-
PDF
- Split View
-
Views
-
Cite
Cite
M. H. Buys, H. J. Flint, E. M. Miller, H. Yao, A. R. Caird, R. J. Ganley, Preparing for the invasion: efficacy of DNA barcoding to discern the host range of myrtle rust (Puccinia psidii) among species of Myrtaceae, Forestry: An International Journal of Forest Research, Volume 89, Issue 3, July 2016, Pages 263–270, https://doi.org/10.1093/forestry/cpw017
Close - Share Icon Share
Abstract
Puccinia psidii is a major threat to members of the Myrtaceae worldwide. New Zealand is still free of P. psidii but possesses significant Eucalyptus plantations as well as valuable native tree species such as Metrosideros excelsa and Leptospermum scoparium that could be at risk in the event of an incursion. As part of preparing for a potential incursion, we generated a barcode reference library for >100 diverse species of Myrtaceae occurring in New Zealand by sequencing internal transcribed spacer (ITS), external transcribed spacer (ETS) and maturase K (matK), or by obtaining relevant sequences from GenBank. The Myrtaceae DNA barcoding database will enable rapid identification of large numbers of host species in the event of a myrtle rust incursion. We undertook a comparative analysis of the ability for the three mentioned loci to discriminate species. Interspecific divergence was assessed by mean interspecific distance, θ prime and minimum interspecific distance; intraspecific variation was evaluated by mean intraspecific difference, θ and coalescent depth. Overall identification efficiency of the three loci sequenced was determined using BLAST1 and Near Distance methods. Barcoding gaps between inter- and intraspecific divergences were also analysed. ITS and ETS share similar mean interspecific distance, θ prime and minimum interspecific distance values – both higher than obtained for matK. In contrast, in terms of intraspecific variation, matK had higher values than ITS and ETS in all three metrics analysed. ITS had the highest identification success rate for species followed by matK and ETS, respectively, as measured by BLAST1 and Near Distance methods. Overall identification success rate increases when a combination of ITS and matK in particular is used. The sequence data are not only a valuable reference collection for a myrtle rust response but also a national botanical resource.
Introduction
Puccinia psidii Winter, causal agent for what is known variously as myrtle rust, eucalyptus rust or guava rust, received particular attention when it was found to affect Eucalyptus L'Hér. in Brazil in the early 1900s (Joffily, 1944) and has since become an important biosecurity threat to many countries. The pathogen has extended its geographic range to other South and Central American countries. It has reached continental US (Marlatt and Kimbrough, 1979) and subsequently most of the Hawaiian Islands (Uchida et al., 2006). Japan (Kawanishi et al., 2009), Australia (Carnegie et al., 2010) and China (Zhuang and Wei, 2011) have not been spared, and most recently, records from South Africa (Roux et al., 2013), New Caledonia (F. Giblin, unpublished data) and Indonesia (A. McTaggart, personal communication) have been reported, despite warnings that P. psidii is a significant threat to Myrtaceae globally (Coutinho et al., 1998; Tommerup et al., 2003). In Australia, the presence of myrtle rust has taken on a new significance because the country represents one of the centres of diversity for Myrtaceae (Wilson et al., 2001), especially for species with capsular fruits such as Eucalyptus, which are almost entirely endemic to the region (Ladiges, 1997). The host range of P. psidii has expanded significantly since its introduction into Australia, infecting >250 species in 56 genera to date (Carnegie, 2015; Machado et al., 2015b) and with the potential to infect more as evidenced by artificial inoculation (Morin et al., 2012).
It is beyond the scope of this article to detail the taxonomy, biology and pathology of P. psidii, and readers are referred to Pegg et al. (2014) and Machado et al. (2015b) for an overview. Initial uncertainty regarding the identification of the pathogen in Australia (Elith et al., 2013) has been recognized as one of the reasons why the window of opportunity to eradicate has passed (Howard et al., 2015). Although a number of strains with differing infecting characteristics have been identified in South America, only a single strain is present in Australia (Loope and La Rosa, 2008; Graça et al., 2013; Machado et al., 2015a, b). Puccinia psidii in general harms young, developing tissue and depending on the host, infects young leaves, shoots, flower buds and/or fruit (Glen et al., 2007; Zauza et al., 2010) causing defoliation, branch dieback and abortion of flowers and fruits (Pegg et al., 2014). Unsurprisingly, myrtle rust has had a negative economic impact worldwide (e.g. Junghans et al., 2003; Ribeiro and Pommer, 2004; Loope and La Rosa, 2008), and in the context of Australia in particular, estimates by Cannon (2011) suggest an annual loss of 1.5 million m3 in plantation volume due to the disease. The full ecological and commercial impact of P. psidii in Australia will become evident within the foreseeable future (Carnegie and Lidbetter, 2012; Carnegie et al., 2015).
New Zealand is ‘myrtle rust’ free, for now. Climate niche modelling, based on recent climatic data, has highlighted regions in New Zealand's North Island that provide optimal conditions for growth and survival year round (Kriticos et al., 2013). However, the spectre of global warming will affect the range and impact of the pathogen (Bebber, 2015). There is concern that P. psidii may soon arrive in New Zealand (Goldson et al., 2015; Teulon et al., 2015) either by anthropogenic introduction (including myrtaceous nursery stock) or by long-distance dispersal (McKenzie, 1998; Clark, 2011). While other rust fungi have blown across the Tasman Sea and infected plants in New Zealand (McKenzie, 1998), there are indications that much of the long-distance spread of myrtle rust within Australia was due to human movement of infected material (Makinson, 2014). A negative correlation between spore release and increasing wind speed and turbulence intensity of the air was found (Zauza et al., 2015), but once released, the spores are capable of moving across smaller distances.
