-
PDF
- Split View
-
Views
-
Cite
Cite
Elizabeth L. Jockusch, Developmental and Evolutionary Perspectives on the Origin and Diversification of Arthropod Appendages, Integrative and Comparative Biology, Volume 57, Issue 3, September 2017, Pages 533–545, https://doi.org/10.1093/icb/icx063
Close - Share Icon Share
Abstract
Jointed, segmented appendages are a key innovation of arthropods. The subsequent diversification of these appendages, both along the body axis and across taxa, has contributed to the evolutionary success of arthropods. Both developmental and fossil data are informative for understanding how these transitions occurred. Comparative analyses help to pinpoint the developmental novelties that distinguish arthropod appendages from the lobopodous appendages of other panarthropods, and that distinguish different appendage types. The fossil record of stem group arthropods is diverse and preserves intermediate steps in these evolutionary transitions, including some that cannot be directly inferred based on extant taxa. These lead to hypotheses that can be tested with comparative developmental data, as well as to reinterpretations of developmental results. One developmental novelty of arthropods is the reiterated deployment of the joint formation network, which divides the appendages into segments. The fossil record raises questions about how this joint formation network was first deployed, given the contrasting morphologies of appendages in stem group versus extant arthropods. The fossil record supports a character tree for appendage diversification showing progressive individuation of appendages in an anterior-to-posterior sequence. However, to date, developmental evidence provides at best limited support for this character tree. Recent interpretations of the fossil record suggest that the labrum of extant arthropods is a greatly reduced protocerebral appendage pair; this hypothesis is consistent with the extensive shared developmental patterning of the labrum and jointed appendages. Reciprocal illumination from fossils and developmental patterning in a phylogenetic context both makes sense of some results and helps motivates questions for future research.
Introduction
Jointed, segmented ventral appendages are the hallmark of arthropods, and they are thought to be a key innovation that has helped fuel the tremendous evolutionary success of this clade (Adamowicz etal. 2008). These arthropodized appendages have diversified in both form and function, contributing to core organismal activities including reproduction, feeding, sensation, locomotion, respiration, and defense. One reason that appendages are so functionally important is that arthropods have many pairs of them and their diversification along the body axis allows for functional diversification within an individual.
Context for the origin and diversification of arthropod appendages is provided by their close relatives, the panarthropod phyla Onychophora (velvet worms) and Tardigrada (water bears) (Rota-Stabelli etal. 2013), as well as by an extensive stem group fossil record (Fig. 1). Onychophorans and tardigrades have lobopodous appendages; these appendages are unsegmented and unbranched, retaining morphological features similar to those inferred for the common ancestor of panarthropods. Multiple major transitions in appendage morphology occurred along the branch that separates extant arthropods from their panarthropod relatives, including arthropodization, the origin of appendage branches, and differentiation of appendage types along the body axis (Legg etal. 2013; Edgecombe and Legg 2014; Smith and Ortega-Hernández 2014; Ortega-Hernández etal. 2017). In keeping with the symposium theme, this review is focused on the origin and diversification of arthropodized appendages because perspectives gained from fossils, phylogeny, and development provide some degree of reciprocal illumination, while stimulating additional questions amenable to developmental research.
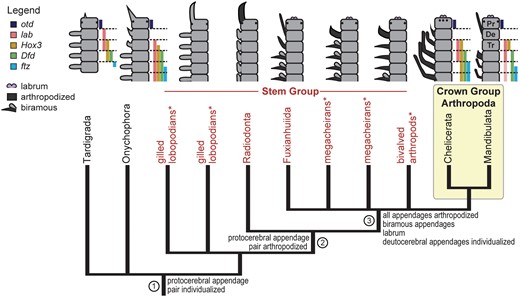
Phylogeny and body plans of extant panarthropods and the arthropod stem group that give insight into arthropod appendage evolution (mapped characters). * indicates named taxa that may not be monophyletic. In the body plan diagrams, gene expression patterns (colored bars, in same order as in legend) show homology of the five anterior segments across extant panarthropods (slightly modified from Smith etal. (2016), with permission from Elsevier). Different shapes indicate different appendage types; arthropodized appendages are dark gray; non-arthropodized ones are light gray. For fossils, segmental affiliations of appendages follow Ortega-Hernández etal. (2017). References to colors in all figures apply to the electronic version of this article. Genes: Dfd, Deformed; ftz, fushi tarazu; lab, labial; otd, orthodenticle. Segment names: Pr, protocerebral; De, deutocerebral; Tr, tritocerebral.
Context from the arthropod stem group
The arthropod stem group includes a morphologically, ecologically, and taxonomically diverse set of fossils (Legg etal. 2013; Ortega-Hernández 2016) extending from the Early Cambrian into at least the Early Devonian (Kühl etal. 2009; Siveter etal. 2014). In phylogenetic analyses, Radiodonta, of which the most famous member is Anomalocaris, is recovered as the most distant relative of crown-group arthropods that bears arthropodized appendages (Legg etal. 2013) (Fig. 1). In these large, soft-bodied predators, only the anterior-most appendage pair was arthropodized, whereas more posterior appendages have instead evolved into flaps (Daley and Edgecombe 2014).
