-
PDF
- Split View
-
Views
-
Cite
Cite
Heike Diekmann, Michael Klinger, Thomas Oertle, Dietmar Heinz, Hans-Martin Pogoda, Martin E. Schwab, Claudia A. O. Stuermer, Analysis of the Reticulon Gene Family Demonstrates the Absence of the Neurite Growth Inhibitor Nogo-A in Fish, Molecular Biology and Evolution, Volume 22, Issue 8, August 2005, Pages 1635–1648, https://doi.org/10.1093/molbev/msi158
Close - Share Icon Share
Abstract
Reticulons (RTNs) are a family of evolutionary conserved proteins with four RTN paralogs (RTN1, RTN2, RTN3, and RTN4) present in land vertebrates. While the exact functions of RTN1 to RTN3 are unknown, mammalian RTN4-A/Nogo-A was shown to inhibit the regeneration of severed axons in the mammalian central nervous system (CNS). This inhibitory function is exerted via two distinct regions, one within the Nogo-A–specific N-terminus and the other in the conserved reticulon homology domain (RHD). In contrast to mammals, fish are capable of CNS axon regeneration. We performed detailed analyses of the fish rtn gene family to determine whether this regeneration ability correlates with the absence of the neurite growth inhibitory protein Nogo-A. A total of 7 rtn genes were identified in zebrafish, 6 in pufferfish, and 30 in eight additional fish species. Phylogenetic and syntenic relationships indicate that the identified fish rtn genes are orthologs of mammalian RTN1, RTN2, RTN3, and RTN4 and that several paralogous fish genes (e.g., rtn4 and rtn6) resulted from genome duplication events early in actinopterygian evolution. Accordingly, sequences homologous to the conserved RTN4/Nogo RHD are present in two fish genes, rtn4 and rtn6. However, sequences comparable to the first ∼1,000 amino acids of mammalian Nogo-A including a major neurite growth inhibitory region are absent in zebrafish. This result is in accordance with functional data showing that axon growth inhibitory molecules are less prominent in fish oligodendrocytes and CNS myelin compared to mammals.
Introduction
In the mammalian central nervous system (CNS), regeneration of severed fiber tracts is impaired by inhibitory proteins associated with CNS myelin (Filbin 2003; Schwab 2004). Nogo-A is one of the CNS myelin components that interferes with axon regrowth in the rat and mouse CNS and provokes growth cone collapse in vitro (Chen et al. 2000; GrandPré et al. 2000; Prinjha et al. 2000; Oertle et al. 2003c). Nogo/RTN4 is the fourth member of the reticulon (RTN) gene family that codes for proteins with a highly conserved carboxy-terminal reticulon homology domain (RHD; Pfam PF02453) and a variable amino-terminus. The RHD is 150–201 amino acids (aa) in length and is characterized by two large (>30 aa) hydrophobic stretches that are responsible for the association of RTN proteins to membranes (van de Velde et al. 1994; Oertle, Merkler, and Schwab 2003). In mammals, four RTN family members are known. RTN1 (formerly neuroendocrine specific protein NSP1), RTN2, and RTN3 are enriched in membranes of the endoplasmic reticulum (van de Velde et al. 1994), but their exact functions have not been elucidated so far. RTN4/Nogo gives rise to a number of different isoforms (Nogo-A, -B, and -C as main transcripts) both through alternative splicing and alternative promoter usage (Chen et al. 2000; Oertle et al. 2003a), and the largest isoform, Nogo-A/RTN4-A, is a potent neurite outgrowth inhibitor (Chen et al. 2000; GrandPré et al. 2000; Prinjha et al. 2000). In vitro assays with recombinant peptides allowed to map the inhibitory function to two different regions of the Nogo-A–protein (Oertle et al. 2003c). One domain provoking growth cone collapse is encoded by a stretch of the Nogo-A–specific exon (NiG-Δ20; aa 544–725 of rat Nogo-A; Oertle et al. 2003c), and antibodies against this region promote in vivo CNS regeneration in rats (Schwab 2004). The second region that induces growth cone collapse is the 66-aa loop between the two C-terminal hydrophobic domains of the RHD (Nogo-66). Nogo-66 is identical in all Nogo/RTN4 isoforms and signals through an interaction with the glycosylphosphatidylinositol-linked Nogo-66 receptor (NgR) (Fournier, GrandPré, and Strittmatter 2001; GrandPré, Li, and Strittmatter 2002).
In contrast to mammals, lesioned axons readily regenerate in the fish CNS (Gaze 1970; Stuermer 1988a, 1988b). Success of CNS axon regeneration correlates with the growth-permissive substrate properties of goldfish CNS myelin in in vitro assays (Bastmeyer et al. 1991; Wanner et al. 1995). In fact, growth cones of fish retinal axons cross fish CNS myelin but collapse when contacting mammalian CNS myelin (Bastmeyer et al. 1991). This implies that fish axons recognize neurite growth inhibitors associated with mammalian CNS myelin but that fish CNS myelin is devoid of such inhibitors (Bastmeyer et al. 1991; Wanner et al. 1995). In this context, it is an intriguing question whether Nogo-A might be the molecule causing growth cone collapse of fish axons upon contact of mammalian CNS myelin and whether Nogo-A is absent from fish myelin. Here, we show that growth of fish axons in vitro is indeed blocked by a rat Nogo-A–specific peptide. To determine whether fish possess an rtn4/nogo ortholog, we cloned rtn family members. Given the high conservation in the RHD of the rtn gene family and the proposed fish-specific genome duplication (Taylor et al. 2003), we analyzed phylogenetic and syntenic relationships within the entire rtn family and unequivocally confirmed the presence of rtn4 orthologs in fish. However, comparison of the exon composition of all fish rtn genes with the respective human orthologs and dissection of sequence homologies within their N-termini argue for fundamental differences in the evolution of rtn1–rtn3 and rtn4: the specific N-termini of fish and mammalian rtn1, rtn2, and rtn3, respectively, evolved from a common ancestor, whereas the rtn4 N-termini must have been acquired independently. Fish RTN4 isoforms have short N-termini without any homology to mammalian Nogo-A, -B, or -C. Finally, the presence of exons homologous to the N-terminal region of mammalian Nogo-A in zebrafish was excluded by aligning the relevant genomic regions of zebrafish and human rtn4. Consequently, our results show that sequences related to the neurite growth inhibitory region of mammalian Nogo-A are absent in fish.
Materials and Methods
Axon Outgrowth Assay Using Goldfish Retinal Explants
Purified recombinant rat NiG-Δ20 peptide (aa 544–725 of rat Nogo-A; 3 mg/ml; Oertle et al. 2003c) was incubated as a sandwich between polylysine-coated 18 × 18-mm coverslips at 4°C overnight. The next day, coverslips were washed three times with cold modified Leibowitz medium L15 (GIBCO). Goldfish retinae were prepared 10 days after conditioning optic nerve lesion as described (Vielmetter and Stuermer 1989) and chopped into 200 × 200-μm pieces. About 50 miniexplants were plated on each NiG-Δ20–coated coverslip and incubated with F12 medium (Invitrogen, Karlsruhe, Germany) at 23°C (Wanner et al. 1995). As controls, coverslips were coated with a noninhibitory rat Nogo-A peptide (NiG-Δ36; aa 260–415 of rat Nogo-A; 3 mg/ml; Oertle et al. 2003c), with a control protein (base pairs 1650–1748 of SC1, a member of the immunoglobulin superfamily, cloned into pTrcHis [Invitrogen] and purified on Ni2+-NTA columns [Qiagen, Hilden, Germany] similar to NiG-Δ20), or with the buffer used for protein purification. After 20 h, the amount of explants with growing axons were counted using a phase contrast microscope (Axiovert Zeiss, Jena, Germany). A total of 1,087 miniexplants in 20 different cultures were evaluated using three independent protein purifications of the NiG-Δ20 peptide of rat Nogo-A, whereas for the SC-1 protein (3 different protein purifications) 669 miniexplants in 18 different cultures were analyzed. Significance of axon growth differences was calculated using Student's t-test.
