-
PDF
- Split View
-
Views
-
Cite
Cite
Lindell Bromham, Molecular Clocks in Reptiles: Life History Influences Rate of Molecular Evolution, Molecular Biology and Evolution, Volume 19, Issue 3, March 2002, Pages 302–309, https://doi.org/10.1093/oxfordjournals.molbev.a004083
Close - Share Icon Share
Abstract
Life history has been implicated as a determinant of variation in rate of molecular evolution amongst vertebrate species because of a negative correlation between body size and substitution rate for many molecular data sets. Both the generality and the cause of the negative body size trend have been debated, and the validity of key studies has been questioned (particularly concerning the failure to account for phylogenetic bias). In this study, a comparative method has been used to test for an association between a range of life-history variables—such as body size, age at maturity, and clutch size—and DNA substitution rate for three genes (NADH4, cytochrome b, and c-mos). A negative relationship between body size and rate of molecular evolution was found for phylogenetically independent pairs of reptile species spanning turtles, lizards, snakes, crocodile, and tuatara. Although this study was limited by the number of comparisons for which both sequence and life-history data were available, the results suggest that a negative body size trend in rate of molecular evolution may be a general feature of reptile molecular evolution, consistent with similar studies of mammals and birds. This observation has important implications for uncovering the mechanisms of molecular evolution and warns against assuming that related lineages will share the same substitution rate (a local molecular clock) in order to date evolutionary divergences from DNA sequences.
Introduction
The inverse relationship between body size and rate of molecular evolution in vertebrates is one of the most consistent patterns of lineage-specific substitution rates described to date. Martin and Palumbi (1993) described the body size gradient in rates of molecular evolution for a range of vertebrate taxa, including mammals, birds, reptiles, amphibians, and fish, and the relationship has since been confirmed for birds and mammals (Mooers and Harvey 1994 ; Bromham, Rambaut, and Harvey 1996 ). Although the inverse body size relationship has been reported for a range of molecular data, including RFLP, DNA-DNA hybridization, and nucleotide sequences (including mitochondrial and nuclear genes, RNA and protein, and both introns and exons; Martin and Palumbi 1993 ; Mooers and Harvey 1994 ; Bromham, Rambaut and Harvey 1996 ), it remains controversial. Examples of lineage-specific rates inconsistent with a body size trend have prompted questions of the generality of the relationship. The cause of the body size trend has also been debated, particularly the relative importance of two competing explanations: the generation time effect and the metabolic rate hypothesis (e.g., Martin 1995 ; Bromham, Rambaut, and Harvey 1996 ; Martin 1999 ). Furthermore, the validity of analyses that have revealed a body size relationship has been questioned (e.g., Slowinski and Arbogast 1999 ).
Resolving the controversy over the association between body size and rate of molecular evolution has important implications for understanding and using the molecular clock (the use of measures of DNA divergence to estimate the age of biological lineages). Clearly the rate of molecular evolution is not uniform across all lineages: for example, viruses can have a rate of molecular evolution a million times faster than mammals (Fitch 1996 ), and mammalian rates are an order of magnitude faster than sharks (Martin 1999 ). However, there has been a tendency to assume a local molecular clock—that closely related species will have approximately the same rate of molecular evolution—in order to use a calibration rate estimated from one lineage to date the origin of other related lineages. But if there is a general body size effect, then even closely related species could differ in substitution rate. Establishing the extent of the body size effect among vertebrate lineages is important in order to critically evaluate the common assumption of a local molecular clock.
The study of patterns of lineage-specific rates of molecular evolution has in the past been hampered by insufficient comparative data and inappropriate analyses. Available gene sequences have been concentrated in a few well-studied taxa. In particular, the most well-studied cases of lineage-specific rates have been for four mammalian species: rat, mouse, human, and cow. Although comparison of a large number of sequences from these species has served to illustrate consistent lineage-specific rates (Li et al. 1990 ; Bulmer, Wolfe, and Sharp 1991 ; Easteal and Collett 1994 ; Yang and Nielsen 1998 ), the limited number of taxa has prevented assessment of the generality of lineage-specific rates and has hampered testing of hypothetical causes of rate differences. For example, the consistently higher rates of molecular evolution in rats and mice than in humans could be caused by their shorter generation time, higher metabolic rate, lower rates of DNA repair, or all or none of these possibilities. The limited number of taxa has also led to an anecdotal approach to assessing the patterns and causes of rate variation, pointing to specific examples as evidence for or against a given hypothesis.