If the current strain of P. psidii in Australia was to arrive in New Zealand, it would threaten the country's mānuka honey industry utilizing Leptospermum scoparium J.R.Forst. & G.Forst and high-value environmental assets such as the iconic native trees Metrosideros excelsa Sol. ex Gaertn. (pohutukawa) and M. robusta A.Cunn. (northern rata), both of which are already under threat from browsing by possum (Trichosurus vulpecula Kerr). New Zealand only has 24 native species of Myrtaceae (De Lange and Rolfe, 2010), compared with more or less 2250 endemic species in Australia (CHAH Australian Plant Census, unpublished data). From cultivated New Zealand natives in Australia, it is known that P. psidii can infect M. excelsa, M. kermadecensis W.R.B.Oliv. and Lophomyrtus bullata Burret (Giblin and Carnegie, 2014). There are, however, numerous cultivated species of Eucalyptus recorded in New Zealand (386 species documented in the National Forestry Herbarium database) as well as a wide range of other genera with horticultural value. The strain of P. psidii present in Australia is currently not causing serious disease in mature eucalypt plantations, affecting only young trees in relatively small numbers (Carnegie, 2015) as opposed to one of the strains found in Brazil that severely affects eucalypt plantations (Graça et al., 2013).
New Zealand has been afforded the opportunity to prepare for a myrtle rust invasion. The Ministry for Primary Industries' (MPI) High Risk Site Surveillance programme is tasked with providing early detections of unwanted plant pests of tree species present in New Zealand (Stevens, 2008), and the programme has been put on a myrtle rust alert. The identification of species of Myrtaceae by morphological means can be time consuming and often requires a combination of characters, which may not all be present at a particular time. The more rapidly species of Myrtaceae can be identified, the more rapidly surveying for myrtle rust and a response to an incursion can take place. Accurate surveillance of infected hosts will enable MPI to consider appropriate biosecurity measures in order to protect threatened plant species and vegetation communities and to reduce the risk of spread to new areas by workers and visitors (Quinn and Buys, 2014). Moreover, transporting infected plants to herbaria across New Zealand for identification poses a risk of spore escape. In contrast, extracted DNA can be more safely transported, and with DNA barcoding, a large number of specimens can be identified in a relatively short space of time.
Collaboration between the National Forestry Herbarium and MPI Plant Health and Environment Laboratory scientists resulted in the development of a plant DNA barcoding database of >100 species of Myrtaceae occurring in New Zealand. DNA barcoding for identifying specimens relies on a reference library of sequences from known species. Sequence data generated from query specimens can then be identified by matching it to sequences in the reference library. The breadth and depth of the reference library determines the degree of identification success (Bergsten et al., 2012). Although there have been calls to improve the DNA barcoding process, e.g. Zhang et al. (2012), the power of current DNA barcoding procedures relies in its standardization and the subsequent scalability associated with that (Collins and Cruickshank, 2014). This work forms a key part of MPI's preparedness and ability to respond to damaging exotic pests and diseases in the event of an incursion (Quinn and Buys, 2014). We report on the efficacy of maturase K (matK), internal transcribed spacer (ITS) and the external transcribed spacer (ETS) to discern the selected species in our database.
Materials and methods
Taxon sampling
We compiled a species list after consulting the New Zealand National Forestry Herbarium (NZFRI) database, the New Zealand Virtual Herbarium and MPI data (species listed in the Plant Biosecurity Index, Plant Pest Information Network and High Risk Site Surveillance database). The following factors were considered when compiling this list: (1) maximization of sampling coverage across the family; (2) inclusion of species present in New Zealand that are known to be susceptible to myrtle rust (based on Australian myrtle rust host lists, e.g. www.dpi.nsw.gov.au/biosecurity/plant/myrtle-rust/hosts); (3) inclusion of species that have high economic significance (including about a dozen of the main species of Eucalyptus used in the forestry industry, Anonymous, 1995); (4) inclusion of species that have cultural significance (i.e. all native species) and (5) inclusion of species where sufficient material was available to provide three or more samples per species in order to identify interspecific variation. During the pilot phase of the project, loci for a small number of additional species other than those ultimately decided upon were found on GenBank or were in some instances generated by us. These additional data were included in the barcode reference library as well as the analysis reported on below. The full list of species in the barcode reference library, as well as corresponding GenBank numbers, is provided in Supplementary Tables S1–S3.
Barcoding loci
The Plant Working Group within the Consortium for the Barcode of Life (CBOL) recommends targeting two loci in the chloroplast genome for barcoding purposes, namely the ribulose-bisphosphate carboxylase (rbcL) gene and the matK gene. The combination of the two aforementioned loci has not proven to be equally informative across the entire plant world and overall has a relatively low discrimination success rate of 72 per cent at the rank of species (CBOL Plant Working Group, 2009). In particular, rbcL has fallen in disfavour when compared with matK (Hollingsworth et al., 2011). In contrast, the ITS from the nuclear genome is widely used in taxonomy and molecular phylogeny because it is easy to amplify and has a high degree of variation even between closely related species (Hollingsworth et al., 2011; Li et al., 2011). In particular, ITS2 has gained patronage for barcoding purposes and Chen et al. (2010) have shown that it has a discrimination success rate of 92.7 per cent in plants.
For the purposes of our work, we used matK and ITS (ITS1 and ITS2) as the primary loci and choose the 5′ ETS as a third region for a subset of taxa (Angophora Cav., Corymbia K.D.Hill & L.A.S.Johnson, Eucalyptus and Metrosideros Banks ex Gaertn.). ETS has not been promoted as a barcoding region (Hollingsworth et al., 2011), but has been used with success in combination with ITS in phylogenetic studies (e.g. Logacheva et al., 2010).
Obtaining samples
The existing collection in the National Forestry Herbarium provided 128 samples for DNA extraction. Together with specimens housed in the Auckland War Memorial Museum Herbarium (AK), herbarium specimens made up 43 per cent of samples obtained for this project. The majority of samples (54 per cent) were obtained from field-based collections and subsequently placed in silica gel. The remaining 3 per cent of the DNA extractions were obtained directly from fresh material. Voucher specimens for newly collected samples have been lodged at the National Forestry Herbarium (NZFRI).