Nomenclature of Fish rtn Transcripts
Zebrafish and fugu transcripts were named according to the nomenclature guidelines for RTN genes (Oertle et al. 2003b). In brief, rtn serves as a gene symbol for chordate RTNs. Paralogous rtn sequences are arbitrarily numbered. To distinguish rtn genes of various species, a prefix according to the identification code proposed by SWISS-PROT is used (e.g., (FUGRU)rtn4). Alternative transcripts generated by alternative promoter usage receive different letters (e.g., (FUGRU)rtn4-l, (FUGRU)rtn4-n), while alternatively spliced transcripts derived from a single promoter have the same letter but are distinguished by consecutive numbering (e.g., (FUGRU)rtn4-l1, (FUGRU)rtn4-l2).
Cloning and Sequence Analysis of Zebrafish and Fugu rtn Genes
Zebrafish and fugu rtn genes and transcripts were uncovered by a combination of library screening and database searches for fish expressed sequence tags (ESTs) and mRNAs with known human RTN protein sequences, reverse transcriptase polymerase chain reaction (RT-PCR), and rapid amplification of cDNA ends (RACE) (supplementary table 1A). To isolate zebrafish rtn cDNAs, two rounds of library screening were performed at the RZPD (Deutsches Ressourcenzentrum für Genomforschung GmbH, Heidelberg, Germany). The RZPD first screened high-density filters of an adult zebrafish retina library and of a late somitogenesis library using 33P-labeled base pairs 2–490 of (DANRE)rtn4-n. In a subsequent approach, the same libraries plus an adult brain library were probed with 33P-labeled base pairs 1739–2422 of (DANRE)rtn1-a1 and base pairs 1624–3157 of (DANRE)rtn6-a1. Obtained clones are listed in supplementary table 1A.
In addition, zebrafish and fugu cDNA and genomic sequences were obtained using Blast algorithms (Altschul et al. 1997) at the National Center for Biotechnology Information (NCBI) (www.ncbi.nlm.nih.gov/BLAST/), Ensembl (www.ensembl.org/Danio_rerio/blastview and www.ensembl.org/Fugu_rubripes/blastview), and the Doe Joint Genome Institute (aluminum.jgi-psf.org/prod/bin/runBlast.pl?db=fugu6) Web pages. Exon-intron structures were examined by comparing genomic sequences against cDNA sequences, respecting the GT-AG rule of splice donor and acceptor sites. For fugu rtn genes, without any ESTs or cDNA information available, exon sequences were deduced from the genomic sequences by comparison with the zebrafish cDNAs. In total, we uncovered 13 different zebrafish and fugu rtn genes with 37 mRNA variants. The sequences surrounding the putative start methionines of most fish rtn transcripts comply with the consensus motif for translation initiation (gccAccATGG) at least at one of the two most important positions (a G following the ATG and a purine at position −3; Kozak 1996). In addition, an upstream stop codon in most sequences ensures that the identified start methionines correspond to the respective N-terminus of the protein (supplementary table 1A). The predicted proteins have a more or less conserved dilysine endoplasmic reticulum membrane retention motif at their C-terminus, and the N-termini of the long splice forms (-a and -l variants) are remarkably acidic (supplementary table 1A).
We used the exon-intron information and the deduced cDNA sequences to amplify zebrafish rtn splice variants from various adult tissues (see RT-PCR) and fugu rtn transcripts from liver and brain cDNA with specific primers (supplementary tables 1A and 1B). Sequences were completed by performing 5′- and 3′-RACE, respectively. In brief, we extracted mRNA from a pool of 48-h postfertilization zebrafish embryos (FastTrack™ 2.0 kit; Invitrogen) and used 0.9 μg per reaction as template for the synthesis of either first-strand 5′-Ready cDNA using 5′-CDS and SMART II (switching mechanism at 5′end of RNA transcript) oligonucleotides or 3′-Ready cDNA using 3′-CDS primer, according to the manufacturer's instructions (SMART RACE cDNA Amplification Kit; BD Biosciences, Erembodegem, Belgium). All polymerase chain reaction (PCR) fragments were directly subcloned into the pCRII cloning vector (Invitrogen), and plasmid DNA was prepared using the QIAprep® 8 Miniprep Kit (Qiagen). Both DNA strands were sequenced using the Abi Prism® BigDye™ Terminator Cycle Sequencing Kit (Applied Biosystems, Foster City, Calif.) and analyzed on an Abi Prism 3100 Genetic Analyzer. Single sequences were assembled using SeqMan™II of the DnaStar software package (GATC Biotech, Konstanz, Germany). Specific informations for the cloning strategy of each gene are available upon request. Sequences were deposited in GenBank and accession numbers are listed in supplementary table 1D.
Sequence Alignments and Phylogenetic Analyses
GenBank accession numbers of sequences used for the different alignments are listed in supplementary table 1D. A total of 46 partial or complete RTN mRNAs were uncovered by database searches in addition to the ones already described (Roebroek et al. 1993, 1996, 1998; Kools et al. 1994; Baka et al. 1996; Stubbs et al. 1996; Ninkina, Baka, and Buchman 1997; Geisler et al. 1998; Nagase et al. 1998; Moreira, Jaworski, and Rodriguez 1999; Morris et al. 1999; GrandPré et al. 2000; Prinjha et al. 2000; Yang et al. 2000; Hamada et al. 2002; Zhang, Harrison, and Gerstein 2002; Oertle et al. 2003a; Klinger et al. 2004a). The single RTN gene of the urochordate Ciona intestinalis was used as an out-group.
Nucleotide sequences of the RHDs were translated using BioEdit (Hall 1999) and aligned as aa using ClustalW (Thompson, Higgins, and Gibson 1994). The alignment was edited by hand and then converted back into nucleotides to produce a codon alignment that was 642 nucleotides long. Due to length variations in the RTN2 genes, all other sequences contain a C-terminal gap of up to 60 nucleotides. Eighteen of the 86 sequences used for the analyses were incomplete either at the N- or C-terminus producing alignment gaps of different length. Phylogenies of RTN sequences were reconstructed using neighbor-joining (NJ) methods with MEGA version 2.1 (Kumar et al. 2001) and pairwise deletion of the aforementioned gaps. Support for nodes in the NJ tree was assessed using 1,000 bootstrap reiterations (Felsenstein 1985). The RHD alignment (ALIGN_000759) and similarly constructed alignments of the various rtn N-termini (ALIGN_000753, ALIGN_000754, ALIGN_000755, ALIGN_000756, ALIGN_000757, ALIGN_000758) were deposited at EMBL-ALIGN (http://www.ebi.ac.uk/embl/Submission/align_top.html).
Molecular evolutionary analyses were conducted using MEGA version 2.1 (Kumar et al. 2001). In brief, sequences of the RHDs were aligned at the aa level by the ClustalW program and gaps were pairwise deleted. Numbers of nonsynonymous substitutions (aa altering) per nonsynonymous site (dN) and synonymous substitutions (silent) per synonymous site were estimated (Nei and Gojobori 1986) for each fish paralog in separate comparisons to the respective human RTN sequence (supplementary table 1E).
Radiation Hybrid Mapping and Synteny Analysis
A conserved synteny is defined by two or more genes located on the same chromosome in fish and their orthologs located on a single chromosome in humans (Barbazuk et al. 2000). Therefore, zebrafish rtns were mapped on the LN54 radiation hybrid panel using standard conditions (Hukriede et al. 1999) and the respective Web interface (http://mgchd1.nichd.nih.gov:8000/zfrh/beta.cgi). Because no unequivocal result was obtained for rtn5 and rtn6 on this panel, these rtns were mapped on the Goodfellow T51 radiation hybrid panel (Research Genetics, Inc., Huntsville, Ala.) by instant mapping at http://134.174.23.167/zonrhmapper/instantMapping.htm.
For synteny analysis (Woods et al. 2000), other zebrafish genes and ESTs already mapped on the LN54 and T51 radiation hybrid panels (http://zfin.org/cgi-bin/mapper_select.cgi) were assigned to putative human orthologs by BlastX searches (Altschul et al. 1997) against the NCBI human nonredundant (nr) protein sequence database (http://www.ncbi.nlm.nih.gov/blast/blast.cgi). For EST clones that have been sequenced at the 5′ and 3′ ends, both sequences were used for BlastX searches. If the results of these searches had expected scores (E values) of ≤−5, the putative orthologs were further tested with reciprocal searches against the zebrafish subset of nr sequences and dbESTs. A human ortholog was confirmed if the original zebrafish gene or EST (or a gene or EST that showed highly significant overlap with the original sequence) was in the top 15 matches of the reciprocal search by TBlastN. Fugu synteny data were retrieved with MartView (http://www.ensembl.org/Fugu_rubripes/martview) for all scaffolds, on which other genes could be predicted in the vicinity of the fugu rtns.