Furthermore, many studies seeking correlates of lineage-specific rates have been limited by failure to adequately account for phylogenetic bias in the data (e.g., Martin and Palumbi 1993 ; Omland 1997 ; Bleiweiss 1998 ; Schmitz and Moritz 1998 ). Species cannot be considered as independently derived data points in a statistical analysis because patterns of descent create hierarchies of similarity. Including multiple nonindependent comparisons could create a false association between rate and body size. But phylogenetic bias cannot wholly account for the observation of a body size effect on rate of molecular evolution because studies that have controlled for phylogenetic nonindependence have also found an inverse body size effect (Mooers and Harvey 1994 ; Bromham, Rambaut, and Harvey 1996 ).
Here, a phylogenetic comparative method has been used to demonstrate a significant association between life history and rate of DNA sequence evolution for a range of reptile lineages for three genes. The power of this study is limited by the available data—it is difficult to find sufficient comparisons between reptile species for which appropriate sequence data, life history, and phylogenetic information are all available—but the significant association between life history and rate of molecular evolution for the genes tested in this study suggests that this may be a general phenomenon in reptiles. As more data become available, the precise nature of the relationship, its taxonomic generality, and the mechanisms behind the pattern may be illuminated.
Data
Data sets for independent comparisons between reptile species were constructed for two mitochondrial protein-coding genes (cytochrome b and NADH4) and a nuclear protein-coding gene (c-mos). These genes were chosen by virtue of having sequences available in GenBank for the largest range of reptile species. Major sources of sequence data are given in the legend to table 1 . The following life-history variables were taken from the literature for each species: body size (mean adult length − maximum adult length was used for both species in a comparison where mean was unavailable); length at maturity; generation time (age at maturity); and clutch size. These variables were chosen as the most commonly available measures of life history for reptiles. Life-history data are given in table 1— not all variables were available for all species.
Methods
In assessing patterns of lineage-specific rate of molecular evolution, it is essential to use phylogenetically independent contrasts. The inclusion of nonindependent pairwise comparisons that overlap on the phylogeny risks generating a spurious association between rate and life history by effectively including the same data (internal nodes of the tree) multiple times in the analysis. This problem can be countered by selecting phylogenetically independent comparisons, such that no lineage is included in the analysis more than once (Felsenstein 1985 ; Harvey and Pagel 1991 ; Harvey and Purvis 1991 ). Lack of a fully resolved phylogeny may limit the number of comparisons that can safely be made, but it does not prevent the application of the comparative method. Here, published phylogenies and taxonomy have been used as a guide to choose conservative monophyletic pairs and outgroups (see Bromham, Rambaut, and Harvey 1996 ).
DNA sequences used in this study were taken from GenBank and aligned by eye using Se-Al (Rambaut 1996 ). Three distance matrices were estimated from each alignment. Genetic distance (D) was estimated using the HKY85 model, with transition-transversion ratio estimated from the data (implemented in PAUP*; Swofford 1999 ). Synonymous (DS) and nonsynonymous (DN) distances were estimated using PAML 2.0k (Yang 1999 ). From each of these distance matrices, the difference in rate of molecular evolution was calculated for each comparison using the relative rates test, which uses triplets of taxa to compare the distance between two ingroup taxa and an outgroup (Sarich and Wilson 1973 ; Wu and Li 1985 ). When used as a clock test to detect significant departures from rate constancy, the relative rates test frequently has poor power (Scherer 1989 ; Avise 1994 ; Robinson et al. 1998 ); for sequences less than 1,000 nucleotides, there is a substantial risk of failure to detect all cases of rate variation (Bromham et al. 2000 ). Although weak as a clock test, relative rates tests provide a simple but effective tool for examining patterns of rate variation across taxa by allowing comparison of the amount of genetic change accumulated in two sister lineages since their last common ancestor from the pairwise genetic distances between taxa. The association between the difference in branch length for each comparison and the ratio of life-history variables was assessed using both parametric (Pearson correlation) and nonparametric tests (Spearman rank correlation).
