in the assay. Anti-GST and anti-His antibodies were used to detect the respective proteins. Asterisk denotes His-TF-PIF3 band, hash indicates additional non-specific band. C, BiFC assay showing the interaction between BBX32 and PIF3, Agrobacterium transformed with N-terminal YFP fused BBX32 and the C-terminal YFP fused PIF3 were infiltrated into N. benthamiana leaves along with empty vector controls in mentioned combinations. The infiltrated plants were incubated in dark for 36 h before capturing the images under confocal microscope. Scale bar indicates 50 µm. D, Colocalization of BBX32 and PIF3 in the nucleus of N. benthamiana leaf epidermal cells. BBX32-GFP and RFP-PIF3 constructs were transformed into Agrobacterium and infiltrated together into 3-week-old leaves and after 2 d, images were captured. The scale bar measures 50 µm. E, Representative images demonstrating cotyledon opening response and the corresponding cotyledon opening angle of dark-grown seedlings of Col-0, pif3-3, 35S:PIF3 #1, 35S:PIF3 #2, and 35S:PIF5 #1 upon BRZ treatment. The seedlings were grown on MS plates with 1.5% sucrose at 2 µM concentrations of BRZ in dark for 7 d (+BRZ). The plate without BRZ was used as control (−BRZ). Scale bar = 2.5 mm. F, Representative images of seedlings of indicated genotypes grown in dark with or without BRZ and their corresponding cotyledon opening angles. The seedlings of Col-0, bbx32, pif3-3, and bbx32pif3-3 were grown in dark with 0 µM (−BRZ) and 2 µM of brassinazole (+BRZ) for 7 d. E, F, Representative seedling images of each genotype were spliced together to prepare the composite image for comparison purpose. In (E) and (F) the measures of cotyledon opening angles given are the mean of 30 seedlings. Error bar = sem. Letters denote the statistical groups obtained using two-way ANOVA and Tukey’s post hoc test (P <0.05).
Since PIF3 and BZR1, the two interacting partners of BBX32, are transcription factors and BBX proteins are also involved in transcriptional regulation, we asked the question if these proteins can modulate the transcriptional activation potential of each other. We co-transfected Arabidopsis protoplasts with the promoter of the anthocyanin biosynthesis gene CHS fused to luciferase (ProCHS:LUC) as the reporter together with 35S:BBX32, 35S:PIF3, and 35S:BZR1 as effectors in different combinations (Supplemental Figure S5F). CHS has been previously shown to be a target of factors involved in light and BR signaling (Chory et al., 1991; Ang et al., 1998). Relative luciferase activity measurements indicated that BBX32 represses while BZR1 and PIF3 promote the transcription of CHS (Supplemental Figure S5F). Interestingly, the transcriptional activation potential of BZR1 and PIF3 is repressed by BBX32 suggesting role of BBX32 in transcriptional modulation (Supplemental Figure S5F).
BBX32 regulates genes promoting cotyledon closure and opening in opposite ways
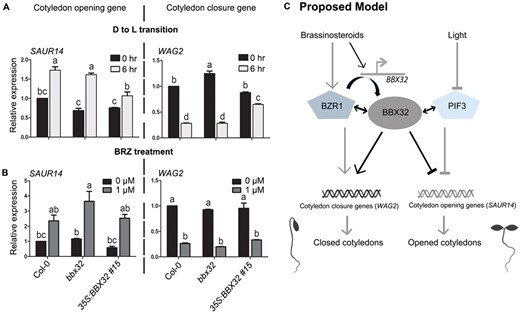
Organ-specific transcriptomic studies have identified a number of SAUR genes that promote cotyledon opening and are repressed by the PIFs in dark (Sun et al., 2016). Among these targets, PIF3 strongly inhibits the activation of SAUR14 in the cotyledons of dark-grown seedlings (Dong et al., 2019). BZR1 inhibits cotyledon opening in dark by regulating several genes involved in auxin transport. One of its targets is the serine–threonine kinase WAG2 that mediates auxin transport and acts as a negative regulator of cotyledon opening (Li and He, 2016). Since BBX32 inhibits cotyledon opening in dark similar to BZR1 and PIF3, and also interacts with the two proteins, we asked the question if these proteins target similar genes to regulate the cotyledon opening and closure. To determine this, we examined the expression levels of SAUR14 (gene promoting cotyledon opening) and WAG2 (gene inhibiting cotyledon opening) in the cotyledons of Col-0, bbx32, and 35S:BBX32 #15 seedlings grown in dark and transferred to light (Figure 6A). In WT seedlings, SAUR14 expression levels were upregulated upon exposure to light, whereas the WAG2 transcript levels were repressed in light (Figure 6A). We did not see significant difference in the expression of SAUR14 and WAG2 in Col-0 and bbx32 exposed to light for 6 h (Figure 6A). This might be because the regulation of these genes might partially depend on the interaction of BBX32 with its partners PIF3 and BZR1 in darkness. However, in 35S:BBX32 #15 seedlings exposed to light, the expression of SAUR14 was reduced and WAG2 was enhanced compared to Col-0, correlating well with its closed cotyledon phenotype in light (Figures 6A and 2, E–G). Similar regulation was found in the light-exposed 35S:BBX32 #15 seedlings for few more genes promoting cotyledon opening (SAUR16, SAUR50) and genes inhibiting cotyledon opening (LAX3, ACS5; Supplemental Figure S6). Further, we quantified the expression levels of SAUR14 and WAG2 in the seedlings of Col-0, bbx32, and 35S:BBX32 #15 grown in dark in the presence or the absence of BRZ (Figure 6B). In the BRZ replete condition, SAUR14 was upregulated and WAG2 was downregulated in Col-0 compared to no-BRZ condition, in agreement with BRZ promoting cotyledon opening in dark (Figure 6B). The expression level of SAUR14 was higher in bbx32 compared to Col-0 under BRZ (Figure 6B). However, the expression of WAG2 did not show a significant difference from Col-0 in the presence of BRZ (Figure 6B). In 35S:BBX32 #15, the expression levels of both these genes were similar to the WT in the presence and the absence of BRZ suggesting that the BRZ-resistant phenotype regulated by BBX32 might involve the regulation of some other target genes (Figures 6B and 3, A–C). Together these data indicate that BBX32 regulates the expression of the targets of BZR1 and PIF3 to modulate the BR-dependent cotyledon closure in dark and opening of cotyledons in light (Figure 6C).
BBX32 regulates the expression of genes involved in opening and closing of cotyledons. A, B, RT-qPCR data showing relative expression levels of the genes regulating cotyledon opening during dark to light (D to L) transition (A) or upon BRZ treatment (B). A, The expression levels of indicated genes in the cotyledons of Col-0, bbx32, and 35S:BBX32 #15 seedlings grown in dark for 3 d and shifted to (80 µmol m−2 s−1) white light for 6 h before harvesting the samples. SAUR14 promotes cotyledon opening and WAG2 inhibits cotyledon opening. B, The relative transcript levels of SAUR14 and WAG2 in detached cotyledons of 5-d-old bbx32 and 35S:BBX32 #15 compared to Col-0 grown in darkness upon 0 µM or 1 µM BRZ treatment. In both (A) and (B), n = 2, error bar = sem. Letters denote the statistically significant groups determined by two-way ANOVA followed by Tukey’s post hoc test (P <0.01). UBQ10 was used for normalization. C, The proposed model to explain the molecular mechanism of BBX32 integrating light and BR signaling pathways to regulate the opening and closing of cotyledons. In dark, BR activates the BBX32 expression in cotyledons. BZR1 can modulate the BBX32 expression and BBX32 physically interacts with BZR1. BBX32 and BZR1 regulate common target genes that promote cotyledon closure like WAG2. BBX32 also interacts with PIF3 and in darkness the two proteins co-repress genes like SAUR14 that promote cotyledon unfolding. Light inhibits PIF3 to activate cotyledon opening genes. The black lines indicate connections identified in this study, whereas the grey lines represent previously published pathways. Here, the lines with arrowheads indicate activation, while lines with inverted T represent inhibition. Double-headed arrows indicate interaction between proteins.