Percent Identity Plots of RTN4
The zebrafish clone BX324134 from nucleotides 1 to 112,195 (linkage group [LG]6), covering the entire (DANRE)rtn4 gene, as well as the orthologous genomic sequences of the mitochondrial translation initiation factor 2 (MTIF2) and RPS27A, was aligned against 1 Mb (nucleotides 33,841,399–34,841,398) of the corresponding human region on chromosome 2p16 from the genomic contig NT_022184.13/Hs2_22340 using the minus-strand sequence. In addition, both sequences were aligned against 400 kb (nucleotides 26,300,000–26,700,000) of the orthologous mouse sequence on chromosome (Chr.) 11 from the genomic contig NT_039515.2/Mm11_39555_32 as previously described in Oertle et al. (2003a). Using a newly generated exon mask for the human genes MTIF2, RPS27A, FLJ31438, and RTN4 and a mask for repetitive sequences using the RepeatMasker software (http://repeatmasker.org), the percent identity plots (PIPs) were generated using MultiPipMaker (http://bio.cse.psu.edu/cgi-bin/multipipmaker) and the dot plot was generated using Advanced PipMaker (http://bio.cse.psu.edu/cgi-bin/pipmaker?advanced) according to the published method (Schwartz et al. 2000). The output was modified as in Oertle et al. (2003a). The same procedure was adopted using the mVista Software (http://genome.lbl.gov/vista.index.shtml) according to the published methodology (Mayor et al. 2000) with comparable results to the plots generated by PipMaker.
Reverse Transcriptase Polymerase Chain Reaction
A total of 100 zebrafish embryos (approximately 100 mg) for each stage or 50 mg of various adult tissues were used for the preparation of total RNA with the RNeasy Mini Prep Kit (Qiagen) following manufacturer's instructions. Muscle tissue was additionally subjected to proteinase K digestion (200 μg per 30 mg tissue). First-strand cDNA was synthesized under standard conditions with the Superscript First-Strand Synthesis System (Invitrogen) using oligo(dT)12–18 primer. Zero transcriptions (without Superscript II in the reaction) were performed in parallel to control for genomic DNA contaminations in subsequent PCRs. Amount and quality of the different cDNA samples were evaluated comparing the expression level of the housekeeping gene actin. Informations on primer sequences and PCR conditions are listed in supplementary tables 1B and 1C.
Results
Growth Inhibition of Fish Axons by Rat Nogo-A
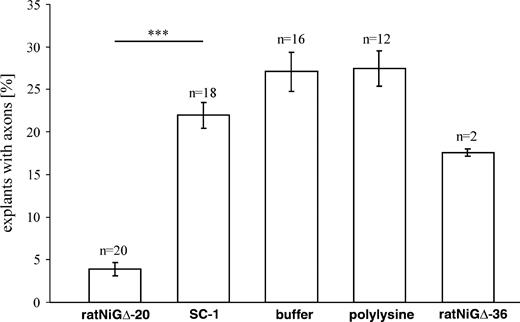
Goldfish retinal axons are able to regenerate and grow over isolated fish oligodendrocytes in vitro. In contrast, growth of fish axons is inhibited by rat oligodendrocytes (Bastmeyer et al. 1991; Wanner et al. 1995). To determine whether this inhibition might be mediated by Nogo-A protein, we evaluated growth of fish axons on the most inhibitory fragment of rat Nogo-A (NiG-Δ20; aa 544–725; Oertle et al. 2003c). Number of retinal explants with axons on recombinantly expressed Nogo-A peptide (NiG-Δ20) was compared to growth on noninhibitory Nogo-A peptide (NiG-Δ36; aa 260–415; Oertle et al. 2003c), unrelated control protein (SC-1), purification buffer, and polylysine (fig. 1). After 20 h in culture, explants with axons on Nogo-A NiG-Δ20–coated coverslips were rare (4%), as opposed to 18% explants with axons on NiG-Δ36 peptide, 22% on control protein (SC-1), 27% on buffer, and 27.5% on polylysine-coated coverslips. This growth inhibition on purified Nogo-A NiG-Δ20 peptide is comparable to that seen on rat CNS myelin (Wanner et al. 1995) and convincingly demonstrated a strong influence of rat Nogo-A on the growth of goldfish retinal axons. An intriguing question now is whether the capacity of fish to regenerate lesioned axons correlates with the absence of Nogo-A.
Axon outgrowth assay. Percentage of goldfish retinal miniexplants with axons grown after 20 h in culture. Axon growth on rat control protein SC-1, elution buffer, and polylysine-treated coverslips was robust and about six times higher than that on the NiG-Δ20 peptide of rat Nogo-A. Moreover, a similar result was obtained in a separate experiment using NiG-Δ36 of rat Nogo-A as control peptide. Bars in each column represent standard error, and asterisks indicate significant difference (P < 0.001) by Student's t-test.
Identification of Fish rtns and Their Phylogenetic Relationships
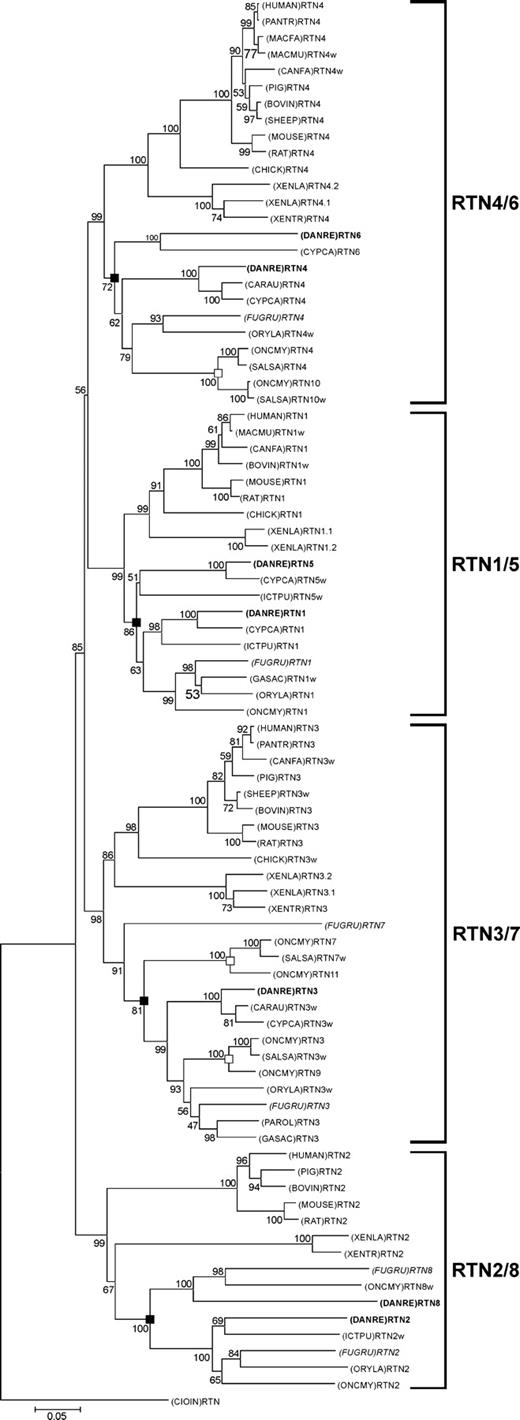
To determine whether or not true rtn4 orthologs exist in fish, we classified newly identified fish rtns by phylogenetic analysis of the vertebrate RTN gene family. Characterization of all fish rtn family members avoided conceivable misidentification due to high conservation of the RHD and ensured the detection of paralogous rtn genes resulting from the proposed fish-specific genome duplication. We physically cloned seven rtn family members from zebrafish (Danio rerio) and five from pufferfish (Takifugu rubripes) by library screening, RT-PCR, and RACE (supplementary table 1A). Database searches uncovered a sixth rtn gene within the fugu genome and 46 new partial or complete vertebrate RTN mRNAs (supplementary table 1D) compared to the ones already described (Oertle et al. 2003b). We produced an unambiguous alignment of all 86 vertebrate RTN sequences, representing the conserved RHD (ALIGN_000759). Phylogenetic reconstruction produced a tree comprising distinct, well-supported RTN1, RTN2, RTN3, and RTN4 clades (fig. 2). Each clade included human, mouse, rat, and several other tetrapod sequences; at least one zebrafish and one fugu sequence; and a variable number of other fish species, indicating that gene duplication events producing RTN1, RTN2, RTN3, and RTN4 occurred before the divergence of ray-finned fish (actinopterygians) and tetrapods (sarcopterygians). The assignment of four distinct RTN clades was supported by the identification of aa residues unique to one subfamily of rtn genes (supplementary fig. 1). The RTN1 and RTN2 subfamilies are each defined by two unique and derived (not ancestral) aa substitutions. RTN3 proteins have one diagnostic and derived residue, whereas no diagnostic residues could be identified for the RTN4 subfamily (supplementary fig. 1).