DNA extraction
DNA was extracted from leaf samples using the NucleoSpin Plant II DNA extraction Kit (Cat. #740770.50, Macherey–Nagel), with minor modifications to the protocol. The modifications were as follows: (1) samples were chopped into small pieces and placed in FastPrep™ tubes from MP Biomedicals (containing ceramic bead and garnet chips); (2) 400 µl of Buffer PL2 and 13 µl RNase A were added and the tubes homogenized in an OmniRuptor from Omni International for two cycles of 5 m/s for 20 sec; (3) after incubation for at least 60 min at 65°C, 100 µl of Buffer PL3 was added and tubes incubated on ice for 5 min and (4) tubes were centrifuged for 5 min at maximum speed and the cleared supernatant was loaded onto the NucleoSpin® Filters. DNA was stored at −20°C.
Polymerase chain reaction and sequencing
Two different polymerase chain reaction (PCR) protocols were used in this study. Generally, ITS PCRs were more difficult to amplify and JumpStart™ REDTaq® ReadyMix (Sigma) was the polymerase that worked best (Protocol 1). matK and ETS PCRs generally worked well using our standard PCR protocol with HOT FIREPol® Blend Master Mix, Ready to Load (Solis BioDyne) (Protocol 2).
Protocol 1: PCR reactions contained 1× JumpStart™ REDTaq® ReadyMix, 10 µg bovine serum albumin, primers (Supplementary Table S4) to a final concentration of 0.3 µM. PCR was carried out on a Perkin Elmer PE9700 thermal cycler using the following programme: initial denaturation at 94°C for 3 min, followed by 35 cycles of 94°C for 30 sec, annealing at 56°C for 30 sec, 72°C for 1 min and a final 10 min extension step at 72°C.
Protocol 2: PCR reactions contained HOT FIREPol® Blend Master Mix, Ready to Load, primers to a final concentration of 0.3 µM. PCR was carried out on a PE9700 thermal cycler using the following programme: initial denaturation at 94°C for 15 min, followed by 35 cycles of 94°C for 30 sec, annealing at 58°C for 40 sec, 72°C for 1 min and a final 10 min extension step at 72°C.
Successful PCR products were treated with Exonuclease I (Fermentas) and FastAP Thermosensitive Alkaline Phosphatase (Fermentas) prior to sequencing. Sequencing was done either by Macrogen (Korea) or in-house on the 3500 Series Genetic Analyzer (Applied Biosystems®) using BigDye® Terminator v3.1 Cycle Sequencing Kit (Applied Biosystems®).
Sequence length, inter- and intraspecific divergence, barcoding gap and specimen identification
The inter- and intraspecific divergences were calculated by first using ClustalW to align sequences and then calculating Kimura 2-parameter distances (Kimura, 1980) using PAUP (Swofford, 2002). Sequences with ambiguous bases or incomplete sequence lengths were excluded from the analyses, e.g. matK data for Blepharocalyx O.Berg., Darwinia capitellata Rye and D. meeboldii C.A.Gardner obtained from GenBank.
Interspecific divergence was assessed by (1) mean interspecific distance (Meier et al., 2008); (2) θ prime (the mean pairwise distances calculated for each species that has more than one representative, thereby eliminating biases associated with uneven sampling among taxa; Meyer and Paulay, 2005) and (3) minimum interspecific distance (Meier et al., 2008).
Intraspecific variation was evaluated by (1) mean intraspecific difference, (2) θ and (3) average coalescent depth (the maximum distance from tips of a node linking all sampled members of a species, i.e. ‘book-ending’ intraspecific variability; Lahaye et al., 2008).
DNA barcoding gaps (difference between inter- and intraspecific genetic distances within a group of organisms) were analysed following Ross et al. (2008 and references therein) and species identification success was calculated using the BLAST1 method (the assigned identification is that of the species associated with the best BLAST hit) as well as Nearest Distance method in which the assigned identification is that of the species of the sequence having the smallest genetic distance from the query (Ross et al., 2008).
Results
Lists of newly obtained barcode sequences (with GenBank accession and herbarium voucher numbers) as well as existing sequences sourced from GenBank (with GenBank numbers only) that make up the barcode reference library are provided for matK, ITS and ETS (Supplementary Tables S1–S3).
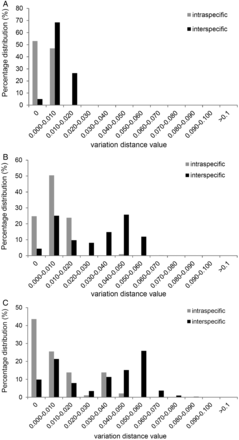
ITS and ETS share similar mean interspecific distance, θ prime and minimum interspecific distance values – both higher than obtained for matK (Table 1). In contrast, in terms of intraspecific variation, matK had higher values than ITS and ETS in all three metrics analysed. The barcoding gap assessments demonstrate that the intra- and interspecific variations of matK and ETS exhibit the most amount of overlap, and ITS the least (Figure 1).