Discussion
Integration of light and BR signals to regulate cotyledon opening
Antagonistic interplay between light and BR modulates the dynamics of the cotyledon development during de-etiolation (Li and He, 2016). While light promotes the opening of cotyledons, BR inhibits cotyledon opening in darkness (Li and He, 2016; Sun et al., 2016; Xin et al., 2018; Dong et al., 2019). Interactions between the components of the light and BR signaling pathways regulate several developmental processes in plants like germination, hypocotyl elongation, cotyledon development, and greening (Nolan et al., 2020). A recent study also suggests that light promotes BR synthesis to regulate hook opening and petiole elongation, thereby making the interplay between light and BR even more complex (Hamasaki et al., 2020). Many of the light signaling modulators play crucial roles in BR signaling. The transcription factors BZR1 and BES1 form a crucial node of interaction with photoreceptors like PHYB, CRY1, CRY2, UVR8, and light signaling intermediates like COP1, PIF4, GATA TRANSCRIPTION FACTOR 2 (GATA2), and HY5 (Oh et al., 2012; Li and He, 2016; Liang et al., 2018; Martinez et al., 2018; Wang et al., 2018; He et al., 2019; Wu et al., 2019; Nolan et al., 2020). It is quite interesting to note that the positive regulators of light signaling like the photoreceptors and HY5 inhibit the activity of BZR1/BES1, while the factors promoting skotomorphogenesis like COP1 and PIF4 act in concert with BR signal transduction regulators in dark. COP1 stabilizes BZR1 in dark to promote BR signaling (Kim et al., 2014). COP1 is also known to interact with BIN2, the GSK kinase that negatively regulates BR signaling, and to inhibit the BIN2-mediated phosphorylation of PIF3, thereby stabilizing PIF3 in dark (Ling et al., 2017). Most of the above interactions have studied the role of the integration of the external light and endogenous BR signaling pathway in regulating hypocotyl elongation. The only exception so far being the study of HY5–BZR1 interaction and its role in mediating cotyledon opening in dark (Li and He, 2016). Here, we identify BBX32 interacting with BZR1 and mediating the cross-talk between light and BR signaling pathways to regulate cotyledon dynamics during de-etiolation (Figures 2–4). The only other BBX protein so far identified as a common regulator of light and BR signaling pathways is BBX20/BZS1 (Fan et al., 2012). It is a positive regulator of photomorphogenesis that is transcriptionally repressed by BZR1 (Fan et al., 2012). Our results also indicate that PIF3 plays a positive role in the BR signaling pathway (Figure 5). The PIF3 overexpressing lines were hyposensitive to BRZ-mediated cotyledon opening in dark (Figure 5). Higher order pif mutants exhibit open cotyledons in dark suggesting their role in regulating cotyledon opening/closure. PIF3 mediates cotyledon closure in darkness by inhibiting the binding of TCP4 on to the promoters of SAUR genes that promote cotyledon opening (Dong et al., 2019). Our study suggests the role of PIF3 in BR-mediated cotyedon closure in dark thereby providing more insight into the regulation of cotyledon separation during de-etiolation.
B-box protein BBX32 positively regulates BR signaling to promote cotyledon closure
BBX proteins play roles in the regulation of several processes like photomorphogenesis, flowering, pigment accumulation, shade avoidance, thermomorphogenesis, and UV-B tolerance (Song et al., 2020). Photomorphogenesis includes morphological changes in the growth and development of hypocotyl, cotyledon, and the root during de-etiolation. Traditionally, the regulators of this process have mostly been identified and characterized based on their role in modulating hypocotyl elongation during the transition from dark to light. However, recent studies have started dissecting each of these processes and identifying their specific regulators. Organ-specific transcriptomic studies have identified genes that are induced in cotyledons and repressed in the hypocotyls upon light exposure and contribute to cotyledon expansion and inhibition of hypocotyl elongation in light (Sun et al., 2016). Cotyledon development itself involves a series of steps like the release of the apical hook, opening, expansion, and greening of the cotyledons; each having independent as well as inter-dependent regulatory processes.
BBX32 was initially identified as a negative regulator of photomorphogenesis promoting hypocotyl elongation by inhibiting HY5 activity (Holtan et al., 2011). Later, its role in the regulation of seed yield, flowering, and root development has also been reported (Preuss et al., 2012; Tripathi et al., 2017; Yeh et al., 2019). Our results indicate that BBX32 promotes BR signaling and negatively regulates light-mediated opening of cotyledons during de-etiolation (Figure 2). To our knowledge, this is the first report of a BBX protein integrating light and BR signals to regulate cotyledon opening/closure. Recently, the microproteins BBX30 and BBX31 were found to regulate apical hook opening and cotyledon development by disrupting the oligomerization of PIFs and EIN3 (Wu et al., 2020). BBX32 appears to be a key regulator of plant development as it modulates both early developmental processes like cotyledon opening in seedlings as well as traits of agronomic importance like flowering and seed yield later during the reproductive development of the adult plant. All our experiments were performed on seeds inoculated on plates containing a solidified growth medium. Since in natural conditions seeds are buried under a layer of soil, in the future it might be interesting to simulate the soil cover and investigate the cotyledon opening of BBX32 mutants together with their emergence into light under mechanical pressure.