Results
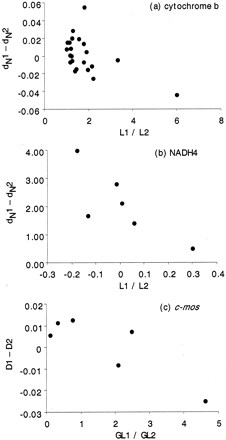
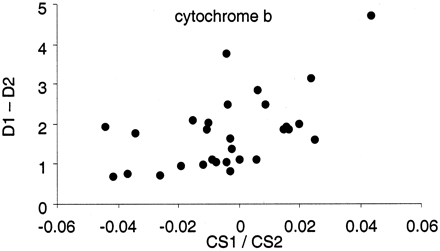
Significant association between body size and rate of molecular evolution was revealed for each of the three genes (fig. 1 ). For both cytochrome b and NADH4, there was a significant negative association between the difference in nonsynonymous branch length and the ratio of body sizes (as measured by snout-vent length: fig. 1a and b ). For c-mos, there was a significant negative association between total genetic distance and the ratio of lengths at maturity (fig. 1c ). Curiously, there was also a positive correlation between clutch size and total genetic distance for cytochrome b (fig. 2 ).
Multiple tests are a problem with this type of analysis. Testing many variables against each other increases the probability of a significant correlation between two variables occurring by chance. Correcting the P-values with a Bonferroni adjustment is overconservative in this case as the variables are not independent; life-history traits are strongly covariable, and the different measures of genetic distance use partitions of the same data. However, chance correlations between variables would not be expected to show a consistent pattern nor would that pattern be consistent with previously reported relationships. The results shown here confirm the negative relationship between rate of molecular evolution and body size reported elsewhere, with the addition of a positive relationship between clutch size and rate of molecular evolution (consistent with a prediction of Britten (1986)— see Discussion). However, these results should be interpreted in the light of the paucity of data, the low statistical power of the tests, and the problem of multiple tests. A larger data set would allow these problems to be overcome.
Discussion
The results presented here, although limited by available data, suggest that the inverse body size effect described by Martin and Palumbi (1993) is likely to be a general feature of molecular evolution in reptiles. This is consistent with body size-generation time effect observed for birds (Mooers and Harvey 1994 ) and mammals (Bromham, Rambaut, and Harvey 1996 ). The generation time hypothesis rests on the assumption that species with shorter generation times will undergo more DNA replications per unit time, and so for any given rate of copy error their DNA will accumulate more replication errors. An effect of the number of DNA replications on substitution rate is supported by the observation of male-driven evolution. It takes more cell divisions to make sperm than eggs in mammals; so genes that spend more time in males ought to accumulate more copy errors. This prediction is borne out by a comparison of sequences on Y-chromosomes (which spend all their time in males), autosomal chromosomes (which spend on an average half their time in males), and X-chromosomes (which spend only a third of their time in males) (Shimmin, Chang, and Li 1993 ; Chang et al. 1994 ; Shimmin, Chang, and Li 1994 ). A test of this hypothesis using birds, in which females are the heterogametic sex (WZ) and males have two Z-chromosomes, confirms the pattern (Ellegren and Fridolfsson 1997).
There are a number of complications of the generation time hypothesis that need clarification. Firstly, the number of germline DNA replications per organism generation varies between species, and it seems likely that it also scales with body size, potentially counteracting any generation time effect. Secondly, although mitochondria must replicate at cell division, they may also replicate throughout the lifetime of the cell at a frequency partly determined by the role and activity of the cell. This suggests that the generation time effect ought to be less obvious for mitochondrial genes, and yet they show the relationship as consistently as nuclear genes. Thirdly, the generation time effect predicts a relationship with synonymous rate (as a reflection of increase in number of mutations per unit time; Li, Tanimura, and Sharp 1987 ; Ohta 1993 ; Eyre-Walker and Gaut 1997 ), and yet the body size effect is also a feature of total genetic distance and nonsynonymous rate (this study; Martin and Palumbi 1993 ; Mooers and Harvey 1994 ). Finally, generation time clearly does not provide a global explanation of lineage-specific rate variation—for example, the range of generation times in sharks overlaps with that of primates; yet shark rates are estimated to be an order of magnitude lower than primate rates (Martin and Palumbi 1993 ; Martin 1995, 1999 ).
