-
PDF
- Split View
-
Views
-
Cite
Cite
R. Aflatoonian, E. Tuckerman, S.L. Elliott, C. Bruce, A. Aflatoonian, T.C. Li, A. Fazeli, Menstrual cycle-dependent changes of Toll-like receptors in endometrium, Human Reproduction, Volume 22, Issue 2, 1 February 2007, Pages 586–593, https://doi.org/10.1093/humrep/del388
Close - Share Icon Share
Abstract
BACKGROUND: Rapid innate immune defences against infection usually involve the recognition of invading pathogens by specific pattern recognition receptors recently attributed to the family of Toll-like receptors (TLRs). Reports from our laboratory and others have demonstrated the existence of TLRs 1–6 in the female reproductive tract. However, little has been done to identify TLRs 7–10 in the female reproductive tract, particularly in the uterus. Also little information exists regarding variation in TLRs in the female reproductive tract during the menstrual cycle. METHOD: The distribution of TLR7–10 protein was detected by immunostaining in timed endometrial biopsies from normal women. RT–PCR was used to show the existence of TLR1–10 genes in endometrial tissue and real-time PCR analysis to investigate the relative expression of these genes during the menstrual cycle in normal human endometrium. RESULTS: TLR7–10 proteins were detected in endometrial epithelium and stroma. TLR1–10 genes were expressed in human endometrial tissue, and the mean relative expression of TLR2–6, 9 and 10 genes was significantly higher during the secretory phase compared with other phases of the menstrual cycle. CONCLUSIONS: TLR7–10 localization is not limited to endometrial epithelium but is also present in the stroma of the endometrial tissue. Endometrial TLR2–6, 9 and 10 genes are cyclically expressed during the menstrual cycle.
Introduction
Sexually transmitted diseases (STDs) are a major worldwide health problem that compromise reproductive fecundity as well as cut short the lives of millions of men, women and children (Cates, 1986; Piot et al., 1988). Despite advances in the management of these infections, only limited success has been achieved in curtailing the morbidity and mortality associated with them. The commoner STDs with the largest health and socio-economic impact include infections by the herpes simplex virus type 2 (HSV-2), Chlamydia trachomatis, the gonococcus and human immunodeficiency viruses (HIVs). In addition, maternal genital tract carriage of the group B streptococcus and bacterial vaginosis seems to be associated with premature birth and neonatal mortality and morbidity (Martius and Eschenbach, 1990). Because of this growing global health problem, prevention and effective treatment methodologies need to be developed. Understanding the mechanisms that regulate the female reproductive tract immune system holds therapeutic promise. If the immunological host defence mechanisms against these infections are clarified, effective vaccines can be developed.
Rapid innate immune defences against infection usually involve the recognition of invading pathogens by specific pattern recognition receptors recently attributed to the family of Toll-like receptors (TLRs) (Medzhitov and Janeway, 2000; Janeway and Medzhitov, 2002). TLRs are expressed by cells involved in the first line of host defence, including neutrophils, macrophages, dendritic cells, dermal endothelial cells and mucosal epithelial cells. Collectively, TLRs function to alert the immune system to the presence of micro-organisms. Different members of the TLR family are expressed on different cell organelles and appear to mediate signal transduction to a range of antigenic stimuli by engaging with specific ligands leading to the production of various proinflammatory cytokines, chemokines and effector molecules, depending on the cell type that is activated. The members of the TLR family, of which at least 11 have been identified, recognize distinct pathogen-associated molecular patterns (PAMPs) produced by various bacterial, fungal and viral pathogens. TLR2 forms heterodimers with TLRs 1 and 6 and recognizes a broad range of microbial products from Gram-positive bacteria (peptidoglycan) (Schwandner et al., 1999), fungi (zymosan) (Underhill et al., 1999) and synthetic lipoproteins (Pam3Cys-Ser-(Lys)4) (Takeuchi et al., 2002). TLR3 recognizes double-stranded RNA of viral or cellular origin (Alexopoulou et al., 2001; Kariko et al., 2004). The major component of the outer membrane of Gram-negative bacteria, lipopolysaccharide (LPS), is recognized by TLR4 in association with CD14 and MD-2 (Akashi et al., 2001; da Silva Correia et al., 2001; Nagai et al., 2002). TLR5 recognizes bacterial flagellin (Hayashi et al., 2001), TLRs 7 and 8 recognize single-stranded RNA (ssRNA) and synthetic nucleotide derivatives (Hemmi et al., 2002; Heil et al., 2004), and TLR9 recognizes non-methylated CpG containing DNA (Hemmi et al., 2000). No specific ligand has yet been identified for TLR10 (Chuang and Ulevitch, 2001). Studies in mice have shown TLR11 binds to a profilin-like protein from Toxoplasma gondii (Yarovinsky et al., 2005; Yarovinsky and Sher, 2006).