BBX32 interacts with BZR1 and PIF3, and modulates genes regulating cotyledon separation
The B-box containing BBX proteins are known to mediate protein–protein interactions. BBX proteins were identified in a yeast-two hybrid screen interacting with COP1 (Holm et al., 2001). BBX4 (COL3), BBX24 (Lee et al., 2007), BBX25 (STH) were identified in this screen and their role in photomorphogenesis was further characterized (Holm et al., 2001; Datta et al., 2006; Indorf et al., 2007; Yan et al., 2011; Gangappa et al., 2013). Since then many more BBX proteins CO/BBX1, BBX10, BBX19, BBX20, BBX22, BBX28 have been found to physically interact with COP1 and act as substrates for degradation via the 26S proteasome pathway (Kim et al., 2017). The bZIP transcription factor HY5 regulates the transcription of several thousand genes but lacks transcriptional activation potential by itself both in yeast and in planta (Ang et al., 1998; Lee et al., 2007; Burko et al., 2020). The BBX proteins BBX20, BBX21, BBX22, BBX23 interact with HY5 and promote the transcriptional activation potential of HY5 (Datta et al., 2007, 2008; Wei et al., 2016; Zhang et al., 2017). BBX20, BBX21, and BBX22 proteins act as rate-limiting cofactors of HY5 and positively regulate photomorphogenesis (Bursch et al., 2020). On the other hand, the negative regulators of photomorphogenesis BBX24, BBX25, and BBX28 physically interact with HY5 and inhibit its transcriptional activity (Jiang et al., 2012; Gangappa et al., 2013; Lin et al., 2018). BBX proteins are also known to interact with other BBX factors to form heterodimers. BBX32 interacts with BBX21 and suppresses the activities of BBX21 and HY5 in promoting photomorphogenesis (Holtan et al., 2011). BBX32 forms heterodimer with BBX4 to negatively regulate FT transcription and delay flowering in Arabidopsis (Tripathi et al., 2017). Here, we report that BBX32 physically interacts with PIF3 and BZR1 (Figures 4 and 5). Recently, interaction between another BBX protein BBX4, and PIF3 was shown (Heng et al., 2019). There are no reports of a physical interaction between BZR1 and a BBX protein. However, BZR1 is known to transcriptionally repress BBX20/BZS1, the suppressor of BZR1 (Fan et al., 2012). Our study indicates that BBX32 physically interacts with BZR1 and PIF3, integrating BR and light signaling pathways to regulate the opening of cotyledons. BZR1 and PIF3 have been shown to physically interact to regulate photomorphogenesis (Zhang et al., 2014). We chose genes regulated by BZR1 and PIF3 to modulate cotyledon opening or closure and examined if they are regulated by BBX32 (Li and He, 2016; Dong et al., 2019). Our results indicate that BBX32 promotes the expression of cotyledon closure genes, while it inhibits the transcription of the cotyledon opening SAUR genes in dark (Figure 6). BBX32 is also expressed in the apical hook region in addition to cotyledons during de-etiolation (Supplemental Figure S1A). We saw that in light the expression of BBX32 is reduced in cotyledons when exposed to continuous light or for longer durations after transition from darkness (Figure 1B). However, in light BBX32 is expressed at high levels in the apical hook or SAM region (Supplemental Figure S1). BBX32 negatively regulates the expression of SAUR50 in light that was recently shown to promote the apical hook opening (Figure 6A; Wang et al., 2020). WAG2 represses apical hook opening and is positively regulated by BBX32 (Figure 6A; Willige et al., 2012). This regulation might enable BBX32 to repress cotyledon opening in light. Further, organ-specific expression studies might be required to better resolve the role of BBX32 in apical hook and cotyledon opening in light when BZR1 and PIF3 are unavailable.