Evolutionary relationships among vertebrate RTN genes. Phylogenetic relationships of vertebrate RTN sequences as determined by NJ method with 1,000 bootstrap reiterations based on a 642-bp-long alignment of the RHDs (ALIGN_000759). Ciona intestinalis was used as an out-group. Nodes that reflect genome duplication early during fish evolution and nodes that reflect the salmonid genome duplication are highlighted with filled and open boxes, respectively. Zebrafish and fugu sequences are written in bold and italics, respectively. Sequences that did not comprise the full RHD received the suffix “w.” The scale represents 5% nucleotide sequence divergence. Abbreviations: BOVIN, Bos taurus; CANFA, Canis familiaris; CARAU, Carassius auratus; CHICK, Gallus gallus; CIOIN, Ciona intestinalis; CYPCA, Cyprinus carpio; DANRE, Danio rerio; FUGRU, Takifugu rubripres; GASAC, Gasterosteus aculeatus; ICTPU, Ictalurus punctatus; HUMAN, Homo sapiens; MACMU, Macaca mulatta; MACFA, Macaca fascicularis; MOUSE, Mus musculus; ONCMY, Oncorhynchus mykiss; ORYLA, Oryzias latipes; PANTR, Pan troglodytes; PAROL, Paralichthys olivaceus; PIG, Sus scrofa; RAT, Rattus norvegicus; SHEEP, Ovis aries; SALSA, Salmo salar; XENLA, Xenopus laevis; XENTR, Silurana tropicalis.
All clades except RTN3 contained two zebrafish genes and the RTN2 and RTN3 clades included two fugu genes (fig. 2). Consequently, the zebrafish and fugu genes were named (DANRE)rtn1–rtn4 and (FUGRU)rtn1–rtn4, respectively, and additional fish genes were consecutively numbered according to the nomenclature guidelines for RTN genes (Oertle et al. 2003b), i.e., (DANRE)rtn5, rtn6, and rtn8, and (FUGRU)rtn7 and rtn8. Fish sequences within one clade did not cluster according to species but subdivided into two subclades, each containing one of the duplicated zebrafish and/or fugu genes (fig. 2). The relationships in the resulting subclades are similar, e.g., the zebrafish genes are closely related to carp ((CYPCA)rtn1, 3, 4, 5, 6) and goldfish ((CARAU)rtn3, 4) and fugu genes are closely related to medaka ((ORYLA)rtn2, 3, 4, 8). This topology is consistent with the hypothesis that the duplicates of rtn1–rtn4 were produced before the ancestors of zebrafish and fugu diverged. The relationships between the 13 trout and salmon sequences included in this study argue for an additional Salmonidae-specific genome duplication.
To examine whether one of the two zebrafish or fugu paralogs ((DANRE)rtn1/5, rtn2/8, rtn4/6 and (FUGRU)rtn2/8, rtn3/7) evolved faster and is therefore more distantly related to the respective human RTN, we calculated the rates of synonymous (silent) and nonsynonymous (aa altering) nucleotide substitutions (dS and dN; Nei and Gojobori 1986). All zebrafish and fugu rtns accumulated similar numbers of silent nucleotide changes, but aa-altering substitutions were retained to a different extent (supplementary table 1E). In particular, twice as many changes were fixed in zebrafish rtn6 compared to rtn4, indicating that the rtn6 duplicate has evolved faster and is therefore more distantly related to human RTN4.
Taken together, our phylogenetic analysis served to assign the identified fish genes to the correct RTN subfamily and provided evidence for the existence of orthologous rtn4 genes in fish. In addition, we found duplications for all mammalian RTN tetralogs in zebrafish (rtn1/5, rtn2/8, rtn4/6) and/or in fugu (rtn2/8, rtn3/7), resulting in two paralogous genes in the respective species.
Conserved Syntenies of Fish and Human rtns
To unequivocally confirm the identity of the fish rtn genes by a second nonbioinformatical method, syntenic relationships were analyzed. Zebrafish rtn1, rtn2, rtn3, rtn4, and rtn8 were mapped on LG20, LG15, LG7, LG6, and LG21, respectively. Both rtn5 and rtn6 lie on LG13. The chromosomal positions of the mapped zebrafish rtn genes were then compared to the locations of human RTNs (fig. 3, supplementary table 1F–K).
Analyses of zebrafish and human syntenic relationships. Map locations of ESTs and genes in the radiation hybrid panels T51 and LN54 were obtained from zebrafish information network (ZFIN). The relative chromosomal locations of the human orthologs were deduced from data in LocusLink. Markers that are syntenic to rtn1, rtn3, rtn4, and rtn5 (red) are shown in green. ESTs supporting a synteny of rtn6 and human Chr. 2 are shown in blue. Markers present on more than one zebrafish panel are connected by dotted lines. (A) Conserved synteny of zebrafish LG20 and human Chr. 14 defined by (DANRE)rtn1 (red). (B) Conserved synteny of zebrafish LG13 and human Chr. 14 defined by (DANRE)rtn5 (red) and conserved synteny of zebrafish LG13 and human Chr. 2 defined by (DANRE)rtn6 (red). (C) Conserved synteny of zebrafish LG7 and human Chr. 11 defined by (DANRE)rtn3 (red). (D) Conserved synteny of zebrafish LG6 and human Chr. 2 defined by (DANRE)rtn4 (red). (E) Conserved synteny of zebrafish LG21 and human Chr. 19 defined by (DANRE)rtn8 (red). (F) Conserved synteny of zebrafish LG15 and human Chr. 19 defined by (DANRE)rtn2 (red). Abbreviations: cR, centiray; K, kilobasepairs; *, EST and genes have been published by (Woods et al. 2000); **, EST and genes have been published by Yoder and Litman (2000).
(DANRE)rtn1 together with 15 additional ESTs and the genes e(r) and esr2a on LG20 and (DANRE)rtn5 with 10 ESTs on LG13 both extend existing syntenies within the same region (14q22–14q33) of human Chr. 14 (fig. 3A and B; Woods et al. 2000). This indicates that zebrafish rtn1 and rtn5 result from a fish-specific duplication of this segment and are therefore both orthologous to human RTN1. (DANRE)rtn2 together with five ESTs (fig. 3F) and (DANRE)rtn8 together with EST fi30c01 (fig. 3E) both define the so far unrecognized small syntenic groups with the region around RTN2 on human Chr. 19. Mapping of (DANRE)rtn3, 12 additional ESTs, and the 2 genes fth1 and men1 confirmed the large syntenic region between zebrafish LG7 and human Chr. 11 (fig. 3C; Yoder and Litman 2000). (DANRE)rtn4 belongs to a known syntenic group on zebrafish LG6 and human Chr. 2 (Woods et al. 2000; fig. 3D), supported by 15 additional ESTs and the gene chma1. (DANRE)rtn6 and eight other ESTs on LG13 are also syntenic to human Chr. 2 (fig. 3B), providing additional evidence that (DANRE)rtn6 resulted from a fish-specific duplication and is therefore a second ortholog of human RTN4.