Three additional hypothesized stem-group taxa, the bivalved arthropods, megacheirans, and fuxianhuiids, are characterized by arthropodization of all appendages (Legg etal. 2013) (Fig. 1). These groups bear classic biramous (branched) limbs on most body segments, but all show a highly morphologically divergent anterior-most appendage pair, along with additional appendage specialization in the head (Yang etal. 2013; Aria and Caron 2015; Liu etal. 2016). Mapping these appendage traits onto the currently accepted hypotheses for relationships indicates that arthropodization arose initially in the context of the frontal appendage, and subsequently spread to (or was independently evolved in) all more posterior appendages (Legg etal. 2013). Analyses of the fossils further suggest that diversification of the anterior appendages occurred in a step-wise fashion, proceeding from anterior to posterior (Ortega-Hernández etal. 2017).
Arthropodization of appendages: the origin of podomeres and joints
Appendages originate, in both development and evolution, as body wall outgrowths. Thus, they add a set of novel proximodistal (PD) axes to the body plan. An early discovery of evolutionary developmental biology was that growth and patterning of this novel axis are regulated by a conserved network of transcription factors (reviewed in Angelini and Kaufman 2005). This network provides a coordinate system that differentiates regions along the PD appendage axis. This network is hypothesized to have been coopted from patterning of a more ancient axis, the anteroposterior nervous system axis (Lemons etal. 2010). Recent work has extended the conservation of this appendage patterning network to the lobopodous appendages of onychophorans (Fig. 2). Thus, overall PD axis patterning predates the origin of arthropodized appendages (Janssen etal. 2010a; Oliveira etal. 2014).
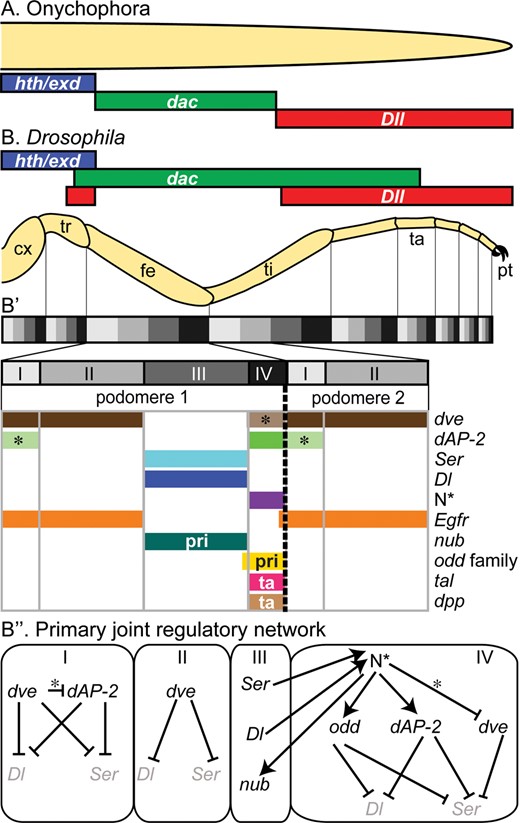
Subdivision of the appendage axis and formation of joints. Initial subdivision of the PD appendage axis is similar in onychophorans ((A), based on Janssen etal. 2010a) and arthropods ((B), Drosophila). (B’, B’’) In Drosophila, the axis is subdivided further by the reiterated expression of a suite of interacting genes that regulates joint formation (modified from Jockusch and Smith (2015) with permission of Springer); some genes have functions restricted to the primary (pri) or tarsal (ta) joints. N* indicates activated form of Notch; elsewhere, * indicates late interactions. Genes: dac, dachshund; dAP-2, Drosophila activator protein-2; Dl, Delta; Dll, Distal-less; dpp, decapentaplegic; dve, defective proventriculus; Egfr, Epidermal growth factor receptor; hth/exd, homothorax/extradenticle coexpression; N, Notch; nub, nubbin; odd, odd-skipped (and paralogs); Ser, Serrate; tal, tarsal-less. Appendage segments: cx, coxa; fe, femur; pt, pretarsus; ta, tarsus (subdivided into 5 tarsomeres); ti, tibia; tr, trochanter.
Central to the transformation of lobopodous appendages into arthropodized appendages is the subdivision of the PD axis into podomeres (segments) separated by joints. Joint formation involves activation of a second coordinate system at repeated positions along the PD axis. Early evidence for the existence of a reiterated coordinate system came from classic regeneration studies that showed a developmental equivalence between tissues from similar PD positions of different podomeres (French etal. 1976). Molecular support comes from the expression of numerous genes in a pattern of one circumferential ring per podomere (reviewed in Jockusch and Smith 2015). How appendages became segmented and jointed is thus in part a question about the origin of this second, reiterated axis patterning network.