Materials and methods
Plant materials and growth conditions
The Arabidopsis (A. thaliana) mutants used in the study are bbx32 (SALK059534), pif3-3 (fast neutron mutagenized; Stephenson et al., 2009), and bzr1-1D (Wang et al., 2002). The transgenic OE lines 35S:BBX32 #13, 35S:BBX32 #15, 35S: BBX32-GFP, 35S:PIF3 #1, 35S:PIF3 #2, 35S:PIF5 and promoter-reporter line ProBBX32:GUS were made using Agrobacterium-mediated floral dip method after cloning the genes in respective vectors using Gateway cloning technology. The entry clones were made in pDONR207; the destination vectors pCAMBIA1300 (35S promoter), pGWB3 (GUS reporter); pGWB6 (35S promoter, N terminal sGFP) were used to generate the lines. All these mutants and transgenic lines are in the Col-0 background. The crosses used in this study include pif3-3bbx32 and bzr1-1Dbbx32. In both bbx32pif3-3 and bzr1-1Dbbx32, bbx32 was used as a male parent. Genotyping or segregation analysis for hygromycin resistance was performed to select homozygous double mutants.
The seeds were sterilized using 5% (v/v) sodium hypochlorite solution for 3 min followed by five washes with autoclaved water. The seeds were grown on Murashige & Skoog (MS) media with 1.5% sucrose for BRZ experiments and 1/2 MS with 1% sucrose for other experiments. BRZ (SIGMA SML1406-5mg) was dissolved in 100% ethanol. After 3 d of stratification at 4°C, the seeds were induced for germination by exposing to white light (80 µmol m−2 s−1) for 6 h. Then the plates were shifted to dark or different monochromatic light conditions according to the experiments. The primer sequences used for genotyping and to generate transgenics are given in Supplemental Table S1.
GUS assay
GUS assay was performed as given in the Arabidopsis Laboratory manual (Cold Spring Harbor Laboratory Press; Weigel and Glazebrook, 2002). The GUS substrate 5-bromo-4-chloro-3-indolyl-β-d-glucuronide (X-Gluc) was provided by MP Biomedicals and dissolved in DMF (Dimethyl formamide). The seedlings were washed in 90% acetone twice. Further, the seedlings were washed with the GUS staining solution (0.5 M sodium phosphate buffer, 4% Triton X-100, 100-mM potassium ferrocyanide, and 100-mM potassium ferricyanide without X-Gluc) twice on ice. The GUS staining solution with GUS substrate was added to the seedlings and vacuum was provided for 30 min. After vacuum infiltration, the seedlings were incubated at 37°C till the blue-colored stain appeared. Once the stain was visible, the seedlings were subjected to destaining. Destaining involved a series of ethanol washes with 20%, 35%, and 50% ethanol for 30 min each at room temperature. Finally, the stain in seedlings was fixed with FAA mixture (10% formaldehyde, 50% ethanol, and 5% acetic acid) and the seedlings were stored in 70% ethanol. The images were taken with a light microscope LEICA DM2500.
Cotyledon angle and hypocotyl length measurements
The cotyledon opening angles were measured as reported before (Li et al., 2014). ImageJ software was used to measure cotyledon angles and hypocotyl length of the seedling images. The mean angle or hypocotyl length of all seedlings of a particular genotype was used to plot the graphs.
Statistical analysis
Graphpad Prism was used to plot graphs and to perform statistical analysis, “n” in the figure legends denote the number of biological replicates. The details of analysis and P-value considered to determine the significance are given in the respective figure legends.
Yeast-two hybrid assay
The protein–protein interaction using yeast two-hybrid assay was performed as explained by Clontech. BBX32 was cloned in pDEST-GBKT7 and BZR1, BES1 (BZR2), PIF1, 3, 4, and 5 were cloned in pDEST-GADT7 vectors. The primer sequences of the cloned genes are given in Supplemental Table S1. The vectors were co-transformed into yeast cells as mentioned in the protocol. The yeast cells were grown overnight and pelleted down at 4,000 rpm for 2 min. The supernatant was discarded and the pellet was washed with sterile H2O. The cells were incubated in LiOAc for 30 min. Then 30 µL cells were added to the transformation buffer (Tris pH 8.0, EDTA pH 8.0, 100-mM LiOAc, and 40% polyethylene glycol [PEG]) and ssDNA (Salmon Sperm) also was added to enhance the transformation efficiency. Finally, the vectors in different combinations including all the necessary controls were added. These cells were incubated at 30°C for 1 h and shifted to 42°C for 25–30 min. The cells were then pelleted down and plated on selection medium. The yeast selection marker for pDEST-GADT7 is leucine and for pDEST-GBKT7 is tryptophan. The entry of both the vectors was confirmed by using double-dropout plates which are lacking both Leucine and Tryptophan. The interaction of two proteins was confirmed by the third marker histidine.