Given these factors which speak against a simple and universal generation time effect, why is a relationship between body size-generation time and lineage-specific rates of molecular evolution so consistently indicated for vertebrates? The first and second complications, which concern the potential decoupling of DNA replication frequency from organismal generation time, might dampen but not negate the generation time effect. For example, the influence of mitochondrial divisions might be minimized if mitochondria are effectively quiescent in female germ cells (Allen 1995 ). There are several possible explanations for the observation of a generation time effect for nonsynonymous substitutions (and total genetic distance) when the predicted effect is for synonymous change. Nearly neutral theory predicts that nonsynonymous substitutions will be approximately even over time because small animals tend to have both shorter generation time and larger population sizes, so that the effect of population size on substitution rates (faster substitution in smaller populations) could be cancelled out by the generation time effect (faster rates in small animals; Chao and Carr 1993 ; Ohta 1993 ). However, the magnitude of the effect of population and generation time on substitution rates is not known for real populations, and so it may be that the effect of generation time is not cancelled out by population size, leaving residual lineage-specific rate variation in the nonsynonymous substitution rate. The amino acid replacement rate might also be raised by selection, if rapid generation turnover speeds the process of selection (but see Rosenheim and Tabashnik 1993 ).
The fourth complication, that generation time does not provide a global explanation for lineage-specific rates in vertebrate molecular evolution, suggests that the effect of generation time on substitution rate is overridden by other effects between widely divergent lineages, such as endothermy versus ectothermy or variation in genome size. In this way, the body size-generation time effect could be considered primarily a feature of the local molecular clock. An alternative explanation for the persistence of a generation time effect, despite complications which might reduce its influence, is that generation time may be a proxy variable for an unknown causal factor that scales with body size and its covarying life-history traits (Bromham, Rambaut, and Harvey 1996 ). One such variable is metabolic rate. The metabolic rate hypothesis suggests that because small animals tend to have higher mass-specific metabolic rates than their larger relatives, they may have higher rates of production of potentially DNA-damaging metabolites, such as free oxygen radicals (Martin and Palumbi 1993 ; Rand 1994 ; Martin 1995, 1999 ). This hypothesis is consistent with the observation that poikilotherms have slower rates of molecular evolution than endotherms. It is also consistent with higher rates of molecular evolution in mitochondrial genes than nuclear genes; mitochondria are the site of oxidative metabolism and hence might be expected to suffer the greatest effect of the production of DNA-damaging metabolites. Because metabolic rate and generation time both tend to scale with body size in vertebrates, as do many life-history traits, their effects on rate of sequence evolution may be conflated. However, two studies that explicitly compared these two hypotheses found significant correlations between rate of molecular evolution and both metabolic rate and generation time but found no evidence of an effect of metabolic rate beyond its covariation with generation time (Mooers and Harvey 1994 ; Bromham, Rambaut, and Harvey 1996 ). Unfortunately, insufficient metabolic rate data prevented a similar test for this study.
Clutch Size
This study revealed a positive relationship between clutch size and rate of molecular evolution for cytochrome b. This relationship could be considered consistent with a prediction of Britten (1986)— that high fecundity will speed rate of molecular evolution by increasing the number of times the genome is copied per generation. This hypothesis shares its basic premise with the generation time effect and male-driven evolution—increasing the number of DNA replications per unit time adds more replication errors to the substitution rate.
The relationship between fecundity and number of DNA replications is not simple, as it varies between species and between sexes within a species. Production of more offspring will only increase the substitution rate by the mechanism described if increase in gamete production is achieved by increasing the average number of cell divisions taken to produce each gamete. For example, this may be true of mammalian spermatogenesis (gametes produced by a chain of cell divisions such that later sperm have undergone more cell divisions than earlier sperm), but it is less likely to be true for mammalian oogenesis. Higher fecundity may contribute to faster substitution rates in eusocial taxa than in their close nonsocial relatives (Schmitz and Moritz 1998 ; L. Bromham and R. Leys, unpublished data). Because oogenesis is continuous in Hymenoptera, eusocial species that produce a large number of sterile workers before their reproductive offspring must have a much higher number of DNA replications per generation than a nonsocial species that produces reproductive offspring directly (L. Bromham and R. Leys, unpublished data). However, it seems likely that oogenesis in reptiles resembles the process in mammals, although the germ cell development differs among reptile groups (Hubert 1985 ). The positive relationship between rate and clutch size may warrant further investigation as more sequence and life-history data become available.