Analysis of inter- and intraspecific divergences in species analysed for barcoding Myrtaceae in New Zealand when including alleles (three loci) and excluding alleles (two loci)
| Divergences . | Including alleles . | Excluding alleles . | |||
|---|---|---|---|---|---|
| matK . | ITS . | ETS . | ITS . | ETS . | |
| Interspecific | |||||
| Mean interspecific distance | 0.0080 ± 0.0048 | 0.0291 ± 0.0196 | 0.0349 ± 0.0232 | 0.0294 ± 0.0192 | 0.0351 ± 0.0241 |
| θ prime | 0.0096 ± 0.0053 | 0.0353 ± 0.0242 | 0.0347 ± 0.0131 | 0.0359 ± 0.0243 | 0.0351 ± 0.0135 |
| Minimum interspecific distance | 0.0042 ± 0.0055 | 0.0179 ± 0.0260 | 0.0098 ± 0.0157 | 0.0181 ± 0.0263 | 0.0097 ± 0.0150 |
| Intraspecific | |||||
| Mean intraspecific distance | 0.0040 ± 0.0158 | 0.0037 ± 0.0056 | 0.0062 ± 0.0119 | 0.0036 ± 0.0054 | 0.0066 ± 0.0123 |
| θ | 0.0043 ± 0.0150 | 0.0030 ± 0.0046 | 0.0026 ± 0.0043 | 0.0032 ± 0.0048 | 0.0029 ± 0.0051 |
| Coalescent depth | 0.0079 ± 0.0279 | 0.0053 ± 0.0075 | 0.0051 ± 0.0087 | 0.0054 ± 0.0074 | 0.0061 ± 0.0123 |
| Divergences . | Including alleles . | Excluding alleles . | |||
|---|---|---|---|---|---|
| matK . | ITS . | ETS . | ITS . | ETS . | |
| Interspecific | |||||
| Mean interspecific distance | 0.0080 ± 0.0048 | 0.0291 ± 0.0196 | 0.0349 ± 0.0232 | 0.0294 ± 0.0192 | 0.0351 ± 0.0241 |
| θ prime | 0.0096 ± 0.0053 | 0.0353 ± 0.0242 | 0.0347 ± 0.0131 | 0.0359 ± 0.0243 | 0.0351 ± 0.0135 |
| Minimum interspecific distance | 0.0042 ± 0.0055 | 0.0179 ± 0.0260 | 0.0098 ± 0.0157 | 0.0181 ± 0.0263 | 0.0097 ± 0.0150 |
| Intraspecific | |||||
| Mean intraspecific distance | 0.0040 ± 0.0158 | 0.0037 ± 0.0056 | 0.0062 ± 0.0119 | 0.0036 ± 0.0054 | 0.0066 ± 0.0123 |
| θ | 0.0043 ± 0.0150 | 0.0030 ± 0.0046 | 0.0026 ± 0.0043 | 0.0032 ± 0.0048 | 0.0029 ± 0.0051 |
| Coalescent depth | 0.0079 ± 0.0279 | 0.0053 ± 0.0075 | 0.0051 ± 0.0087 | 0.0054 ± 0.0074 | 0.0061 ± 0.0123 |
Analysis of inter- and intraspecific divergences in species analysed for barcoding Myrtaceae in New Zealand when including alleles (three loci) and excluding alleles (two loci)
| Divergences . | Including alleles . | Excluding alleles . | |||
|---|---|---|---|---|---|
| matK . | ITS . | ETS . | ITS . | ETS . | |
| Interspecific | |||||
| Mean interspecific distance | 0.0080 ± 0.0048 | 0.0291 ± 0.0196 | 0.0349 ± 0.0232 | 0.0294 ± 0.0192 | 0.0351 ± 0.0241 |
| θ prime | 0.0096 ± 0.0053 | 0.0353 ± 0.0242 | 0.0347 ± 0.0131 | 0.0359 ± 0.0243 | 0.0351 ± 0.0135 |
| Minimum interspecific distance | 0.0042 ± 0.0055 | 0.0179 ± 0.0260 | 0.0098 ± 0.0157 | 0.0181 ± 0.0263 | 0.0097 ± 0.0150 |
| Intraspecific | |||||
| Mean intraspecific distance | 0.0040 ± 0.0158 | 0.0037 ± 0.0056 | 0.0062 ± 0.0119 | 0.0036 ± 0.0054 | 0.0066 ± 0.0123 |
| θ | 0.0043 ± 0.0150 | 0.0030 ± 0.0046 | 0.0026 ± 0.0043 | 0.0032 ± 0.0048 | 0.0029 ± 0.0051 |
| Coalescent depth | 0.0079 ± 0.0279 | 0.0053 ± 0.0075 | 0.0051 ± 0.0087 | 0.0054 ± 0.0074 | 0.0061 ± 0.0123 |
| Divergences . | Including alleles . | Excluding alleles . | |||
|---|---|---|---|---|---|
| matK . | ITS . | ETS . | ITS . | ETS . | |
| Interspecific | |||||
| Mean interspecific distance | 0.0080 ± 0.0048 | 0.0291 ± 0.0196 | 0.0349 ± 0.0232 | 0.0294 ± 0.0192 | 0.0351 ± 0.0241 |
| θ prime | 0.0096 ± 0.0053 | 0.0353 ± 0.0242 | 0.0347 ± 0.0131 | 0.0359 ± 0.0243 | 0.0351 ± 0.0135 |
| Minimum interspecific distance | 0.0042 ± 0.0055 | 0.0179 ± 0.0260 | 0.0098 ± 0.0157 | 0.0181 ± 0.0263 | 0.0097 ± 0.0150 |
| Intraspecific | |||||
| Mean intraspecific distance | 0.0040 ± 0.0158 | 0.0037 ± 0.0056 | 0.0062 ± 0.0119 | 0.0036 ± 0.0054 | 0.0066 ± 0.0123 |
| θ | 0.0043 ± 0.0150 | 0.0030 ± 0.0046 | 0.0026 ± 0.0043 | 0.0032 ± 0.0048 | 0.0029 ± 0.0051 |
| Coalescent depth | 0.0079 ± 0.0279 | 0.0053 ± 0.0075 | 0.0051 ± 0.0087 | 0.0054 ± 0.0074 | 0.0061 ± 0.0123 |
The barcoding gap between interspecific (black) and intraspecific (grey) divergences for the three analysed loci for barcoding Myrtaceae in New Zealand. (A) matK, (B) ITS and (C) ETS.
At the species level, ITS had the highest identification success rate followed by matK and ETS, respectively, as measured by both methods (Table 2). The 90.4 per cent success rate of ITS (ITS1 and ITS2) determined by the BLAST1 method is considerably higher than the 76.1 per cent obtained by Yao et al. (2010) based on an analysis of ITS2 alone and a sample size of 34 676 dicotyledons. Because we sampled multiple samples per species, we were able to identify different alleles of ITS and ETS. Excluding alleles from the calculations did not have a large effect on the divergences and identification efficiency rates (Tables 1 and 2).