A key output of this reiterated patterning system is activation of Notch signaling immediately proximal to each joint (Fig. 2B’). In Drosophila, this is accomplished via spatially restricted expression of both Notch ligands, Delta and Serrate (Rauskolb 2001). Comparative data support a conserved, ancestral role for Notch signaling during joint formation in arthropods, although which ligand is used appears to vary across species and appendage types (Prpic and Damen 2009; Mito etal. 2011; Turchyn etal. 2011; Angelini etal. 2012a, 2012b; Chesebro 2013; Coulcher etal. 2015) (Fig. 3). The joint formation network both up- and downstream of Notch signaling is relatively well understood in Drosophila (reviewed in Jockusch and Smith 2015) (Fig. 2B”). Comparative data on the expression and function of these genes remain piecemeal, but suggest that their roles may vary across taxa (Fig. 3). Several genes inferred to have ancestral roles in arthropod joint formation (Prpic and Damen 2009) have been studied in other contexts in onychophorans (Treffkorn and Mayer 2013; Janssen and Budd 2013, 2016; Janssen etal. 2015). These give no evidence of repeated rings of expression (Fig. 3). This absence is unsurprising given the lobopodous appendage morphology, and is consistent with the hypothesis that the joint formation network was newly deployed in appendage development in the common ancestor of arthropods (Prpic and Damen 2009).
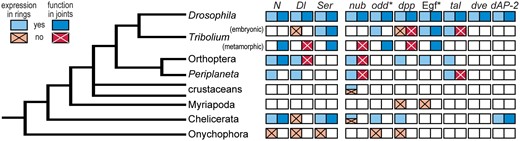
Comparative data on candidate genes for joint formation based on the Drosophila model. Color and pattern indicate evidence supporting or not supporting a joint formation role based on expression (in one ring per segment) or function (in most cases, RNAi-mediated downregulation resulting in loss of all or most leg joints). Multiple colors within a cell indicate variation across surveyed species. Empty cells indicate species/gene combinations for which relevant data were not found. * indicates that information from multiple genes is taken into account: both receptor- and ligand-encoding genes for the epidermal growth factor pathway and multiple paralogs in the odd gene family. References and additional details are provided in Supplementary Table S1. Gene abbreviations as in Fig. 2.
Extant arthropods have a relatively small number of podomeres in most appendage types, and the ancestral number of leg podomeres for crown group arthropods is inferred to have been seven (Boxshall 2004). Each podomere is typically both morphologically identifiable and evolutionarily conserved within major clades, although secondary subdivision has occurred in some lineages and appendage types (e.g., as illustrated by the insect tarsus, Fig. 4C). In Drosophila, there is evidence that joint formation at each podomere boundary may be independently regulated downstream of the conserved PD axis patterning network. For example, rings of dAP-2 (Ahn etal. 2011) and Serrate expression (Bachmann and Knust 1998; Rauskolb 2001) in different podomeres are regulated by different cis-regulatory elements. The stereotyped nature of appendage segmentation patterns, combined with the irregular but conserved order in which joints are defined developmentally (Rauskolb 2001), suggests that this independent regulation became codified early in the history of different major lineages, and thus may be amenable to reconstruction via comparative analysis.
Fossils reveal an intermediate step in the origin of segmented appendages. (A) Lobopodous appendages illustrated by the onychophoran Peripatoides novaezealandiae (photo copyright Gil Wizen) were transformed into homonomously segmented appendages in stem group arthropods, illustrated by (B) the fuxianhuiid Chengjiangocaris kunmingensis (Yang etal. 2013; photo courtesy of Xi-guang Zhang and Jie Yang). Subsequently, a reduced number of distinct podomere types evolved in the common ancestor of extant arthropods ((C) leg of the flour beetle Tribolium castaneum, reprinted from Smith and Jockusch (2014), with permission from Elsevier).
Fossils and the origin of segmented appendages
When segmented trunk appendages first appear in the fossil record, their morphology is quite different from that of extant arthropod appendages. Instead of being formed by a small set of uniquely identifiable podomeres, they are formed by a graded series of relatively homonomous (similar) podomeres (Fig. 4B). The ancestral number of podomeres remains uncertain, but the number is inferred to have undergone reduction along the lineage leading to the arthropod crown group (Legg etal. 2012). The number of podomeres varies across closely related species, and can be large. In an extreme case (presumed to result from an evolutionary increase), upwards of 30 podomeres are inferred in the legs of the bivalved arthropod Nereocaris briggsi (Legg and Caron 2014). Megacheirans and fuxianhuiids also have representatives with nine or more similar podomeres (Legg and Vannier 2013; Yang etal. 2013; Aria and Caron 2015; Liu etal. 2016).