The six fugu rtn genes were identified within the genomic sequences of scaffolds 188 (rtn1), 257 (rtn2), 346 (rtn3), 2616 (rtn4), 2117 (rtn8), 3887 (rtn7, all scaffolds Ensembl release 19.2.1), and 6658 (rtn7, NCBI). Predicted genes in the vicinity of the fugu rtns were compared with the chromosomal localization of their respective mammalian ortholog. Again conserved syntenies were revealed for genes near rtn1 and human Chr. 14, for rtn2 and human Chr. 19, and for rtn3 and human Chr. 11 (supplementary table 1L).
Summarizing, the comparative mapping results in both species support the phylogenetic classification of fish rtns into four RTN subfamilies (RTN1, RTN2, RTN3, and RTN4) and the fish-specific duplications leading to rtn5, rtn8, rtn7, and rtn6. Both methods emphasize that two zebrafish orthologs of human RTN4 have been identified.
Nonhomogeneous Evolution of rtn Genes
Having ascertained the presence of orthologous rtn4 genes in fish (based on the conserved RHD), we determined the genomic organization and the N-terminal sequences of all fish rtn genes in comparison to the respective human orthologs to analyze the evolution of the variable N-termini and to examine whether domains homologous to the growth inhibitory regions in the N-terminus of mammalian Nogo-A (i.e., NiG-Δ20) are present in fish.
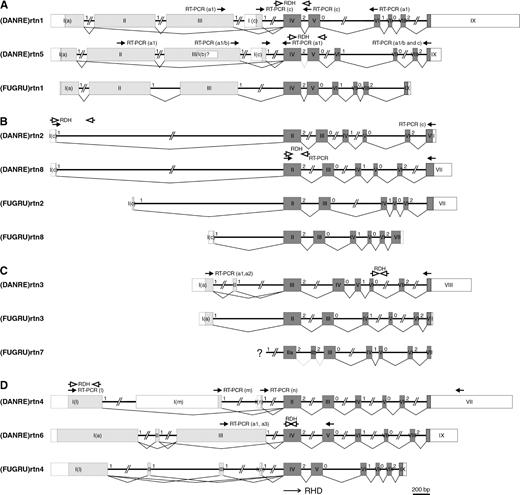
Zebrafish rtn1 and rtn5 are formed by at least 10 exons that can give rise to several isoforms (rtn1-a, rtn1-c and rtn5-a, rtn5-c, respectively) by alternative promoter usage (fig. 4A). For rtn1-a and rtn5-a, two splice forms each were detected, either with (-a1) or without (-a2) exons II + III. Cryptic splicing generated additional rtn5-a1 variants (supplementary fig. 2C). In the fugu genome, the one rtn1 ortholog comprises nine exons coding for the -a isoform (fig. 4A).
Analysis of zebrafish and pufferfish rtn genes. Exon-intron arrangements are schematically shown. Exons are drawn as roman-numbered boxes. The coding regions for paralog-specific amino-termini are shaded light gray and exons of the RHDs are shaded dark gray. UTRs are depicted as open boxes. Intron phases are specified with arabic numerals. Localizations of primers used for analyses of the zebrafish genes/transcripts are indicated by closed (RT-PCR) and open (radiation hybrid mapping) arrows, respectively. (A) Exon-intron arrangements of (DANRE)rtn1, (DANRE)rtn5, and (FUGRU)rtn1 genes in comparison to (Human)RTN1. (B) Exon-intron arrangements of (DANRE)rtn8, (FUGRU)rtn2, and (FUGRU)rtn8 genes in comparison to (Human)RTN2. Human exons 1A, 2, 3, 4, and 5 (Oertle et al. 2003b) have been omitted from this scheme for simplicity. (C) Exon-intron arrangements of (DANRE)rtn3, (FUGRU)rtn3, and (FUGRU)rtn7 genes in comparison to (Human)RTN3. For fugu (FUGRU)rtn7, the specific N-terminal exon could not be identified within the genomic sequences (marked with ?). (D) Exon-intron arrangements of (DANRE)rtn4, (DANRE)rtn6, and (FUGRU)rtn4 genes in comparison to (Human)RTN4. Human exons 1D, 1E, 1F, and 1G (Oertle et al. 2003b) have been omitted for simplicity.
Rtn2 is the most divergent of the RTN tetralogs with a longer RHD due to nucleotide insertions in the last exon. We found two paralogs each in zebrafish and fugu (rtn2 and rtn8). Based on sequence analyses, one could speculate that the insertions present in mammalian RTN2 (36 or 39 bp) and fish rtn2/8 (33–66 bp) occurred independently (ALIGN_000759). All identified fish rtn2 and rtn8 genes are built by at least seven exons, and the resulting transcripts (zebrafish and fugu rtn2-c and rtn8-c) bear only a short specific N-terminus of four aa in both species (fig. 4B, ALIGN_000755).
The eight exons of (DANRE)rtn3 can give rise to two different splice variants (fig. 4C), encompassing either all exons (rtn3-a2) or lacking exon II (rtn3-a1). In fugu, only the rtn3-a1 transcript was found (fig. 4C). The sequence of the second paralog, (FUGRU)rtn7, was deduced from genomic scaffolds because no transcripts could be found in databases or amplified from liver and brain cDNA. Therefore, we cannot exclude the possibility that it represents a nontranscribed pseudogene. Interestingly, (FUGRU)rtn7 is the only vertebrate RTN identified so far, in which the first exon of the RHD is subdivided by an extra intron (fig. 4C).
Rtn4 is formed by at least nine exons in zebrafish and eight exons in fugu. In zebrafish, three different mRNAs (rtn4-l, -m, and -n) are generated by alternative promoter usage, each consisting of one specific exon and the RHD with six exons (fig. 4D). From fugu cDNA, only the -l and -n transcripts were isolated. Fugu rtn4-l differs by the presence of two additive small exons (fig. 4D), leading to three splice variants (rtn4-l1, -l2, and -l3). The second zebrafish rtn4 ortholog, (DANRE)rtn6, comprises nine exons, and three alternative splice forms (rtn6-a1, -a2, and -a3) are produced by inclusion or exclusion of exon II or exon III (fig. 4D). Several other zebrafish rtn4 and rtn6 mRNA variants spliced at cryptic donor and acceptor sites were detected by RT-PCR (supplementary fig. 2A and B).
These extensive analyses of the genomic organization revealed that all identified zebrafish and fugu rtn genes share the same exon-intron structure in the C-terminal RHD (fig. 4, supplementary table 2), which is also conserved in mammals and reflects the high degree of sequence similarity in this region (ALIGN_0007591). The RHDs are each encoded by six exons of identical sizes (208, 139, 70, 47, 59, and 40/43/49 bp), except for rtn2 orthologs which have a longer last exon and for (FUGRU)rtn7 which has an additional intron in the first RHD exon. Intron phases between the RHD-encoding exons show a consistent pattern of 2-0-1-0-2, while the amino-terminal exons are all symmetrical 1-1 exons (fig. 4, supplementary table 2).
In contrast, the N-terminal sequences of rtn1–rtn8 are less conserved compared to the RHD and show no homology between the tetralogs (rtn1/5 to rtn2/8 to rtn3/7 to rtn4/6). However, the fish N-termini of rtn1/5-a, rtn1/5-c, rtn2/8-c, rtn3/7-a1, and rtn3/7-a2 are each orthologous to their mammalian counterparts. First of all, the exon-intron arrangements for the aforementioned transcripts are comparable to those of the respective mammalian isoforms (fig. 4A–C; Oertle et al. 2003b). For example, both N-termini of (DANRE)rtn1-a1 and (Human)RTN1-A consist of three exons each, with similar size. The same holds true for rtn3-a2, in which particularly exon 2 is conserved (fig. 4C, ALIGN_000756). Secondly, although the N-terminal sequences have diverged between different species, at least stretches of conserved aa can be identified, demonstrating a common predecessor of these N-termini (ALIGN_000753, ALIGN_000754, ALIGN_000755, ALIGN_000756).
Interestingly, the N-termini of mammalian RTN4 and fish rtn4/6 show no indication for a common ancestor. Stretches of conserved aa could neither be identified between the two fish paralogs (rtn4, rtn6) nor be identified between either of the fish duplicates (rtn4/6) and mammalian RTN4 (ALIGN_000757, ALIGN_000758). Moreover, the exon-intron arrangement of the fish rtn4 gene markedly differs from mammalian RTN4. Three alternative N-termini (-l, -m, -n) are generated by the use of three different promoters and are each encoded by a single specific exon, whereas the N-termini of mammalian RTN4 (-A, -B, -C, -D, -E, -F, -G) are represented by one to three exons (fig. 4D; Oertle et al. 2003a).