Several reports exist on the determination and characterization of TLRs in different tissues and organs (Bsibsi et al., 2002; Zarember and Godowski, 2002; Backhed and Hornef, 2003; Basu and Fenton, 2004). However, little has been done to identify TLRs in the human female. Recently, we demonstrated the in vivo distribution of TLRs 1–6 in the human female reproductive tract using immunohistochemical techniques (Fazeli et al., 2005). With the exception of TLR4, all other TLRs studied were uniformly distributed throughout the tract. TLR7–10 mRNA has been shown to be expressed in human uterine tissue (Nishimura and Naito, 2005). TLRs have been predominantly described in the epithelial cells of the human endometrium. Primary human uterine epithelial cell cultures and the uterine epithelial cell line ECC-1 express TLR1–9 genes (Schaefer et al., 2004, 2005). However, in contrast to the studies utilizing cultured cells, Young et al. (2004) could not detect the expression of TLR7, 8 and 10 genes in endometrial tissue samples. No information exists regarding the potential immunohistochemical localization of TLR7–10 in the female reproductive tract and in particular in the endometrium in vivo. In the present investigation, we report the gene expression and the in vivo localization of TLR7–10 molecules in healthy human endometrial tissue biopsies.
Although reports from our laboratory and others have demonstrated the existence of TLRs in the female reproductive tract, little information exists regarding variation in TLR presence in the female reproductive tract during the menstrual cycle. Recently, Jorgenson et al. (2005) demonstrated the cycle-dependent expression of TLR3 in primary endometrial epithelial tissue. One can hypothesize that other TLR molecules may have a cycle-dependent expression in the endometrial tissue as well. We tested this hypothesis, and here, we report the alteration in the expression of TLR1–10 genes during the menstrual cycle in normal human endometrial tissue.
Materials and methods
Tissue collection for immunostaining and genomic investigations
This investigation was approved by the Local Ethics Committee, and written informed consent was obtained prior to the collection of tissue samples. For immunohistochemical investigations, tissue samples were obtained from six fertile women, and for genomic studies, endometrial biopsies were obtained from 21 fertile women. All the women taking part in the investigation had regular cycles, showed no evidence of any pathological uterine disorder and had not used oral contraception or an intrauterine device in the previous 3 months. Biopsies were obtained in the operating theater between 1 and 29 days after the last menstrual period (LMP). The mean age of the women taking part in the study was 35 (range 24–40) years, and each had had at least one previous successful pregnancy. Endometrial biopsies for immunocytochemistry were immediately snap-frozen and stored in liquid nitrogen until processed. Cryosections were cut at 5 mm and stored at −70°C until use. For genomic studies, endometrial biopsies were immediately placed in RNAlater (Ambion, Huntingdon, UK) followed by immediate immersion in liquid nitrogen until processed.
Antibodies and peptides
Antibodies and peptides used in the experiments were obtained from Santa Cruz Biotechnology (CA, USA). These were goat polyclonal antibodies specific for N-terminal domains of TLRs 7 and 9 (catalogue number sc13207 and sc13212, respectively), goat polyclonal antibody specific for V-terminal domains of TLR10 (catalogue number sc23577) and rabbit polyclonal specific for D-terminal domains of TLR8 (catalogue number sc13212-R). Blocking peptides specific for the respective antibodies were used to detect non-specific staining.