The contrasting morphologies and patterns of variation in fossil and extant forms suggest that different mechanisms were employed to establish the podomere boundaries ancestrally versus in extant arthropods. In particular, it is unlikely that a system in which boundaries are individually placed via different components of the PD axis patterning network could have been the ancestral mechanism that gave rise to relatively homonomously segmented appendages. Rather, it seems more likely that ancestrally podomeres were generated via one of two distinct mechanisms that can generate repeating patterns: either a growth zone that forms appendage segments via molecular cycling or alternatively by wiring the joint formation network downstream of a regulatory gene that was already expressed in a reiterated PD pattern in ancestral lobopodous appendages. These considerations give rise to a two-step model for the transformation of lobopodous appendages into arthropod appendages (Fig. 4). In the first step, ancestrally lobopodous appendages acquired a joint formation network, which was under similar developmental regulation in each podomere. In the second step, regulation of podomere boundaries became individuated, stabilizing the podomere composition.
Such a scenario has been supported for analogous changes in mechanisms generating body segments. Ancestrally, most body segments are inferred to have been generated via the cycling of a molecular segmentation clock in the posterior growth zone (Peel etal. 2005; El-Sherif etal. 2012; Schönauer etal. 2016). Subsequently, in long germ insects such as Drosophila, this mechanism was replaced by the independent establishment of body segment boundaries, via a mechanism that is dissociated from growth, and involves multiple independent cis-regulatory regions. This transformation is perhaps best illustrated by the regulation of eve-skipped (eve) expression. In both beetles and spiders, eve stripes are established as part of or downstream of the segmentation clock (El-Sherif etal. 2012; Schönauer etal. 2016). By contrast, in Drosophila, seven similar stripes of eve expression are generated by different combinations of body axis patterning genes binding to five separate enhancers, each of which generates one or two stripes (Vincent etal. 2016).
Open questions about the evolution of joints and segments
The origin of appendage segmentation and the expected contrast in its developmental regulation in the arthropod stem group versus extant arthropods raise a series of questions and hypotheses that can be investigated with comparative developmental data. One question is which components of joint patterning were present in the ancestral network. More systematic comparative analysis of the candidate genes across major arthropod lineages, including chelicerates, crustaceans, and myriapods, is an important step towards reconstructing the ancestral network. A second question is how the joint formation network originated. Morphologically, a natural hypothesis is that it was coopted from a similar, evolutionarily more ancient process operating to divide the body axis into segments (although these are not separated by true joints) (Prpic and Damen 2009). Current data provide some support for this hypothesis, as Notch signaling is required for segmentation in a diverse set of arthropods, suggesting that it was a component of the ancestral body segmentation pathway (e.g., Damen 2007; Pueyo etal. 2008; Eriksson etal. 2013; Schönauer etal. 2016, but see also Kainz etal. 2011). The pair rule gene odd-skipped (odd) is also required for both body and appendage segmentation in some arthropods (e.g., Angelini etal. 2012b; Sarrazin etal. 2012). However, Wnt signaling, another common and possibly ancestral element in body segmentation (Janssen etal. 2010b; Chesebro etal. 2013; Schönauer etal. 2016), has not been implicated in appendage segmentation. Alternatively, the joint formation network may have been activated ancestrally at multiple positions via a gene that was already expressed in a reiterated pattern along the PD axis of lobopodous appendages. Appendages of both extant onychophorans (Fig. 4A) and some fossil lobopodians (Ortega-Hernández 2016) display morphological repetition along the PD axis. These morphological patterns suggest that repeated molecular addresses may already have been present along the PD appendage axis in the ancestor of panarthropods. Finally, one notable exception to individualized appendage podomere identities is flagellate appendages, such as orthopteran antennae, which have large numbers of similar podomeres. Such flagellate appendages offer a suitable system for studying how homonomously segmented appendages are generated in extant arthropods.
Diversification of serial homologs through individuation of appendage identities
A second aspect of arthropod appendage evolution is diversification along the body axis from an initially similar series. Individual elements in a homonomous series cannot be directly homologized across lineages. Unique identities emerge when elements of a homonomous series become individuated (Wagner 2014). This process creates appendage types that are directly homologous across taxa and can be traced through evolution. Serial homologs that have evolved distinct identities are interesting from an evo-devo perspective because of their combination of evolutionary independence and developmental interdependence (Jockusch etal. 2004; Wagner 2014).
One complication in developing a model for the evolution of appendage identities has been homologizing body segments across lineages, both extant and fossil. A significant success of comparative evo-devo has been resolving the segmental homologies for living taxa, solving one aspect of the long-standing arthropod head problem (reviewed in Scholtz and Edgecombe 2006; Ortega-Hernández etal. 2017). Comparative gene expression data provide a strongly supported model for conserved segmental identities, in a 1:1 alignment, across the anterior five segments of crown group arthropods (Damen etal. 1998; Telford and Thomas 1998) (Fig. 1). Recent comparative data have extended this 1:1 alignment to onychophorans (Eriksson etal. 2010; Janssen etal. 2014) and tardigrades; in tardigrades, the first of these segments forms the head and the rest constitute virtually the entire trunk (Smith etal. 2016). The distinct, conserved patterning of anterior segments, regulated in part by Hox genes, predates the specialization of anterior appendage types and provides a molecular substrate for the establishment of unique appendage identities.