Overall identification efficiency (%) of the loci sequenced in species analysed for barcoding Myrtaceae in New Zealand using the BLAST1 and Near Distance methods when including alleles (three loci) or excluding alleles (two loci)
| Method . | Including alleles . | Excluding alleles . | |||
|---|---|---|---|---|---|
| matK . | ITS . | ETS . | ITS . | ETS . | |
| BLAST1 | 81.8 | 90.4 | 76.0 | 90.3 | 75.4 |
| Near distance | 63.1 | 74.7 | 47.5 | 76.3 | 49.1 |
| Method . | Including alleles . | Excluding alleles . | |||
|---|---|---|---|---|---|
| matK . | ITS . | ETS . | ITS . | ETS . | |
| BLAST1 | 81.8 | 90.4 | 76.0 | 90.3 | 75.4 |
| Near distance | 63.1 | 74.7 | 47.5 | 76.3 | 49.1 |
Overall identification efficiency (%) of the loci sequenced in species analysed for barcoding Myrtaceae in New Zealand using the BLAST1 and Near Distance methods when including alleles (three loci) or excluding alleles (two loci)
| Method . | Including alleles . | Excluding alleles . | |||
|---|---|---|---|---|---|
| matK . | ITS . | ETS . | ITS . | ETS . | |
| BLAST1 | 81.8 | 90.4 | 76.0 | 90.3 | 75.4 |
| Near distance | 63.1 | 74.7 | 47.5 | 76.3 | 49.1 |
| Method . | Including alleles . | Excluding alleles . | |||
|---|---|---|---|---|---|
| matK . | ITS . | ETS . | ITS . | ETS . | |
| BLAST1 | 81.8 | 90.4 | 76.0 | 90.3 | 75.4 |
| Near distance | 63.1 | 74.7 | 47.5 | 76.3 | 49.1 |
Of the 43 genera analysed, identification success of specimens to species was 100 per cent for those genera represented by a relatively small number of species in all three analysed loci (Table 3). Kunzea and Eucalyptus, represented by 5 and 27 species, respectively, in matK, had the two lowest overall success rates (20and 22 per cent, respectively) because sequences were shared by more than one species. Success rates in ITS were on average higher in species where matK success rate values were <100 per cent. Metrosideros and Syzygium were the only genera where the success rate of identifying specimens was lower with ITS compared with matK. While the taxon sampling for ETS was small, lower success rates were also present in the more species-rich genera. The 100 per cent success rate of ETS in identifying species of Metrosideros is noteworthy.
Identification efficiency (%) of the three loci per genus analysed for barcoding Myrtaceae in New Zealand using the BLAST1 method
| Genus . | matK . | ITS . | ETS . | ||||||
|---|---|---|---|---|---|---|---|---|---|
| No. of species . | No. of samples . | Success rate at the species level (%) . | No. of species . | No. of samples . | Success rate at the species level (%) . | No. of species . | No. of samples . | Success rate at the species level (%) . | |
| Acca | 1 | 4 | 100 | 1 | 4 | 100 | |||
| Agonis | 1 | 4 | 100 | 1 | 4 | 100 | |||
| Amomyrtus | 1 | 2 | 100 | 2 | 4 | 100 | |||
| Angophora | 3 | 8 | 67 | 3 | 8 | 100 | 2 | 6 | 100 |
| Astartea | 1 | 3 | 100 | 1 | 3 | 100 | |||
| Backhousia | 1 | 3 | 100 | 1 | 3 | 100 | 1 | 2 | 100 |
| Baeckea | 1 | 2 | 100 | 1 | 1 | 100 | |||
| Beaufortia | 1 | 4 | 100 | 1 | 4 | 100 | |||
| Blepharocalyx | – | – | – | 1 | 3 | 100 | |||
| Callistemon | 4 | 15 | 75 | 4 | 18 | 75 | 1 | 2 | 100 |
| Calothamnus | 1 | 3 | 100 | 1 | 3 | 100 | |||
| Calytrix | 1 | 3 | 100 | 1 | 2 | 100 | |||
| Carpolepis | 1 | 1 | 100 | 1 | 1 | 100 | |||
| Chamelaucium | 3 | 7 | 100 | 1 | 3 | 100 | |||
| Corymbia | 4 | 15 | 100 | 4 | 13 | 100 | 4 | 16 | 75 |
| Darwinia | 2 | 4 | 100 | 2 | 4 | 100 | |||
| Eucalyptus | 27 | 96 | 22 | 26 | 105 | 50 | 28 | 87 | 32 |
| Eugenia | 1 | 5 | 100 | 1 | 5 | 100 | |||
| Heteropyxis | 1 | 5 | 100 | 1 | 2 | 100 | |||
| Hypocalymma | 1 | 4 | 100 | 1 | 4 | 100 | |||