Immunostaining
Cryosections were removed from −70°C freezer, fixed in 4% paraformaldehyde for 15 min, washed twice in phosphate-buffered saline (PBS) for 5 min and then immersed in methanol at −20°C for 4 min followed by 2 min in acetone at −20°C before being finally washed in PBS. Cryopreserved slides were timed according to LMP and morphology and divided into three groups (menstrual, proliferative or secretory phase). Slides were stained using a Vectorstain Elite ABC peroxidase kit, according to the manufacturer’s instructions (Vector Laboratories, Peterborough, UK). To avoid the non-specific binding of biotin, we used an avidin/biotin blocking kit (Vector Laboratories). Briefly, slides were blocked for 1 h at room temperature in PBS containing 0.2% appropriate serum and 250 µl/ml of avidin. The block was removed, and slides were incubated overnight at 4°C in primary antibody at an appropriate dilution using antibody diluent media (Dakocytomation, Ely, UK) containing 250 µl/ml of biotin. Binding was visualized by incubation with peroxidase substrate 3-amino-9-ethylcarbazole (AEC) (Vector Laboratories) for 10 min, washed in distilled water for 3 min and counterstained in 10% haematoxylin for 10 min. Slides were washed in tap water for 2 min and mounted with Aquamount (VWR).
Optimum staining was achieved by incubating tissue sections with 10 µg/ml of the specific TLR antibody. Negative control sections were obtained by blocking of primary antibody with its specific peptide. Immunostained sections were examined using an Olympus BH2 microscope (Olympus, London, UK).
RNA isolation, cDNA production and quantitative PCR
Tissues were removed from RNAlater and homogenized in 3 ml of TRI reagent (Sigma, Pool, UK) using an Ultra Turrax homogenizer (VWR) for ∼2 min. Total RNA was extracted using TRI reagent standard protocol supplied by the manufacturer. Total RNA was treated with DNase I (DNA-free Kit; Ambion) to remove genomic DNA contamination from samples. First-strand cDNA synthesis was performed using oligo dT primers and the Superscript II reverse transcriptase system (Invitrogen, Paisley, UK).
Reverse-transcription polymerase chain reaction (RT–PCR) was performed using the prepared cDNA, primers for TLRs 1–10 (Table I) and Platinum Blue PCR Super Mix (Invitrogen) under the following conditions: 40 cycles at 95°C for 30 s, 59–70°C for 1 min (different temperature for different TLRs) and 72°C for 2 min.
Sequence of primers
| Gene . | Forward primer (5′-3′) . | Reverse primer (5′-3′) . | Annealing temperature (°C) . | Accession number . | Product size (bp) . | Reference . |
|---|---|---|---|---|---|---|
| TLR1 | GGGTCAGCTGGACTTCAGAG | AAAATCCAAATGCAGGAACG | 61 | U88540.1 | 250 | |
| TLR2 | FTCGGAGTTCTCCCAGTTCTCT | TCCAGTGCTTCAACCCACAA | 59 | AF051152.1 | 175 | |