There have also been significant advances in our understanding of the homologies of segments in fossil taxa, which show a diverse set of anterior appendage morphologies and specialization patterns. An important new source of data is fossilized traces of the nervous system (reviewed in Edgecombe etal. 2015), which has greatly reduced the reliance on similarity in appendage morphology as an indicator of segmental identity. The new interpretations suggest that robust, raptorial appendages arose multiple times independently and are associated with the protocerebral, deutocerebral or tritocerebral segment in different stem group taxa (Fig. 1) (Ortega-Hernández etal. 2017).
A character tree for the diversification of appendage identities and patterning
Comparisons of appendage diversity patterns along the body axis of living panarthropods and the arthropod stem group furnish a model for the evolution of distinct appendage identities, which can be tested with developmental data. The evolutionary scenario can be represented as a “character tree” (Musser and Wagner 2015), showing the predicted history of individuation of appendage types from the ancestrally homonomous series as lineages diversified (Fig. 5). The origin of a new character identity is represented as a branch point in the tree. Anteroposterior diversification of appendages appears to have occurred in a step-like fashion, with a unique protocerebral appendage identity extending back to the common ancestor of panarthropods (Ortega-Hernández etal. 2017). Both tardigrades (Smith etal. 2016) and radiodontans (Daley etal. 2013; Van Roy etal. 2015) retain the predicted ancestral morphology of a differentiated protocerebral appendage pair followed by a relatively homonomous series of more posterior appendages (Fig. 1).
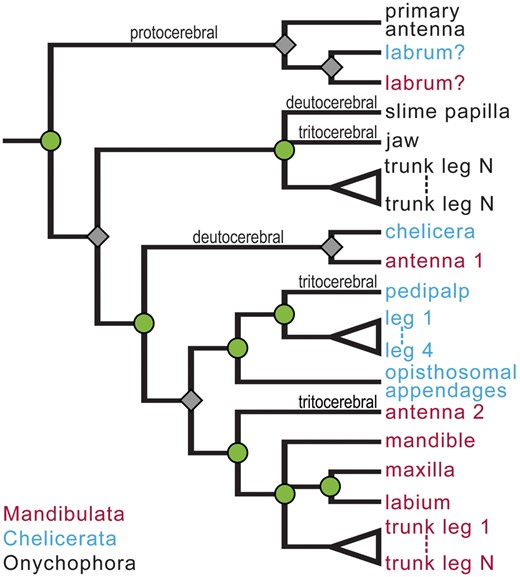
Character tree showing the hypothesized history of individuation of appendage identities from an ancestrally homonomous series. Splits occur by two separate processes: the individuation of a new appendage identity (nodes marked by green circles) and speciation (gray diamonds). This distinction is similar to the more familiar distinction between gene duplication (creating paralogs) and speciation (creating orthologs) as different causes of divergence in gene family trees. Both kinds of splits identify sets of appendages predicted to uniquely share developmental features. The clustering of appendages associated with the protocerebral segment of all three lineages indicates that this appendage identity evolved in a common ancestor of the included taxa, while separation of deutocerebral appendages occurs because individuation of the deutocerebral appendages is hypothesized to have occurred independently in onychophorans versus arthropods. Similarly, individuation of the tritocerebral appendages may have occurred independently in three lineages.
The next appendage to acquire a distinct identity is inferred to have been the deutocerebral appendage. This is the anterior-most jointed appendage in extant arthropods, forming the (first) antenna of mandibulates and chelicera of chelicerates. A differentiated deutocerebral appendage type is reconstructed for all stem-group arthropods except the radiodontans (Ortega-Hernández etal. 2017). Some megacheirans display the inferred transitional pattern of differentiated deutocerebral appendages followed by a relatively homonomous series of biramous appendages (Ortega-Hernández etal. 2017), as do trilobites (Hughes 2003).
Fuxianhuiids (Yang etal. 2013), some bivalved arthropods (e.g., Loricicaris, Legg and Caron [2014]), and crown group arthropods all possess morphologically distinctive tritocerebral appendages. However, fossil forms show a variety of patterns of specialized anterior appendages (i.e., tagmosis patterns). This, combined with uncertainty about phylogenetic resolution of some of these lineages, complicates inferences about the evolution of identities of appendages on the tritocerebral and subsequent segments. It is likely that unique tritocerebral appendage identities evolved independently multiple times, including in crown group chelicerates and mandibulates. Diversification of the post-oral appendages is also inferred to have occurred independently in the Chelicerata and the Mandibulata (Fig. 5).