Taken together, the N-terminal parts of the rtns show higher divergence compared to the RHD, demonstrating nonhomogeneous evolution of the RTN family genes. Although not all rtn splice forms present in mammals were found in fish, detailed genomic and sequence analyses indicate that the specific N-termini of fish and mammalian rtn1/5, rtn2/8, and rtn3/7, respectively, evolved from a common ancestor. In contrast, the rtn4/6 N-termini are completely different and must have been acquired independently. Stretches of aa comparable to the mammalian neurite growth inhibitory region NiG-Δ20 were neither found in the three alternative N-termini of zebrafish rtn4 (-l, -m, and -n) nor found in rtn6-a1.
Absence of Exons Homologous to Mammalian Nogo-A
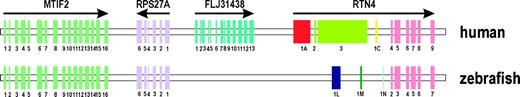
To unequivocally exclude the presence of exons homologous to the N-terminal region of mammalian Nogo-A in zebrafish, we compared the genomic region of (DANRE)rtn4 against the orthologous human and mouse genomic regions using an ungapped alignment of zebrafish versus mammalian genomic sequences ranging from the nearest common gene lying 5′ to RTN4 on the same coding strand to the six exons encoding the RHD (fig. 5, supplementary fig. 3). We identified MTIF2 (Bonner, Wiley, and Farwell 1998) as the nearest gene lying 5′ to rtn4 on the same coding strand in zebrafish, mouse, and human. The two human genes FLJ31438 and RPS27A are in fact located closer to rtn4, but an orthologous FLJ31438 gene is not present in zebrafish, and RPS27A is encoded by the complementary strand. As shown in the PIP (supplementary fig. 3A and B), the coding exons of the MTIF2 gene and the six exons of the rtn4 RHD are well conserved in both mammals and zebrafish. However, within the genomic region 5′ of the RHD, no stretches homologous to the N-terminal exons of mammalian Nogo-A (human RTN4 exons 1A, 2, 3, and 1C–G) could be identified in zebrafish (fig. 5, supplementary fig. 3). Consequently, the ungapped alignment from the MTIF2 gene to RTN4 unequivocally proved the absence of exons homologous to any known human isoforms in zebrafish.
Schematic comparison of the human and zebrafish rtn4 gene locus. Scheme of the pairwise comparison between the human-zebrafish RTN4 gene locus based on an ungapped alignment from the MTIF2 gene to RTN4 (supplementary fig. 3). Human RTN4 exons are numbered according to Oertle et al. (2003a). Note that the MTIF2 gene and the exons coding for the conserved C-terminal RHD (exons 4–9) are conserved in human and the orthologous fish locus. In contrast, no homology of the human RTN4 exons 1A, 2, and 3 could be identified in zebrafish, unequivocally proving the absence of sequences orthologous to the N-termini of mammalian RTN4 isoforms in zebrafish.
In addition, Blast searches with the sequence encoding the N-terminal region of human Nogo-A did not reveal any significant homology (>20% identity) to any other (non-rtn) zebrafish gene (data not shown). Thus, the neurite outgrowth inhibitory region in the N-terminus of Nogo-A (NiG-Δ20; Oertle et al. 2003c) is not present in zebrafish.
RT-PCR Analyses of Zebrafish rtn Expression
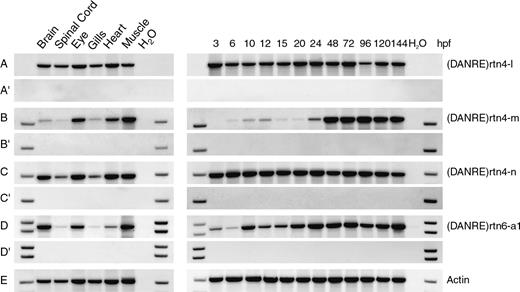
The finding that rtn4 transcripts with functions similar to mammalian Nogo-A are absent in zebrafish would be substantiated by the verification of dissimilar expression patterns of fish rtn4-l, -m, and -n in comparison to mammalian RTN4-A/Nogo-A. In addition, transcription of the other fish rtn mRNAs during zebrafish development and in different adult tissues was analyzed (fig. 6A–L) not only to characterize each rtn splice form but also to compare the expression of recently duplicated gene paralogs (e.g., rtn4–rtn6) and to obtain information about potential subfunctionalizations. The observed temporal expression patterns of the rtns can be assigned to three distinct groups. In zebrafish embryos and larvae (fig. 6, right panel), many rtn cDNAs are ubiquitously expressed without temporal variations (rtn1-a1, rtn5-b, rtn5-c, rtn3-a1, rtn4-l, and rtn4-n). Four transcripts show no (rtn5-a1, rtn2-c) or less (rtn4-m, rtn6-a1) expression at early developmental stages, whereas quite dynamic patterns are observed for rtn1-c and rtn8. All transcripts that are omnipresent during zebrafish development are also ubiquitously expressed in various adult tissues (rtn1-a1, rtn5-b, rtn5-c, rtn3-a1, and rtn4-l) with the exception that rtn1-a1 and rtn5-c are hardly detectable in gills (fig. 6, left panel). Rtn4-n and rtn6-a1 transcripts were found in all tissues analyzed, but to a varying degree. Six of the analyzed zebrafish rtn transcripts show rather tissue-specific expression (rtn1-c, rtn5-a1, rtn2-c, rtn8, rtn3-a2, and rtn4-m).
RT-PCR analyses of zebrafish rtn mRNA expression. Expression of zebrafish rtn mRNAs was examined by RT-PCR (A–L). A reverse transcriptase–negative control (without Superscript II enzyme) was performed with each primer pair (A′–L′). RT-PCR with actin-specific primers (Actin) served as a positive control and to ensure that equal amounts of cDNA template were put into each reaction (M). During development (3–144 hpf, right panel), rtn1-a1, rtn5-b, rtn5-c, rtn3-a1, rtn4-l, and rtn4-n are ubiquitously expressed (A, I, J, D, E, and G, respectively). Rtn5-a1, rtn2-c, rtn4-m, and rtn6-a1 show less expression at early developmental stages (H, C, F, and K, respectively). Rtn1-c expression is quite dynamic with repeatedly low mRNA levels at 3 and 15 hpf (B). The level of rtn8 mRNA is significantly decreased between 6 and 20 hpf (L). Note that the rtn8 RT-PCR detects the RHD and is not specific for a single isoform, but only rtn8-c is known so far. In the adult tissues analyzed (left panel), rtn1-a1, rtn5-b, rtn5-c, rtn3-a1, rtn4-l, rtn4-n, and rtn6-a1 are omnipresent, but some show varying expression levels (A, I, J, D, E, G, and K, respectively). Rtn1-c, rtn5-a1, rtn2-c, rtn8, rtn3-a2, and rtn4-m are rather tissue-specifically expressed (B, H, C, L, D, and F, respectively). Note that rtn5 RT-PCRs with a sense primer located either in exon II or at the 3′ end of exon III (fig. 1A) give totally different patterns (H, I). We therefore propose the existence of a third alternative promoter giving rise to a transcript that we call rtn5-b (fig. 1A). Abbreviations: hpf, hours postfertilization; H2O, no template control.