Bi-fluorescence complementation assay
The protocol for BiFC was followed as suggested by Gampala (Gampala et al., 2007). The vectors used in the experiment are pCL112 and pCL113. The pCL112 vector has N-terminal part of YFP and the pCL113 vector has C-terminal part of YFP. The genes of interest were cloned into the pCL vectors and through Agrobacterium-mediated infiltration the cloned vectors were carried into the N. benthamiana plants. BZR1 and PIF3 were cloned into pCL112 and pCL113, respectively. BBX32 was cloned in both of the vectors. The primer sequences are listed in Supplemental Table S1. Agrobacterium cells transformed with these constructs were treated with activation media comprising 10 mM MgCl2, 10 mM MES, and 20 µM acetosyringone. After 36 h to 48 h in dark, the plants were screened for the fluorescence. Zeiss Apotome microscope was used to capture the fluorescence images (Halogen/Arc lamp as light source) and the excitation and emission wavelengths used were 488 nm and 510–525 nm, respectively. PMT voltage (gain) was not exceeded above 750 V.
Sub-cellular localization
The 35S:BBX32-GFP seedlings were grown for 4 d in dark and shifted to 180 µmol m−2 s−1 white light for 2 h before screening for the sub-cellular localization using Olympus live cell microscope. The Ar-ion gas laser was used for excitation at 488 nm to capture GFP signal with an emission wavelength of 520 nm. PMT voltage (gain) was not exceeded above 750 V.
Protoplast isolation, PEG transfection, and luciferase assay
The protoplast isolation and PEG-mediated transformation were carried out as per the procedures provided by Sheen (Sheen, 2002). Col-0 plants grown in short day for 3 weeks were used for protoplast isolation. BBX32, PIF3, BZR1 cloned in pCAMBIA1300 as effector constructs and the ∼2 kb from TSS of CHS promoter cloned in pGreen II 0800-LUC vector (Lin et al., 2018) as a reporter construct were used to analyze the binding and the effect of BBX32 on the transcriptional activation potential of PIF3 and BZR1. These vectors were transfected into protoplast using PEG solution. Then the protoplasts were incubated overnight in dark and the luciferase activity was analyzed using Promega kit (E1910). The primer sequences are provided in Supplemental Table S1.
Colocalization experiments
The destination vector pGWB661 was used to clone PIF3 and BZR1 to generate RFP-fused PIF3 and BZR1 constructs. The 3-week-old N. benthamiana plants were infiltrated with Agrobacterium transformed with BBX32-GFP and RFP-PIF3 or BBX32-GFP and RFP-BZR1 vectors. After 2 d incubation in dark, the colocalization was observed using an Olympus live cell microscope. The excitation wavelengths used were 488 nm and 561 nm for GFP and RFP, respectively. The Ar-ion and Kr-ion gas lasers were used for excitation at 488 nm and 561 nm to capture GFP and RFP signals with emission wavelengths of 520 nm and 583 nm, respectively. PMT voltage (gain) was not exceeded above 750 V.
In vitro pull-down assay
BBX32 and BZR1 or PIF3 were cloned in pGEX4T1 and pCOLD, respectively, using BamH1 and ECoR1 as restriction sites. The primer details are given in Supplemental Table S1. BL21 cells transformed with GST-BBX32 were induced using 0.5 mM IPTG for 4 h at 28°C. The proteins were purified by incubating the bacterial lysate with GST beads (Glutathione Sepharose 4B, GE Healthcare) for 3 h at 4°C. In case of His-TF-BZR1 and His-TF-PIF3, the BL21 cells were induced with 0.5 mM IPTG for 16 h at 14°C. The bacterial lysate was incubated with Ni-NTA beads (Ni Sepharose 6 Fast flow, GE Healthcare) for 45 min. The unbound proteins in GST-BBX32 and His-TF-BZR1 or His-TF-PIF3 were washed with phosphate buffer (1X PBS + 300-mM NaCl) and tris buffer containing 20-mM Imidazole, respectively. GST-BBX32 was eluted using 30 mM glutathione and His-TF-BZR1 or His-TF-PIF3 was eluted using 300-mM imidazole. The concentrators (MERCK) were used to remove excessive glutathione or imidazole from protein elutes.