Developmental predictions
This character tree makes predictions about conservation and divergence of appendage patterning across arthropods which can be evaluated with comparative data on gene expression and function. A shared identity is expected to manifest itself through uniquely shared aspects of developmental patterning across taxa, even when that appendage type takes highly diverse forms. For example, the morphologically highly divergent deutocerebral appendages of crown group chelicerates and mandibulates are predicted to share derived developmental features that differentiate them from more posterior appendage types (Sharma etal. 2015). A likely target for changes in patterning when novel identities evolve is the early-acting PD axis patterning genes, which were expressed ancestrally in all appendages and interact with the identity-regulating Hox genes. The character tree further predicts that such sharing of unique developmental features will not occur across appendages that are located on homologous segments, but that are inferred to have evolved their identities independently, such as the deutocerebral appendages of arthropods versus onychophorans.
Another consequence of this character tree is that crown group arthropods descended from an ancestor with a differentiated pair of jointed protocerebral appendages, directly homologous to the protocerebral appendages of onychophorans and tardigrades. Some current interpretations of head segmentation of fossils support the hypothesis that the labrum is the unjointed remnant of the ancestral protocerebral appendages (Ortega-Hernández etal. 2017). An alternative hypothesis is that the frontal filaments, found in both some stem group fossils (Ortega-Hernández and Budd 2016) and crustaceans, are what remain of the protocerebral appendages (Frase and Richter 2013). This leads to the prediction that the labrum or frontal filaments will develop via a modified appendage patterning network and, furthermore, that the protocerebral appendage derivates will uniquely share developmental patterning similarities with the primary antennae of onychophorans (Janssen 2017).
Testing predictions of conservation and divergence of appendage patterning
Developmental genetic work in Drosophila provides a model for how PD axis patterning differs in antennae (the deutocerebral appendages of mandibulates) versus legs, regulating their different morphologies. The key early difference is co-expression of the PD axis patterning genes homothorax (hth) and Distal-less (Dll) in the distal antenna, but repression of hth by Dll in the distal legs (Dong etal. 2002). Loss of hth expression in the antenna transforms it to leg identity. Recently, a chelicera-to-leg transformation was reported in response to hth knockdown in a chelicerate. This result suggests that the function of hth in determining deutocerebral appendage identity could be ancestral for arthropods (Sharma etal. 2015). As predicted by the hypothesized character tree (Fig. 5), this trait is not shared with onychophorans, in which deutocerebral (jaw) hth expression resembles that of more posterior appendages (Janssen etal. 2010a).
However, a complication of this hypothesis for the origin of arthropod deutocerebral appendage identity is that hth knockdown can also lead to homeotic transformations of other anterior appendage types in some arthropods (Inbal etal. 2001; Ronco etal. 2008; Sharma etal. 2015). Additional work is needed to determine whether, contrary to the character tree prediction, these result from a developmental role that is shared between the Hox-less deutocerebral appendages and other anterior appendage types. Alternatively, the effects of hth knockdown on post-deutocerebral appendages may be mediated through the function of Hth as a Hox cofactor, reflecting a different developmental role (Ronco etal. 2008).
Another candidate gene that may have contributed to the ancestral divergence of arthropod deutocerebral appendages from more posterior appendages is spineless. In Drosophila, coexpression of hth and Dll activates expression of spineless, which regulates additional antenna-specific patterning (reviewed in Jockusch and Smith 2015). Strong expression of spineless along much of the PD axis distinguishes the antennae from more posterior appendages in multiple hexapods (reviewed in Jockusch and Smith 2015), with RNA interference data confirming its role in determining antennal identity in the beetle Tribolium (e.g., Shippy etal. 2009). However, the lack of data from chelicerates, along with expression of ss in other mandibulate head appendages (e.g., Shippy etal. 2009; Janssen 2013), leaves it unclear whether the deutocerebral appendage function is ancestral for arthropods and unique to this appendage. In onychophorans, spineless shows a restricted tip expression in all appendages (Oliveira etal. 2014), a pattern consistent with the prediction that modifications of deutocerebral appendage patterning are not shared between onychophorans and arthropods.
Models for how PD axis patterning is modified to give rise to other distinct appendage morphologies, such as the mouthparts of mandibulates, are less well developed (but see Scholtz etal. 1998; Joulia etal. 2006; Lebreton etal. 2008; Angelini etal. 2012a; Doumpas etal. 2013). Multiple factors contribute to this. One factor is the evolutionary suppression of tritocerebral appendages in both hexapods and myriapods. Another is the highly modified nature of the mouthparts in Drosophila. A third factor is that the expression and function of PD axis patterning genes during mouthpart development appear to vary across the small number of mandibulate species for which data are available (Prpic and Tautz 2003; hexapod data reviewed in Jockusch and Smith 2015). This is likely explained by the many ways in which mouthpart morphologies have been modified. A final factor is that much of the work on mouthpart development has focused on endite-derived structures (e.g., Prpic and Tautz 2003; Jockusch etal. 2004; Angelini etal. 2012a; Coulcher and Telford 2013), which are a component of the ancestral appendage morphology that has been retained in these appendages, but suppressed in the walking legs of many lineages. The developmental basis for diversification of the morphologically diverse posterior appendage derivatives in chelicerates also remains poorly studied, offering opportunities for future work (Pechmann etal. 2010).