| TLR3 | CGGGCCAGCTTTCAGGAACCTG | GGCATGAATTATATATGCTGC | 59 | U88879.1 | 400 | Schaefer et al., 2004 |
| TLR4 | CGTGGAGACTTGGCCCTAAA | TTCACACCTGGATAAATCCAGC | 59 | U88880.1 | 301 | |
| TLR5 | CCTCATGACCATCCTCAC AGTCAC | GGCTTCAAGGCACCAGCCATCTC | 65 | AF051151.1 | 355 | Schaefer et al., 2004 |
| TLR6 | CCAAGTGAACATATCAGTTAATACTTTAGGGTGC | CTCAGAAAACACGGTGTACAAAGCTG | 63 | AB020807.1 | 358 | Schaefer et al., 2004 |
| TLR7 | CCTTGAGGCCAACAACATCT | GTAGGGACGGCTGTGACATT | 65 | AF240467.1 | 285 | |
| TLR8 | GTCCTGGGGATCAAAGAGG GAAGAG | CTCTTACAGATCCGCTGCCGTAGCC | 63 | AK075117.1 | 581 | Schaefer et al., 2004 |
| TLR9 | GCGAGATGAGGATGCCCTG CCCTACG | TTCGGCCGTGGGTCCCTGGCAGAAG | 70 | AY359085.1 | 510 | Schaefer et al., 2004 |
| TLR10 | CAGAGGTCATGATGGTTG GATGG | GACCTAGCATCCTGAGATACCAGGGCAG | 63 | AY358300.1 | 256 | Schaefer et al., 2004 |
| β-Actin | GCCAGCTCACCATGGATCAT | CAAACATGATCTGGGTCATCTTC | 59 | BC013380.2 | 384 |
| Gene . | Forward primer (5′-3′) . | Reverse primer (5′-3′) . | Annealing temperature (°C) . | Accession number . | Product size (bp) . | Reference . |
|---|---|---|---|---|---|---|
| TLR1 | GGGTCAGCTGGACTTCAGAG | AAAATCCAAATGCAGGAACG | 61 | U88540.1 | 250 | |
| TLR2 | FTCGGAGTTCTCCCAGTTCTCT | TCCAGTGCTTCAACCCACAA | 59 | AF051152.1 | 175 | |
| TLR3 | CGGGCCAGCTTTCAGGAACCTG | GGCATGAATTATATATGCTGC | 59 | U88879.1 | 400 | Schaefer et al., 2004 |
| TLR4 | CGTGGAGACTTGGCCCTAAA | TTCACACCTGGATAAATCCAGC | 59 | U88880.1 | 301 | |
| TLR5 | CCTCATGACCATCCTCAC AGTCAC | GGCTTCAAGGCACCAGCCATCTC | 65 | AF051151.1 | 355 | Schaefer et al., 2004 |
| TLR6 | CCAAGTGAACATATCAGTTAATACTTTAGGGTGC | CTCAGAAAACACGGTGTACAAAGCTG | 63 | AB020807.1 | 358 | Schaefer et al., 2004 |
| TLR7 | CCTTGAGGCCAACAACATCT | GTAGGGACGGCTGTGACATT | 65 | AF240467.1 | 285 | |
| TLR8 | GTCCTGGGGATCAAAGAGG GAAGAG | CTCTTACAGATCCGCTGCCGTAGCC | 63 | AK075117.1 | 581 | Schaefer et al., 2004 |
| TLR9 | GCGAGATGAGGATGCCCTG CCCTACG | TTCGGCCGTGGGTCCCTGGCAGAAG | 70 | AY359085.1 | 510 | Schaefer et al., 2004 |
| TLR10 | CAGAGGTCATGATGGTTG GATGG | GACCTAGCATCCTGAGATACCAGGGCAG | 63 | AY358300.1 | 256 | Schaefer et al., 2004 |
| β-Actin | GCCAGCTCACCATGGATCAT | CAAACATGATCTGGGTCATCTTC | 59 | BC013380.2 | 384 |
Those data obtained from other reports are referenced.
Sequence of primers
| Gene . | Forward primer (5′-3′) . | Reverse primer (5′-3′) . | Annealing temperature (°C) . | Accession number . | Product size (bp) . | Reference . |
|---|---|---|---|---|---|---|
| TLR1 | GGGTCAGCTGGACTTCAGAG | AAAATCCAAATGCAGGAACG | 61 | U88540.1 | 250 | |
| TLR2 | FTCGGAGTTCTCCCAGTTCTCT | TCCAGTGCTTCAACCCACAA | 59 | AF051152.1 | 175 | |
| TLR3 | CGGGCCAGCTTTCAGGAACCTG | GGCATGAATTATATATGCTGC | 59 | U88879.1 | 400 | Schaefer et al., 2004 |