Comparative Method and Molecular Rates
Studies of determinants of lineage-specific rates of molecular evolution have been limited by the perceived lack of a comparative method for testing correlates of rate variation (Slowinski and Arbogast 1999 ). The method described here and in earlier studies (Mooers and Harvey 1994 ; Bromham, Rambaut, and Harvey 1996 ) provides a general statistical framework for testing for correlates of rate of molecular evolution, for assessing the generality of relationships across taxa, as well as for comparing hypotheses. This approach is limited by the number of taxa with available DNA sequences and life-history data and by the phylogenetic resolution needed to choose independent comparisons, but growth of sequence databases will allow this test to be extended to more taxa and to more genes.
Recently, Slowinski and Arbogast (1999) suggested that the inverse relationship between body size and rate of molecular evolution revealed by Martin and Palumbi (1993) was an artifact of their analysis for two reasons: (1) failure to account for phylogenetic bias, and (2) bias toward underestimation of substitution rate in larger-bodied taxa. The comparative method employed here overcomes both these problems. Selection of phylogenetically independent pairs ensures that any differences between members of a pair must have arisen since their last common ancestor, and this will have occurred independently of any other pair. Such differences provide phylogenetically independent data for use in statistical tests. The comparative relative rates approach also overcomes the problem of biases in the calculation of absolute rates from fossil-based dates of divergence, which could lead to consistent underestimation of rates for larger-bodied lineages with deeper divergences (Slowinski and Arbogast 1999 ). Because each data point is the comparison between two lineages of the same age, any age bias in estimation of genetic distance must apply equally to both members of a pair. Bias in estimating distance could arise if the lineage with the faster rate saturates more rapidly, but this would tend to reduce the observed difference in branch length, dampening any observed relationship with size.
Implications for the Molecular Clock
The observation of a correlation between body size and rate of molecular evolution for several genes for a range of reptilian taxa echoes the body size-generation time effect observed for mammals and birds and suggests a general trend in vertebrate molecular evolution. Clearly, the data are currently limited, and the relationship must be tested for a greater number of taxa and genes before general conclusions can be drawn. Nevertheless, these results suggest that the inverse body size relationship should be explored further because it has two important implications: it provides a window on mechanisms of substitution, and it cautions reliance on a local molecular clock without rigorous testing.
The cause of the inverse body size relationship remains uncertain. The influence of number of DNA replications per unit time on substitution rate is supported by a number of observations and thus provides a plausible explanation for the influence of body size on vertebrate substitution rates. Although both generation time and metabolic rate may influence rate of molecular evolution, it is clear that neither provides a global explanation of lineage-specific substitution rates in vertebrates. It seems likely that many factors contribute to different rates of molecular evolution. Because body size correlates with a large suite of life-history traits, it is important that other traits that might influence rate of DNA change are also considered.
Whatever the cause, the observation of an inverse body size relationship for some taxa and genes suggests caution in assuming a local molecular clock for reptile species without rigorous testing. It should be acknowledged that testing for rate constancy is problematic because clock tests designed to detect lineage-specific rate variation generally have low power for most sequences used in phylogenetic analysis (Bromham et al. 2000 ). This is critical for the accuracy of molecular date estimates as failure to detect sequences with lineage-specific rate variation can result in consistent overestimation of molecular date estimates (Bromham et al. 2000 ). A life-history trend in rate of molecular evolution suggests that rates of molecular evolution must evolve over trees as life-history traits evolve, posing a dynamic picture of rate of molecular evolution, which complicates molecular phylogenetic inference (Sanderson 1997 ).
Edward Holmes, Reviewing Editor
Keywords: molecular clock molecular evolution generation time effect substitution rate relative rates comparative method
Address for correspondence and reprints: Lindell Bromham, School of Biological Sciences, University of Sussex, Brighton BN1 9QG, United Kingdom. l.d.bromham@sussex.ac.uk .