| Kunzea | 5 | 15 | 20 | 4 | 19 | 50 | |||
| Leptospermum | 5 | 18 | 80 | 5 | 21 | 100 | 1 | 1 | 100 |
| Lophomyrtus | 2 | 7 | 100 | 2 | 6 | 100 | 1 | 1 | 100 |
| Lophostemon | 1 | 3 | 100 | 1 | 3 | 100 | |||
| Luma | 1 | 3 | 100 | 1 | 3 | 100 | |||
| Melaleuca | 7 | 19 | 100 | 6 | 19 | 100 | |||
| Metrosideros | 14 | 52 | 86 | 14 | 62 | 79 | 14 | 61 | 100 |
| Micromyrtus | 1 | 4 | 100 | 1 | 1 | 100 | |||
| Myrciaria | 1 | 3 | 100 | 1 | 3 | 100 | |||
| Myrteola | 1 | 1 | 100 | 1 | 1 | 100 | |||
| Myrtus | 1 | 4 | 100 | 1 | 12 | 100 | |||
| Neomyrtus | 1 | 3 | 100 | 1 | 5 | 100 | |||
| Pimenta | 1 | 3 | 100 | 1 | 3 | 100 | |||
| Psidium | 1 | 4 | 100 | 1 | 4 | 100 | 1 | 2 | 100 |
| Sannantha | 1 | 3 | 100 | 1 | 2 | 100 | |||
| Syncarpia | 1 | 3 | 100 | 1 | 2 | 100 | |||
| Syzygium | 6 | 22 | 100 | 6 | 19 | 83 | 1 | 1 | 100 |
| Taxandria | 3 | 9 | 33 | 4 | 12 | 100 | |||
| Thaleropia | 1 | 3 | 100 | 1 | 2 | 100 | |||
| Thryptomene | 1 | 3 | 100 | 1 | 4 | 100 | |||
| Tristaniopsis | 1 | 4 | 100 | 1 | 3 | 100 | |||
| Ugni | 1 | 3 | 100 | 1 | 3 | 100 | |||
| Xanthostemon | 1 | 3 | 100 | 1 | 4 | 100 | |||
| Genus . | matK . | ITS . | ETS . | ||||||
|---|---|---|---|---|---|---|---|---|---|
| No. of species . | No. of samples . | Success rate at the species level (%) . | No. of species . | No. of samples . | Success rate at the species level (%) . | No. of species . | No. of samples . | Success rate at the species level (%) . | |
| Acca | 1 | 4 | 100 | 1 | 4 | 100 | |||
| Agonis | 1 | 4 | 100 | 1 | 4 | 100 | |||
| Amomyrtus | 1 | 2 | 100 | 2 | 4 | 100 | |||
| Angophora | 3 | 8 | 67 | 3 | 8 | 100 | 2 | 6 | 100 |
| Astartea | 1 | 3 | 100 | 1 | 3 | 100 | |||
| Backhousia | 1 | 3 | 100 | 1 | 3 | 100 | 1 | 2 | 100 |
| Baeckea | 1 | 2 | 100 | 1 | 1 | 100 | |||
| Beaufortia | 1 | 4 | 100 | 1 | 4 | 100 | |||
| Blepharocalyx | – | – | – | 1 | 3 | 100 | |||
| Callistemon | 4 | 15 | 75 | 4 | 18 | 75 | 1 | 2 | 100 |
| Calothamnus | 1 | 3 | 100 | 1 | 3 | 100 | |||
| Calytrix | 1 | 3 | 100 | 1 | 2 | 100 | |||
| Carpolepis | 1 | 1 | 100 | 1 | 1 | 100 | |||
| Chamelaucium | 3 | 7 | 100 | 1 | 3 | 100 | |||
| Corymbia | 4 | 15 | 100 | 4 | 13 | 100 | 4 | 16 | 75 |
| Darwinia | 2 | 4 | 100 | 2 | 4 | 100 | |||
| Eucalyptus | 27 | 96 | 22 | 26 | 105 | 50 | 28 | 87 | 32 |
| Eugenia | 1 | 5 | 100 | 1 | 5 | 100 | |||
| Heteropyxis | 1 | 5 | 100 | 1 | 2 | 100 | |||
| Hypocalymma | 1 | 4 | 100 | 1 | 4 | 100 | |||
| Kunzea | 5 | 15 | 20 | 4 | 19 | 50 | |||
| Leptospermum | 5 | 18 | 80 | 5 | 21 | 100 | 1 | 1 | 100 |
| Lophomyrtus | 2 | 7 | 100 | 2 | 6 | 100 | 1 | 1 | 100 |
| Lophostemon | 1 | 3 | 100 | 1 | 3 | 100 | |||
| Luma | 1 | 3 | 100 | 1 | 3 | 100 | |||
| Melaleuca | 7 | 19 | 100 | 6 | 19 | 100 | |||
| Metrosideros | 14 | 52 | 86 | 14 | 62 | 79 | 14 | 61 | 100 |
| Micromyrtus | 1 | 4 | 100 | 1 | 1 | 100 | |||
| Myrciaria | 1 | 3 | 100 | 1 | 3 | 100 | |||
| Myrteola | 1 | 1 | 100 | 1 | 1 | 100 | |||
| Myrtus | 1 | 4 | 100 | 1 | 12 | 100 | |||
| Neomyrtus | 1 | 3 | 100 | 1 | 5 | 100 | |||
| Pimenta | 1 | 3 | 100 | 1 | 3 | 100 | |||
| Psidium | 1 | 4 | 100 | 1 | 4 | 100 | 1 | 2 | 100 |
| Sannantha | 1 | 3 | 100 | 1 | 2 | 100 | |||
| Syncarpia | 1 | 3 | 100 | 1 | 2 | 100 | |||
| Syzygium | 6 | 22 | 100 | 6 | 19 | 83 | 1 | 1 | 100 |
| Taxandria | 3 | 9 | 33 | 4 | 12 | 100 | |||
| Thaleropia | 1 | 3 | 100 | 1 | 2 | 100 | |||
| Thryptomene | 1 | 3 | 100 | 1 | 4 | 100 | |||
| Tristaniopsis | 1 | 4 | 100 | 1 | 3 | 100 | |||
| Ugni | 1 | 3 | 100 | 1 | 3 | 100 | |||
| Xanthostemon | 1 | 3 | 100 | 1 | 4 | 100 | |||
Calculations based on a subset of sequences provided in Supplementary Tables S1–S4 due to partial sequences and sequences with ambiguous bases being excluded.