| TLR4 | CGTGGAGACTTGGCCCTAAA | TTCACACCTGGATAAATCCAGC | 59 | U88880.1 | 301 | |
| TLR5 | CCTCATGACCATCCTCAC AGTCAC | GGCTTCAAGGCACCAGCCATCTC | 65 | AF051151.1 | 355 | Schaefer et al., 2004 |
| TLR6 | CCAAGTGAACATATCAGTTAATACTTTAGGGTGC | CTCAGAAAACACGGTGTACAAAGCTG | 63 | AB020807.1 | 358 | Schaefer et al., 2004 |
| TLR7 | CCTTGAGGCCAACAACATCT | GTAGGGACGGCTGTGACATT | 65 | AF240467.1 | 285 | |
| TLR8 | GTCCTGGGGATCAAAGAGG GAAGAG | CTCTTACAGATCCGCTGCCGTAGCC | 63 | AK075117.1 | 581 | Schaefer et al., 2004 |
| TLR9 | GCGAGATGAGGATGCCCTG CCCTACG | TTCGGCCGTGGGTCCCTGGCAGAAG | 70 | AY359085.1 | 510 | Schaefer et al., 2004 |
| TLR10 | CAGAGGTCATGATGGTTG GATGG | GACCTAGCATCCTGAGATACCAGGGCAG | 63 | AY358300.1 | 256 | Schaefer et al., 2004 |
| β-Actin | GCCAGCTCACCATGGATCAT | CAAACATGATCTGGGTCATCTTC | 59 | BC013380.2 | 384 |
| Gene . | Forward primer (5′-3′) . | Reverse primer (5′-3′) . | Annealing temperature (°C) . | Accession number . | Product size (bp) . | Reference . |
|---|---|---|---|---|---|---|
| TLR1 | GGGTCAGCTGGACTTCAGAG | AAAATCCAAATGCAGGAACG | 61 | U88540.1 | 250 | |
| TLR2 | FTCGGAGTTCTCCCAGTTCTCT | TCCAGTGCTTCAACCCACAA | 59 | AF051152.1 | 175 | |
| TLR3 | CGGGCCAGCTTTCAGGAACCTG | GGCATGAATTATATATGCTGC | 59 | U88879.1 | 400 | Schaefer et al., 2004 |
| TLR4 | CGTGGAGACTTGGCCCTAAA | TTCACACCTGGATAAATCCAGC | 59 | U88880.1 | 301 | |
| TLR5 | CCTCATGACCATCCTCAC AGTCAC | GGCTTCAAGGCACCAGCCATCTC | 65 | AF051151.1 | 355 | Schaefer et al., 2004 |
| TLR6 | CCAAGTGAACATATCAGTTAATACTTTAGGGTGC | CTCAGAAAACACGGTGTACAAAGCTG | 63 | AB020807.1 | 358 | Schaefer et al., 2004 |
| TLR7 | CCTTGAGGCCAACAACATCT | GTAGGGACGGCTGTGACATT | 65 | AF240467.1 | 285 | |
| TLR8 | GTCCTGGGGATCAAAGAGG GAAGAG | CTCTTACAGATCCGCTGCCGTAGCC | 63 | AK075117.1 | 581 | Schaefer et al., 2004 |
| TLR9 | GCGAGATGAGGATGCCCTG CCCTACG | TTCGGCCGTGGGTCCCTGGCAGAAG | 70 | AY359085.1 | 510 | Schaefer et al., 2004 |
| TLR10 | CAGAGGTCATGATGGTTG GATGG | GACCTAGCATCCTGAGATACCAGGGCAG | 63 | AY358300.1 | 256 | Schaefer et al., 2004 |
| β-Actin | GCCAGCTCACCATGGATCAT | CAAACATGATCTGGGTCATCTTC | 59 | BC013380.2 | 384 |
Those data obtained from other reports are referenced.
Quantitative real-time PCR was performed using the prepared cDNA and primers for TLRs 1–10 and human β-actin. The forward and reverse primer sequences used are depicted in Table I. All experiments included negative controls with no cDNA. SYBR Green Jump Start (Sigma) master mix was added to each well of the PCR plate (10 µl of SYBR Green, 7 µl of water, 2 µl of primers and 1 µl of cDNA), and PCR was performed under the following conditions: 50 cycles at 95°C for 30 s, 59–70°C for 30 s (different temperature for different TLRs) and 72°C for 30 s. Samples were run in triplicate. Results were analyzed using an iCycler (Biorad laboratories, Hemel Hempstead, UK).