Overall, the extensive RT-PCR analyses revealed differences in the spatial and/or temporal expression patterns of corresponding paralogous transcripts (rtn1-a1/5-a1; rtn1-c/5-c). For example, onset of rtn5-a1 transcription is delayed with respect to rtn1-a1, meaning that only rtn1-a1 is present at early developmental stages. High-level rtn5-c in contrast to low rtn1-c expression in muscle and heart reveals tissue-specific discrimination. Because the structure of the paralogous rtn4 and rtn6 genes differ for the N-terminal exons (three alternative promoters for rtn4 compared to one promoter but alternative splicing of three exons for rtn6; fig. 4D), the resulting transcripts are not orthologous and therefore not directly comparable. Nevertheless, the expression patterns of rtn4-l, -m, or -n are not similar to rtn6-a1. These divergences in the expression of duplicated genes indicate a potential subfunctionalization. We therefore conclude that none of the gene copies is redundant and that, in particular, the paralogous rtn4 and rtn6 proteins exert different functions in fish. Moreover, none of the patterns of the four fish rtn4-homologous transcripts (rtn4-l, -m, -n and rtn6-a1) fits to the expression of the axon growth inhibitory mammalian RTN4-A/Nogo-A in neurons and oligodendrocytes (Oertle and Schwab 2003), although both mammal and fish rtn4 isoforms are expressed widely and early in development. This provides additional evidence that the identified fish rtn4 transcripts do not exert neurite growth inhibitory functions similar to mammalian Nogo-A.
Discussion
The rtn gene family member Nogo-A/RTN4-A has been widely described as a potent inhibitor of axon regeneration in mammals (Chen et al. 2000; GrandPré et al. 2000; Prinjha et al. 2000; Brittis and Flanagan 2001). In the fish CNS, however, lesioned axons readily regenerate, and fish CNS myelin has been proven to be a permissive substrate for the growth of axons (Bastmeyer et al. 1991; Wanner et al. 1995). Interestingly, fish axons are nevertheless repelled by rat oligodendrocytes in vitro. Here, we showed that growth inhibition of fish retinal axons is mediated by the inhibitory activity of the Nogo-A–specific peptide NiG-Δ20. Together with earlier findings (Bastmeyer et al. 1991; Wanner et al. 1995), this result led to the speculation that Nogo-A may not be present in the fish CNS.
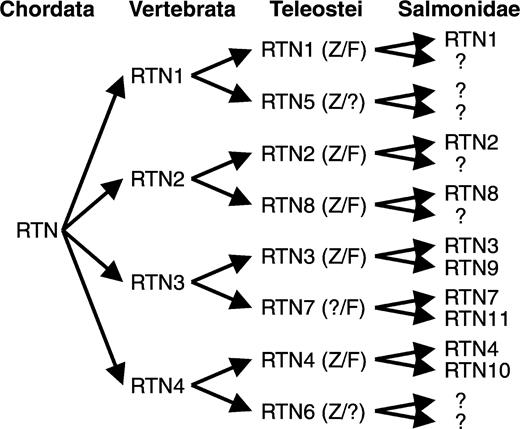
Because rtn4/nogo is a member of a large gene family with the highly conserved C-terminal RHD (Oertle et al. 2003b), phylogenetic analyses of all vertebrate rtn members coupled with gene mapping and synteny data were necessary for the correct assignment of a true fish rtn4 ortholog. We therefore cloned seven zebrafish and five pufferfish rtn genes and uncovered 30 additional rtn genes in eight different fish species by database searches. Our phylogenetic analyses based on the conserved RHD indicate that the four subgroups of the vertebrate RTN family (RTN1–4) arose by duplication events before the divergence of tetrapods (sarcopterygians) and ray-finned fish (actinopterygians) and that we identified at least one zebrafish and fugu ortholog for each subgroup. The presence of additional genes for each of the tetralogs rtn1–4 either in zebrafish and/or pufferfish that can also be seen in other fish species (e.g., carp rtn1/5 and carp rtn4/6) is consistent with the prediction of the fish-specific genome duplication hypothesis (Amores et al. 1998; Taylor et al. 2003). The phylogenetic relationships of the rtn genes identified in Salmonidae, however, cannot solely be explained by this 2R hypothesis. An additional genome duplication leading to possibly 16 different rtn genes (fig. 7) has to be proposed in this fish subgroup (Brunelli, Robison, and Thorgaard 2001; Rise et al. 2004). Assuming the validity of the genome duplications early in actinopterygian evolution and in Salmonidae, it has to be postulated that several zebrafish, fugu, and salmon rtn genes have yet to be discovered or specific gene losses have occurred (fig. 7). The differences in gene expression of duplicated rtn genes and in some cases the rapid sequence divergence of duplicated genes potentially generating alternative functional properties (e.g., relatively low sequence homology of rtn4 and rtn6) could explain why highly related paralogs were maintained in the fish genomes instead of rapidly mutating to pseudogenes (Meyer and Schartl 1999).
Schematic overview of duplicated fish rtn genes. A single rtn gene is present in the urochordate Ciona intestinalis. The divergence of the rtn family in fish was produced by separate duplication events in the ancestors of vertebrates and early in actinopterygian evolution, leading to rtn1 to rtn8 in teleostei. For Salmonidae, an additional genome duplication has to be postulated. Abbreviations: Z, the respective rtn gene has been identified in zebrafish; F, the respective rtn gene has been identified in fugu; ?, the respective rtn gene has yet to be identified or has specifically been lost.
The identity of the zebrafish rtn orthologs was confirmed by radiation hybrid mapping and synteny analyses. All seven zebrafish rtn genes are positioned within syntenic gene clusters, i.e., genes in the vicinity of a given rtn are the same on the zebrafish and the respective human chromosomes. Similar results were obtained for three of the six fugu genes. Moreover, our phylogenetic result of duplication of rtn1–4 in fish is substantiated by the synteny data that zebrafish paralogs map to regions syntenic with the same human chromosome, e.g., zebrafish rtn1 and rtn5 both with human Chr. 14. Although rtn5 and rtn6 both map to the same zebrafish chromosome (LG13), it is unlikely that these genes are the result of a tandem duplication. The distance between rtn5 and rtn6 is rather large, and each gene is located in a region that is syntenic to a different human chromosome (Chr. 14 and Chr. 2, respectively). Taken together, our detailed phylogenetic and syntenic analyses clearly demonstrate that a true fish ortholog of the mammalian rtn4/nogo gene exists and that the fish gene has been duplicated, leading to the two paralogs rtn4 and rtn6. Due to the increased accumulation of aa-altering substitutions, rtn6 is yet more distantly related to human RTN4.
To obtain indications whether the fish orthologs would exert a similar axon growth inhibitory function as the mammalian Nogo-A protein, we dissected the N-termini of fish rtn4 and rtn6 for the presence of homologous aa stretches that have been shown to exert the inhibitory function of mammalian Nogo-A (i.e., NiG-Δ20) and compared their evolution with the evolution of the rtn1–3 N-termini. The N-termini of all rtn genes are far more divergent than their RHDs; still for rtn1/5, rtn2/8, and rtn3/7, the genomic structures of the fish and the respective mammalian genes are comparable and stretches of conserved aa are detectable within these N-termini. Consequently, the N-termini of fish and mammalian rtn1–3 isoforms are orthologous and derived from a common ancestor. These findings are comparable with the nonhomogenous evolution of different parts of the annexin proteins, which has also led to evolutionary conserved C-termini and highly variable N-termini. The annexin N-termini show recognizable homology only in functionally important domains, e.g., phosphorylation sites (Farber et al. 2003). The various alternative fish and mammalian rtn4/6 N-termini, in contrast to rtn1–3, have conserved neither genomic organizations nor stretches of similar aa, which argues against a common ancestor of fish and mammalian rtn4/6 N-termini. Moreover, examination of the genomic region upstream of the zebrafish rtn4 gene proves the absence of exons homologous to the mammalian RTN4-A/B–specific exons 1A, 2, and 3 (Oertle et al. 2003a). Thus, the N-termini of fish and mammalian rtn4 have a different evolutionary origin, and the neurite outgrowth inhibitory region NiG-Δ20 of mammalian Nogo-A (rat aa 544–575) is not present in zebrafish. This result correlates with the acquisition of Nogo-A–specific sequences and the concurrent loss of the ability for axon regeneration during the transition from fish to land vertebrates (Stuermer et al. 1992). In comparison, myelin-associated glycoprotein and oligodendrocyte myelin glycoprotein, two other molecules identified as mammalian CNS inhibitors, are present in zebrafish (Lehmann et al. 2004; data not shown). This suggests that the absence of Nogo-A is causally related to successful axonal regeneration in fish. On the other hand, yet unidentified functions not related to axonal growth inhibition are expected for fish rtn4. Owing to the fact that orthologous N-termini for mammalian Nogo-A have been identified in amphibian and avian organisms (Oertle et al. 2003b; Klinger et al. 2004a), future studies will have to concentrate on the evolutionary origin of this region in organisms linking the transition between fish and amphibians.