Identification efficiency (%) of the three loci per genus analysed for barcoding Myrtaceae in New Zealand using the BLAST1 method
| Genus . | matK . | ITS . | ETS . | ||||||
|---|---|---|---|---|---|---|---|---|---|
| No. of species . | No. of samples . | Success rate at the species level (%) . | No. of species . | No. of samples . | Success rate at the species level (%) . | No. of species . | No. of samples . | Success rate at the species level (%) . | |
| Acca | 1 | 4 | 100 | 1 | 4 | 100 | |||
| Agonis | 1 | 4 | 100 | 1 | 4 | 100 | |||
| Amomyrtus | 1 | 2 | 100 | 2 | 4 | 100 | |||
| Angophora | 3 | 8 | 67 | 3 | 8 | 100 | 2 | 6 | 100 |
| Astartea | 1 | 3 | 100 | 1 | 3 | 100 | |||
| Backhousia | 1 | 3 | 100 | 1 | 3 | 100 | 1 | 2 | 100 |
| Baeckea | 1 | 2 | 100 | 1 | 1 | 100 | |||
| Beaufortia | 1 | 4 | 100 | 1 | 4 | 100 | |||
| Blepharocalyx | – | – | – | 1 | 3 | 100 | |||
| Callistemon | 4 | 15 | 75 | 4 | 18 | 75 | 1 | 2 | 100 |
| Calothamnus | 1 | 3 | 100 | 1 | 3 | 100 | |||
| Calytrix | 1 | 3 | 100 | 1 | 2 | 100 | |||
| Carpolepis | 1 | 1 | 100 | 1 | 1 | 100 | |||
| Chamelaucium | 3 | 7 | 100 | 1 | 3 | 100 | |||
| Corymbia | 4 | 15 | 100 | 4 | 13 | 100 | 4 | 16 | 75 |
| Darwinia | 2 | 4 | 100 | 2 | 4 | 100 | |||
| Eucalyptus | 27 | 96 | 22 | 26 | 105 | 50 | 28 | 87 | 32 |
| Eugenia | 1 | 5 | 100 | 1 | 5 | 100 | |||
| Heteropyxis | 1 | 5 | 100 | 1 | 2 | 100 | |||
| Hypocalymma | 1 | 4 | 100 | 1 | 4 | 100 | |||
| Kunzea | 5 | 15 | 20 | 4 | 19 | 50 | |||
| Leptospermum | 5 | 18 | 80 | 5 | 21 | 100 | 1 | 1 | 100 |
| Lophomyrtus | 2 | 7 | 100 | 2 | 6 | 100 | 1 | 1 | 100 |
| Lophostemon | 1 | 3 | 100 | 1 | 3 | 100 | |||
| Luma | 1 | 3 | 100 | 1 | 3 | 100 | |||
| Melaleuca | 7 | 19 | 100 | 6 | 19 | 100 | |||
| Metrosideros | 14 | 52 | 86 | 14 | 62 | 79 | 14 | 61 | 100 |
| Micromyrtus | 1 | 4 | 100 | 1 | 1 | 100 | |||
| Myrciaria | 1 | 3 | 100 | 1 | 3 | 100 | |||
| Myrteola | 1 | 1 | 100 | 1 | 1 | 100 | |||
| Myrtus | 1 | 4 | 100 | 1 | 12 | 100 | |||
| Neomyrtus | 1 | 3 | 100 | 1 | 5 | 100 | |||
| Pimenta | 1 | 3 | 100 | 1 | 3 | 100 | |||
| Psidium | 1 | 4 | 100 | 1 | 4 | 100 | 1 | 2 | 100 |
| Sannantha | 1 | 3 | 100 | 1 | 2 | 100 | |||
| Syncarpia | 1 | 3 | 100 | 1 | 2 | 100 | |||
| Syzygium | 6 | 22 | 100 | 6 | 19 | 83 | 1 | 1 | 100 |
| Taxandria | 3 | 9 | 33 | 4 | 12 | 100 | |||
| Thaleropia | 1 | 3 | 100 | 1 | 2 | 100 | |||
| Thryptomene | 1 | 3 | 100 | 1 | 4 | 100 | |||
| Tristaniopsis | 1 | 4 | 100 | 1 | 3 | 100 | |||
| Ugni | 1 | 3 | 100 | 1 | 3 | 100 | |||
| Xanthostemon | 1 | 3 | 100 | 1 | 4 | 100 | |||
| Genus . | matK . | ITS . | ETS . | ||||||
|---|---|---|---|---|---|---|---|---|---|
| No. of species . | No. of samples . | Success rate at the species level (%) . | No. of species . | No. of samples . | Success rate at the species level (%) . | No. of species . | No. of samples . | Success rate at the species level (%) . | |
| Acca | 1 | 4 | 100 | 1 | 4 | 100 | |||
| Agonis | 1 | 4 | 100 | 1 | 4 | 100 | |||
| Amomyrtus | 1 | 2 | 100 | 2 | 4 | 100 | |||
| Angophora | 3 | 8 | 67 | 3 | 8 | 100 | 2 | 6 | 100 |
| Astartea | 1 | 3 | 100 | 1 | 3 | 100 | |||
| Backhousia | 1 | 3 | 100 | 1 | 3 | 100 | 1 | 2 | 100 |
| Baeckea | 1 | 2 | 100 | 1 | 1 | 100 | |||
| Beaufortia | 1 | 4 | 100 | 1 | 4 | 100 | |||
| Blepharocalyx | – | – | – | 1 | 3 | 100 | |||
| Callistemon | 4 | 15 | 75 | 4 | 18 | 75 | 1 | 2 | 100 |
| Calothamnus | 1 | 3 | 100 | 1 | 3 | 100 | |||
| Calytrix | 1 | 3 | 100 | 1 | 2 | 100 | |||
| Carpolepis | 1 | 1 | 100 | 1 | 1 | 100 | |||
| Chamelaucium | 3 | 7 | 100 | 1 | 3 | 100 | |||
| Corymbia | 4 | 15 | 100 | 4 | 13 | 100 | 4 | 16 | 75 |
| Darwinia | 2 | 4 | 100 | 2 | 4 | 100 | |||
| Eucalyptus | 27 | 96 | 22 | 26 | 105 | 50 | 28 | 87 | 32 |
| Eugenia | 1 | 5 | 100 | 1 | 5 | 100 | |||
| Heteropyxis | 1 | 5 | 100 | 1 | 2 | 100 | |||
| Hypocalymma | 1 | 4 | 100 | 1 | 4 | 100 | |||
| Kunzea | 5 | 15 | 20 | 4 | 19 | 50 | |||
| Leptospermum | 5 | 18 | 80 | 5 | 21 | 100 | 1 | 1 | 100 |
| Lophomyrtus | 2 | 7 | 100 | 2 | 6 | 100 | 1 | 1 | 100 |
| Lophostemon | 1 | 3 | 100 | 1 | 3 | 100 | |||
| Luma | 1 | 3 | 100 | 1 | 3 | 100 | |||
| Melaleuca | 7 | 19 | 100 | 6 | 19 | 100 | |||
| Metrosideros | 14 | 52 | 86 | 14 | 62 | 79 | 14 | 61 | 100 |
| Micromyrtus | 1 | 4 | 100 | 1 | 1 | 100 | |||
| Myrciaria | 1 | 3 | 100 | 1 | 3 | 100 | |||
| Myrteola | 1 | 1 | 100 | 1 | 1 | 100 | |||
| Myrtus | 1 | 4 | 100 | 1 | 12 | 100 | |||
| Neomyrtus | 1 | 3 | 100 | 1 | 5 | 100 | |||
| Pimenta | 1 | 3 | 100 | 1 | 3 | 100 | |||
| Psidium | 1 | 4 | 100 | 1 | 4 | 100 | 1 | 2 | 100 |
| Sannantha | 1 | 3 | 100 | 1 | 2 | 100 | |||
| Syncarpia | 1 | 3 | 100 | 1 | 2 | 100 | |||
| Syzygium | 6 | 22 | 100 | 6 | 19 | 83 | 1 | 1 | 100 |
| Taxandria | 3 | 9 | 33 | 4 | 12 | 100 | |||
| Thaleropia | 1 | 3 | 100 | 1 | 2 | 100 | |||
| Thryptomene | 1 | 3 | 100 | 1 | 4 | 100 | |||
| Tristaniopsis | 1 | 4 | 100 | 1 | 3 | 100 | |||
| Ugni | 1 | 3 | 100 | 1 | 3 | 100 | |||
| Xanthostemon | 1 | 3 | 100 | 1 | 4 | 100 | |||
Calculations based on a subset of sequences provided in Supplementary Tables S1–S4 due to partial sequences and sequences with ambiguous bases being excluded.