The labrum as the protocerebral appendage pair
The labrum (or more correctly, the labrum/hypostome complex) is an enigmatic preoral structure, whose origin and homology have been the subject of extensive debate among arthropod biologists; even its segmental affiliation has been debated (Scholtz and Edgecombe 2006). Functionally, the labrum forms an “upper lip.” Morphologically, it is a relatively anterior, median, unjointed, and unpaired structure. Developmentally, it originates from two limb-like buds that rotate and fuse (Kimm and Prpic 2006). Recent interpretations of fossils support the hypothesis (Budd 2002) that the labrum represents the remnants of the protocerebral appendage pair (Ortega-Hernández and Budd 2016; Ortega-Hernández etal. 2017). The loss of this appendage pair—corresponding to the primary antennae of onychophorans and the great appendages of radiodontans—corresponds in time with the gain of the unpaired median labrum (Fig. 1).
Development of the labrum has been studied in greatest detail in the flour beetle Tribolium castaneum. This work revealed extensive similarities in molecular patterning and gene function of the labrum and jointed appendages during both embryogenesis (Posnien etal. 2009; Siemanowski etal. 2015) and metamorphosis (Smith etal. 2014). Based on prior views of the labrum as not serially homologous to paired appendages, the patterning similarities were extensive enough that they were interpreted as resulting from co-option (either from labrum to legs or from legs to labrum (Posnien etal. 2009; Smith etal. 2014)).
With the conclusion that the labrum may instead be a serial homolog of paired appendages, the developmental similarities and differences can be evaluated from the perspective that they result from modification of the ancestral appendage patterning network. The labrum retains expression and function of the full complement of early PD axis patterning genes (Posnien etal. 2009; Smith etal. 2014), which, interestingly, would be unusual among morphologically highly reduced appendages. One of these, Dll, may have a unique function in labrum development. Dll downregulation leads to loss of the complete labrum in multiple lineages of both chelicerates (Schoppmeier and Damen 2001; Sharma etal. 2013) and mandibulates (Beermann etal. 2001; Angelini and Kaufman 2004); the proximal regions of other appendages are retained after Dll downregulation.
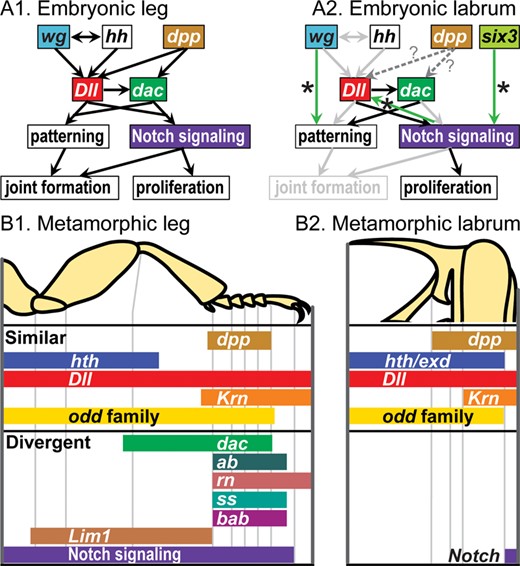
Embryonically, the most striking differences between the labrum and more posterior appendages lie in (1) activation of the early PD axis patterning genes Dll and dachshund (dac) via a novel mechanism, rather than via a suite of conserved body axis patterning genes (Ober and Jockusch 2006; Posnien etal. 2009) and (2) the failure of Notch signaling to activate joint formation (Siemanowski etal. 2015) (Fig. 6A). At metamorphosis, the appendage patterning network appears to be truncated in the labrum relative to jointed appendages, with early axis patterning genes functioning in similar domains along the PD axis, but no evidence for a conserved function for later-acting genes, primarily those required to subdivide the distal limb and form joints (Smith etal. 2014) (Fig. 6B). At both stages, the labrum displays a set of developmental features that would be predicted of a true appendage that had lost joints, consistent with the hypothesis that it represents the protocerebral appendages.
Comparison of leg and labrum patterning in the beetle Tribolium castaneum. (A) Embryonic gene interaction networks (based on Siemanowski etal. 2015); in (A2), light gray arrows indicate interactions that are present in the legs (A1) but absent in the labrum; green arrows with stars indicate novel interactions detected in the labrum; dotted arrows with question marks indicate interactions that were not assessed. (B) Metamorphic functional domains, as evidenced by RNAi (adapted from Smith etal. 2014 with permission from Wiley).