The purified proteins were used in pull-down assays. The GST or GST-BBX32 at 0.5-µM concentration was allowed to bind to the GST beads for 2 h at 4°C. After two washes, the prey protein His-TF or His-TF-BZR1 or His-TF-PIF3 at 1 µM according to the combinations mentioned in Figures 4H and 5B was added and incubated for 1 h at 4°C. These samples were further washed, mixed with 6X protein loading dye, and boiled for 2 min before loading on the gel for western blotting. The GST proteins were detected using anti-GST (Bethyl) and the prey proteins bound to them were detected using anti-His antibody (Bethyl anti-His).
RNA isolation from detached cotyledon and hypocotyl
All the RNA expression analyses except the overexpression levels of transgenic lines of BBX32 or PIF3 were performed by isolating cotyledons and hypocotyls separately from the seedlings in low fluence green light and freezing the samples immediately in liquid N2. The (GENAXY RNeasy Plant mini) kit was used for RNA extraction.
cDNA synthesis and RT-qPCR analysis
One microgram of total RNA was used to make cDNA. cDNA synthesis was performed using TAKARA PrimescriptTM I strand cDNA synthesis kit. RT-qPCR analysis was done using SYBR Green provided by TAKARA and the protocols were followed accordingly. The genes tested using RT-qPCR and their primer sequences are given in Supplemental Table S1.
Accession numbers
BBX32 (AT3G21150), BZR1 (AT1G75080), PIF1 (AT2G20180), PIF3 (AT1G09530), PIF4 (AT2G43010), PIF5 (AT3G59060), WAG2 (AT3G14370), LAX3 (AT1G77690), ACS5 (AT5G65800), SAUR14 (AT4G38840), SAUR16 (AT4G38860) and SAUR50 (AT4G34760).
Supplemental data
The following materials are available in the online version of this article.
Supplemental Figure S1. Spatial expression pattern of BBX32 in 5-d old dark and light-grown seedlings.
Supplemental Figure S2. Relative expression levels and cotyledon and hypocotyl phenotypes of loss-of- and gain-of-function mutants of BBX32.
Supplemental Figure S3. The hypocotyl elongation phenotype of 35S:BBX32 is suppressed by BRZ application in a dose-dependent manner.
Supplemental Figure S4. BR regulates BBX32 expression, and BBX32 does not interact with BES1.
Supplemental Figure S5. Relative expression level, cotyledon, and hypocotyl phenotypes of loss-of- and gain-of-function mutants of PIF3 and interactions of BBX32 with PIF3 and its role in affecting transcriptional activation potential.
Supplemental Figure S6. Transcriptional regulation of genes involved in opening and closing of cotyledons by BBX32 during Dark to Light transition.
Supplemental Table S1. List of primers used in the study.
S.D. and N.R. conceived the project and designed the experiments. N.R. performed all the experiments with partial help from H.R. N.J., A.Y., and K.P.V. generated or maintained resources used in this study. N.R., H.R., and S.D. analyzed the data. S.D. wrote the manuscript with help from N.R. All the authors revised the final manuscript.
The author responsible for distribution of materials integral to the findings presented in this article in accordance with the policy described in the Instructions for Authors (https://academic.oup.com/plphys/pages/general-instructions) is: Sourav Datta (sdatta@iiserb.ac.in).
Acknowledgments
The authors thank Deeksha Singh for providing pDONR207-BZR1 and pGADT7-BES1 for the study. They also thank Chetana and Sameena for the technical assistance during the study. They thank Henrik Johansson for providing pif3-3 seeds and critical reading of the manuscript. They thank Yadukrishnan and other PCDB lab members for providing valuable comments.
Funding
N.R. and H.R. acknowledge CSIR (Council of Scientific and Industrial Research) and DST-INSPIRE (Department of Science and Technology – Innovation in Science Pursuit for Inspired Research) respectively for fellowships. N.J. acknowledges DST-INSPIRE, K.P.V. acknowledges KVPY (Kishore Vaigyanik Protsahan Yojana). and A.Y. acknowledges UGC (University Grants Commission), Government of India for their fellowships. S.D. acknowledges Department of Biotechnology (BT/PR19193/BPA/118/195/2016) and MHRD-STARS (Ministry for Human Resource Development – Scheme for Transformational and Advanced Research in Sciences; STARS/APR2019/BS/245/FS), Government of India for funding.
Conflict of Interest statement. The authors declare that there is no conflict of interest.
References
Author notes
Senior author.