Discussion
The main aim of DNA barcoding is to establish a shared community resource of DNA sequences that can be used for organismal identification and taxonomic clarification (Hollingsworth et al., 2011). Unlike taxon- or clade-based approaches where all the species of a clade are barcoded, e.g. Seberg and Petersen (2009), others follow a floristic approach by targeting a variety of species in a particular area, e.g. de Vere et al. (2012). Our DNA barcode reference library encompasses both approaches. We focussed on the Myrtaceae occurring in New Zealand and therefore did not barcode all the taxa in the Myrtaceae clade. However, we targeted all native species and ensured at least all known exotic genera of Myrtaceae in New Zealand were represented in the reference library with particular emphasis on species commonly utilized in the forestry industry. The choice of loci ensures that our reference library is scalable and it can accommodate improved taxon sampling in the future.
Despite the near-universal usage of ITS sequence data in plant phylogenetic studies, its complex and unpredictable evolutionary behaviour can reduce its utility for phylogenetic or barcoding analyses (Álvarez and Wendel, 2003). While the extent of paralogy, polymorphic sites and sequence quality can be quantified (Bailey et al., 2003), it has been shown that where the presence of rDNA pseudogenes has gone undetected in previous studies of eucalypt phylogeny, the impact on reconstructions of taxon relationships was minimal where taxa were sampled sparsely (Bayly et al., 2008). There are Myrtaceae datasets that are free of pseudogenes (Steane et al., 2002; Murillo-Aldana et al., 2011), and there are those that are not (Bayly et al., 2008). One method to initially determine the presence of pseudogenes is to visually search for unusual sequences in the 5.8S rRNA gene (Bayly et al., 2008). An inspection of our entire ITS dataset revealed no unusual 5.8S rRNA sequences comparable with those reported by Bayly et al. (2008).
Like all other barcoding projects, we present relative rather than absolute discriminatory power of different barcoding regions because our taxon sampling is not complete for the Myrtaceae. Furthermore, different taxonomic opinions exist and can influence the taxon sampling and hence efficacy values. The efficacy (accuracy) of identifying species via barcoding data is reliant on the extent of, and separation between, intra- and interspecific divergences in the selected loci for a given set of taxa. The more overlap there is between genetic variation within species and genetic variation between related species, the less effective barcoding becomes. In our analysis, although ITS and ETS share similar average inter- and intraspecific divergences compared with matK, ITS possesses the largest gap in the distribution of intra- and interspecific divergences. ITS also outperforms matK and ETS in terms of overall identification efficiency. Overall, identification success increases when a combination of ITS and matK, in particular, is used in a reasoned way.
Besides the generated DNA barcode data, a benefit of our work has been the deposition of voucher specimens in Scion's National Forestry Herbarium; this improved resource will assist future identification of Myrtaceae based on morphology. Additionally, 22 existing herbarium specimens that were sampled for barcoding purposes and 8 existing GenBank sequences have been redetermined as a result of our work (M.H. Buys, unpublished data). No new definitive phylogenetic insights were gleaned from the DNA barcode data due to only a subsample of species per genus having being sampled and because only matK and ITS (and to a limited extent ETS) were analysed.
The Myrtaceae DNA barcode database will enable rapid identification of large numbers of myrtaceous specimens in the event of a myrtle rust incursion. This will help MPI to accurately survey the spread of the disease and thereby enable timely decision making and consideration of biosecurity measures in order to protect threatened plant species and reduce the risk of spread to new areas. As such, the Myrtaceae DNA barcode database is a national botanical resource.
Conflict of interest statement
None declared.
Funding
We acknowledge funding from New Zealand's Ministry of Business, Innovation and Employment core funding to Scion and the Ministry for Primary Industries’ Operational Research Programme through the Barcoding Myrtaceae-16209 contract.
Acknowledgements
The following nurseries and gardens are thanked for supplying material of lesser-known species in New Zealand: Te Horo Nurseries, Masons Garden Centre and Nurseries, Oratia Native Plant Nursery, Global Plants Garden Centre, Bason Botanic Gardens, Eastwoodhill Arboretum, Hamilton Gardens, Auckland Domain Nursery and Auckland Botanic Gardens, University of Auckland Grounds Department, Pukekura Park, Wellington Botanic Garden; in Australia: Kings Park, Perth and Royal Botanic Gardens, Sydney; in South Africa: Kirstenbosch Botanical Gardens. The following people assisted in obtaining plant material and their contribution is acknowledged: John Adam, Jessie Balding, Ian and Jocelyn Bell, Colin Bradshaw, Benny Bytebier, Bill Clarkson, Russell Fransham, Keith Hammett, Chris Inglis, Colin Ogle, Barbara Parris, John Rogers, Ernst van Jaarsveld, Mike Wilcox and Peter Wilson.
References