The sequence of the second potential growth inhibitory domain, Nogo-66, which is located in the C-terminal RHD of all RTN4/Nogo isoforms, is present in fish, and the conservation throughout the whole RHD is rather high. No changes in key residues or differing domains within either the tetrapods or the fish rtn4/6 RHD could be identified that could explain the growth inhibitory function of Nogo-66 on one side (tetrapods) and the lack of inhibition on the other (fish). Still, several results and considerations argue against an inhibitory influence of fish Nogo-66. The widespread expression of all zebrafish rtn4/6 transcripts including a variety of nonneuronal tissues rather contradicts an axon growth inhibitory function. Because rtn4 transcripts are also present in goldfish oligodendrocytes (M. Klinger, unpublished data) and fish express the corresponding receptor NgR (Klinger et al. 2004b), a functional receptor-ligand interaction is conceivable. In contrast to mammals (Fournier, GrandPré, and Strittmatter 2001; GrandPré, Li, and Strittmatter 2002), however, this interaction obviously does not lead to the inhibition of axon growth in fish because fish axons grow over isolated fish oligodendrocytes in vitro (Bastmeyer et al. 1991; Stuermer et al. 1992) and readily regenerate in vivo (Stuermer et al. 1992). Therefore, if Nogo-66 interacts with NgR in fish, this interaction is coupled to an intracellular signaling cascade different from that in mammals. Alternatively, the homologous fish Nogo-66 region might not be exposed to the extracellular surface of fish oligodendrocytes and therefore not be accessible for an interaction with NgR. Future studies will show if a molecular interaction between fish Nogo-66 and NgR mediates any inhibitory activity or serves totally different purposes. This could shed light on the physiological importance of Nogo-66 as an inhibitor of axonal regeneration also in mammals.
In summary, evolution of the fish rtn4/6 genes has combined exons encoding the RHD with exons entirely different from mammalian and amphibian Nogo-A, -B, and -C–encoding exons. Sequences related to the Nogo-A major inhibitory domain NiG-Δ20 are absent from fish. While this is congruent with the growth permissiveness of fish CNS myelin, the question why fish axons respond to mammalian Nogo-A remains to be addressed in future studies.
Both authors contributed equally to this work.
Claudia Kappen, Associate Editor
This work was supported by grants of the Deutsche Forschungsgemeinschaft (DFG) and of the Fonds der Chemischen Industrie to C.A.O.S. M.K. was supported by the Stiftung der Deutschen Wirtschaft für Qualifizierung und Kooperation e.V. and H.-M.P. by the DFG (DFG PO807/1-1). We thank M.-A. Cahill for technical assistance, C. Haenisch for her help in the RT-PCR analysis, A. Y. Loos for fish care, and J. S. Taylor for critical reading of the manuscript.
References
Altschul, S. F., T. L. Madden, A. A. Schaffer, J. Zhang, Z. Zhang, W. Miller, and D. J. Lipman.
Amores, A., et al. (13 co-authors).
Baka, I. D., N. N. Ninkina, L. G. Pinon, J. Adu, A. M. Davies, G. P. Georgiev, and V. L. Buchman.
Barbazuk, W. B., I. Korf, C. Kadavi, J. Heyen, S. Tate, E. Wun, J. A. Bedell, J. D. McPherson, and S. L. Johnson.
Bastmeyer, M., M. Beckmann, M. E. Schwab, and C. A. Stuermer.
Bonner, D. S., J. E. Wiley, and M. A. Farwell.
Brittis, P. A., and J. G. Flanagan.
Brunelli, J. P., B. D. Robison, and G. H. Thorgaard.
Chen, M. S., A. B. Huber, M. E. van der Haar, M. Frank, L. Schnell, A. A. Spillmann, F. Christ, and M. E. Schwab.
Farber, S. A., R. A. De Rose, E. S. Olson, and M. E. Halpern.
Felsenstein, J.
Filbin, M. T.
Fournier, A. E., T. GrandPré, and S. M. Strittmatter.
Geisler, J. G., L. J. Stubbs, W. W. Wasserman, and M. L. Mucenski.
GrandPré, T., S. Li, and S. M. Strittmatter.
GrandPré, T., F. Nakamura, T. Vartanian, and S. M. Strittmatter.
Hall, T. A.
Hamada, N., J. Iwahashi, K. Suzuki, H. Ogi, T. Kashiwagi, K. Hara, M. Toyoda, T. Yamada, and T. Toyoda.
Hukriede, N. A., et al. (17 co-authors).
Klinger, M., H. Diekmann, D. Heinz, C. Hirsch, S. Hannbeck von Hanwehr, B. Petrausch, T. Oertle, M. E. Schwab, and C. A. O. Stuermer.
Klinger, M., J. S. Taylor, T. Oertle, M. E. Schwab, C. A. Stuermer, and H. Diekmann.
Kools, P. F., A. J. Roebroek, H. J. Van de Velde, P. Marynen, J. Bullerdiek, and W. J. Van de Ven.
Kozak, M.
Kumar, S., K. Tamura, I. B. Jakobsen, and M. Nei.
Lehmann, F., H. Gathje, S. Kelm, and F. Dietz.
Mayor, C., M. Brudno, J. R. Schwartz, A. Poliakov, E. M. Rubin, K. A. Frazer, L. S. Pachter, and I. Dubchak.
Meyer, A., and M. Schartl.
Moreira, E. F., C. J. Jaworski, and I. R. Rodriguez.
Morris, N. J., S. A. Ross, J. M. Neveu, W. S. Lane, and G. E. Lienhard.
Nagase, T., K. Ishikawa, N. Miyajima, A. Tanaka, H. Kotani, N. Nomura, and O. Ohara.
Nei, M., and T. Gojobori.
Ninkina, N. N., I. D. Baka, and V. L. Buchman.
Oertle, T., C. Huber, H. van der Putten, and M. E. Schwab.
Oertle, T., M. Klinger, C. A. Stuermer, and M. E. Schwab.
Oertle, T., D. Merkler, and M. E. Schwab.
Oertle, T., et al. (13 co-authors).
Prinjha, R., S. E. Moore, M. Vinson, S. Blake, R. Morrow, G. Christie, D. Michalovich, D. L. Simmons, and F. S. Walsh.
Rise, M. L., et al. (29 co-authors).
Roebroek, A. J., T. A. Ayoubi, H. J. Van de Velde, E. F. Schoenmakers, I. G. Pauli, and W. J. Van de Ven.
Roebroek, A. J., B. Contreras, I. G. Pauli, and W. J. Van de Ven.
Roebroek, A. J., H. J. van de Velde, A. Van Bokhoven, J. L. Broers, F. C. Ramaekers, and W. J. Van de Ven.
Schwartz, S., Z. Zhang, K. A. Frazer, A. Smit, C. Riemer, J. Bouck, R. Gibbs, R. Hardison, and W. Miller.
Stubbs, L., et al. (12 co-authors).
Stuermer, C. A., M. Bastmeyer, M. Bahr, G. Strobel, and K. Paschke.
Stuermer, C. A. O.
———.
Taylor, J. S., I. Braasch, T. Frickey, A. Meyer, and Y. Van De Peer.
Thompson, J. D., D. G. Higgins, and T. J. Gibson.
van de Velde, H. J., A. J. Roebroek, N. H. Senden, F. C. Ramaekers, and W. J. Van de Ven.
Vielmetter, J., and C. A. Stuermer.
Wanner, M., D. M. Lang, C. E. Bandtlow, M. E. Schwab, M. Bastmeyer, and C. A. Stuermer.
Woods, I. G., P. D. Kelly, F. Chu, P. Ngo-Hazelett, Y. L. Yan, H. Huang, J. H. Postlethwait, and W. S. Talbot.
Yang, J., L. Yu, A. D. Bi, and S. Y. Zhao.
Yoder, J. A., and G. W. Litman.
Author notes
*Department of Biology, University of Konstanz, Konstanz, Germany; †Brain Research Institute, University of Zurich, Zurich, Switzerland; ‡Department of Biology, Swiss Federal Institute of Technology Zurich, Zurich, Switzerland; and §Department of Developmental Biology, Stanford University School of Medicine