Table 1 Phylogenetically Independent Comparisons Between Reptile Species Used in this Study
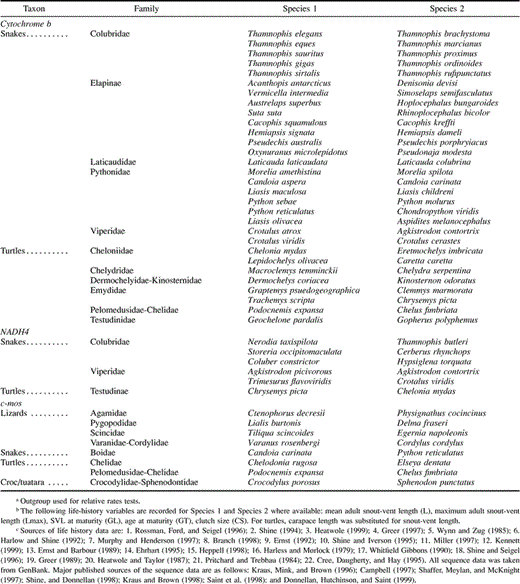
Table 1 Phylogenetically Independent Comparisons Between Reptile Species Used in this Study

Fig. 1.—Difference in relative rate of molecular evolution against the ratio of measures of body size for three genes; each point represents a phylogenetically independent comparison of two reptile species. For a list of comparisons see table 1 . Results of both Spearman and Pearson tests are given for each gene. (a) Cytochrome b (alignment length: 1,116), difference in nonsynonymous distances (dN1 − dN2) against ratio of snout-vent length (L1/L2). Spearman: rS = −0.54, P = 0.03; Pearson: r = −0.49, P = 0.05. (b) NADH4 (alignment length: 694), difference in nonsynonymous distances (dN1 − dN2) against ratio of snout-vent length (L1/L2). Spearman: rS = −0.83, P = 0.04; Pearson: r = −0.79, P = 0.059. (c) c-mos (alignment length: 375), difference in total genetic distances (D1 − D2) against ratio of snout-vent length at maturity (GL1/GL2). Spearman: rS = −0.46, P = 0.29; Pearson: r = −0.86, P = 0.03
Fig. 2.—Difference in total genetic distances (D1 − D2) against ratio of clutch sizes (CS1/CS2) for cytochrome b (alignment length: 1,116). Spearman: rS = 0.47, P = 0.03; Pearson: r = −0.42, P = 0.05. The comparison between Dermochelys coriacea and Kinosternon odoratus is not shown as it was a substantial outlier. The relationship is significant with this comparison included (Spearman: rS = 0.48, P = 0.01; Pearson: r = 0.56, P = 0.02)
The author wishes to thank Andrew Rambaut for writing useful programs, Marcel Cardillo for help with the analysis, and Paul Harvey, Simon Blomberg, Devi Stuart-Fox, Mike Lee, Meg Woolfit, and Steve Donnellan for assistance and encouragement.
References
Bleiweiss R.,
Branch B.,
Britten R. J.,
Bromham L., A. Rambaut, P. H. Harvey,
Bromham L., A. Rambaut, M. D. Hendy, D. Penny,
Bulmer M., K. Wolfe, P. M. Sharp,
Campbell B. N.,
Chang B. H.-J., L. C. Shimmin, S.-K. Shyue, D. Hewett-Emmett, W.-H. Li,
Chao L., D. E. Carr,
Cree A., C. H. Daugherty, J. M. Hay,
Donnellan S. C., M. N. Hutchinson, K. M. Saint,
Easteal S., C. Collett,
Ehrhart L. M.,
Ellegren H., A. K. Fridolfsson,
Eyre-Walker A., B. S. Gaut,
Greer A. E.,
———.
Harlow P., R. Shine,
Harvey P. H., M. Pagel,
Heppell S. S.,
Hubert J.,
Kennett R.,
Keogh J. S., R. Shine, S. Donnellan,
Kraus F., W. M. Brown,
Kraus F., D. G. Mink, W. M. Brown,
Li W.-H., M. Gouy, P. Sharp, C. O'Huigin, Y.-W. Yang,
Li W.-H., M. Tanimura, P. M. Sharp,
Martin A. P.,
———.
Martin A. P., S. R. Palumbi,
Miller J. D.,
Mooers A. Ø., P. H. Harvey,
Murphy J. C., R. W. Henderson,
Ohta T.,
Pritchard P. C. H., P. Trebbau,
Rambaut A. E.,
Rand D. M.,
Robinson M., M. Gouy, C. Gautier, D. Mouchirod,
Rossman D. A., N. B. Ford, R. A. Seigel,
Saint K. M., C. C. Austin, S. C. Donnellan, M. N. Hutchinson,
Sanderson M. J.,
Sarich V. M., A. C. Wilson,
Scherer S.,
Schmitz J., R. F. A. Moritz,
Shaffer H. B., P. Meylan, M. L. McKnight,
Shimmin L. C., B. H.-J. Chang, W. H. Li,
———.
Shine R., R. A. Seigel,
Slowinski J. B., B. S. Arbogast,
Swofford D. L.,
Whitfield Gibbons J.,
Wu C.-I., W.-H. Li,
Wynn A. H., G. R. Zug,
Yang Z.,