Complicating factors: evolutionary homeosis and “concerted evolution”
Two factors have the potential to modify appendage development in ways that complicate interpretation of the evolutionary history of appendage identities and patterning based on developmental data. The first factor is the occurrence of evolutionary homeosis, which changes the identity of an appendage associated with a particular segment. This is well illustrated by work on maxillipeds (mouthpart-like appendages that develop in place of legs) in crustaceans (Martin etal. 2016). The second factor is that even after acquiring distinct identities, divergent appendages types within a lineage are patterned by a shared genome and thus are expected to remain developmentally linked to varying degrees throughout the history of a lineage (Jockusch etal. 2004; Angelini etal. 2012b; Wagner 2014). This developmental linkage motivates a prediction of “concerted evolution” in patterning, resulting from changes that simultaneously affect multiple appendage types within a lineage. Such concerted changes do not map to branches in the hypothesized character tree.
The extent of linkage has been characterized in a large-scale enhancer screen in imaginal discs of D. melanogaster; these structures give rise to the adult body (Jory etal. 2012). More than half of the enhancers showing expression in any surveyed imaginal disc drove similar expression in the legs and antennae. Many of these enhancers also drove similar expression in the genitalia, a morphologically highly modified posterior appendage derivative (Jory etal. 2012). Mutations in these enhancer elements, or in genes encoding proteins that interact with these enhancer elements, are expected to modify development of multiple appendage types simultaneously in a lineage-specific fashion.
Such changes are expected to give rise to lineage-specific features of appendage development, aspects of the developmental regulatory network that characterize multiple appendage types within a lineage, but that distinguish appendage development in that lineage from patterning in relatives and inferred patterning in ancestors (Musser and Wagner 2015). One well-documented case of this predicted “concerted evolution” comes from correlated changes across appendage types in the expression domains of the PD axis patterning genes hth and extradenticle (exd); these changes appear to have happened multiple times. Hexapods (reviewed in Jockusch and Smith 2015) share with onychophorans (Janssen etal. 2010a) a restriction of hth to proximal regions of most appendage types at most developmental stages, whereas exd expression extends distally to (near) the tip. By contrast, in chelicerates and myriapods, hth expression extends distally to exd expression (Prpic etal. 2003; Sharma etal. 2012). In spiders, these genes are both duplicated, and one copy shows repeated rings of expression in multiple appendage types (Prpic etal. 2003).
Such concerted evolution has multiple consequences. One consequence is that overall similarity (including derived similarity) of developmental patterning may not match the character tree for appendage identity. Instead, greater developmental similarity may be observed between evolutionarily divergent appendage types within a lineage than is observed between appendages that share an evolutionary identity but that occur in distantly related lineages. Another consequence is that the sharing of gene expression or function across morphologically divergent serial homologs within a lineage does not necessarily provide evidence that the developmental feature was ancestral. The occurrence of concerted evolution can be identified with sufficient taxon sampling, but has the potential to complicate inferences about the evolution of patterning across serial homologs.
Conclusion
In this review, I have aimed to highlight several ways in which thinking about developmental and fossil data in concert promotes insights into major events in the evolution of arthropod appendages, while also identifying areas for future research. It focuses on subdivision of the appendages into segments separated by joints as the true arthropod novelty, and one whose developmental origin remains obscure. It further highlights a likely transformation in how joints are positioned based on comparisons of the morphology of fossil versus extant arthropod appendages. The inferred sequence of divergence of appendage identities leads to hypotheses about developmental similarities which are just beginning to be tested, and which may be complicated by patterns of concerted evolution. The review also suggests that patterning similarities between the labrum and jointed appendages are consistent with interpretations of the fossil record according to which the protocerebral appendage pair was transformed into the labrum.
One morphological feature for which similar reciprocal illumination would be hoped is another key innovation in arthropod appendages: the biramous appendage. Theories for how this came about have been put forth based on developmental data (e.g., Olesen etal. 2001; Wolff and Scholtz 2008) and the fossil record has also yielded insights (Budd 1996; Zhang and Briggs 2007). While some extant crustaceans retain biramous appendages with well-developed exopods and endopods (Boxshall and Jaume 2009), these remain underrepresented in developmental analyses of appendage patterning.
Acknowledgments
My thinking about arthropod appendages has been shaped by discussions and collaborative work with David Angelini, Frank Smith, and Prashant Sharma. Conversations at the SICB conference with Javier Ortega-Hernández helped guide my thinking about the arthropod fossil record. I very much appreciate reviews from Graham Budd, Ralf Janssen and an anonymous reviewer, as well as feedback from Frank Smith, all of which improved the manuscript. Special thanks are due to Xi-guang Zhang and Jie Yang for the photograph of Chengjiangocaris, to Gil Wizen for the onychophoran photograph, and to Ariel Chipman and Doug Erwin for organizing a stimulating symposium.
Funding
ELJ’s work on arthropod appendages has been supported by the United States Department of Agriculture (Cooperative State Research, Education, and Extension Service seed grant 2006-35604-16746).
Supplementary data
Supplementary data available at ICB online.
References
Author notes
From the symposium “The Evolution of Arthropod Body Plans–Integrating Phylogeny, Fossils and Development” presented at the annual meeting of the Society for Integrative and Comparative Biology, January 4–8, 2017 at New Orleans, Louisiana.