To compare the relative quantities of different TLRs during the menstrual cycle, we divided biopsies into the following three groups: menstrual (LMP+1–4; n = 3; consisting of patients at LMP+1, +4 and +4), proliferative (LMP+5–14; n = 9; consisting of patients at early proliferative LMP+5, +5, +7, mid-proliferative LMP+8, +9, +10 and late proliferative LMP+11, +12 and +13) and secretory (LMP+15–29; n = 9; consisting of patients at early secretory LMP+16, +16 and +17, mid-secretory LMP+20, +21 and +22 and late secretory LMP+26, +28 and +29). Relative TLR expression quantities were compared between these groups. The threshold cycle values were normalized against the threshold value of human β-actin. Differences in normalized expression values between samples were tested for significance using ANOVA statistical test. The results were expressed as mean ± SEM. The level of statistical significance was set at P < 0.05.
Results
Immunostaining
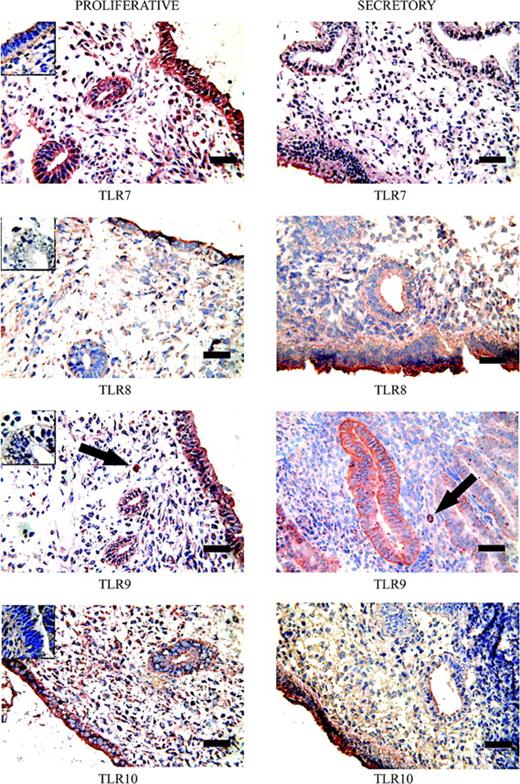
Cryosections were used to study the distribution of TLRs 7–10 in timed endometrial biopsies. Positive immunostaining for all these TLRs was observed in endometrial gland epithelium, luminal epithelium and stroma throughout the menstrual cycle (Figure 1). Moderate TLR7 staining was observed in gland and luminal proliferative epithelium with slight down-regulation of gland staining in the secretory phase. Weak TLR8 immunostaining was present in proliferative luminal and gland epithelium; up-regulation was observed in secretory luminal epithelium. Strong staining for TLR9 was observed in proliferative luminal epithelial cells with weaker staining in proliferative gland cells; gland staining increased in the secretory phase. Proliferative TLR10 immunostaining was similar to that of proliferative TLR9 staining. In contrast to TLR9, TLR10 epithelial gland cell staining decreased in the secretory phase although TLR10 luminal epithelium staining remained high. Diffuse stromal staining was observed for TLRs 7–10. In addition, specific intense staining of a sparse population of individual stromal cells was observed in both proliferative and secretory endometrial tissues stained for TLR9 (arrows, Figure 1).
Immunohistochemical staining of Toll-like receptor (TLR) 7–10 expression in human endometrium during secretory and proliferative phases of the menstrual cycle. Positive staining is red and negative staining blue. Insets show blocking of the anti-TLR7–10 antibodies with its specific peptides. Arrows depict cells in endometrial stroma with intensive staining with TLR9 antibody. Bar = 50 µm.
RT–PCR and quantitative PCR
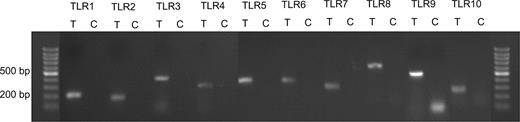
Figure 2 shows the results of RT–PCR for mRNA expression of TLR1–10 genes in human endometrial tissue. All amplified products were the predicted size for that particular gene. There was no product amplified in control samples indicative of the absence of genomic DNA contamination.
The expression of Toll-like receptor (TLR) 1–10 genes in human endometrial tissue. Each pair of primers produced a specific product with the specific predicted size in the test (T) samples. C = control samples.
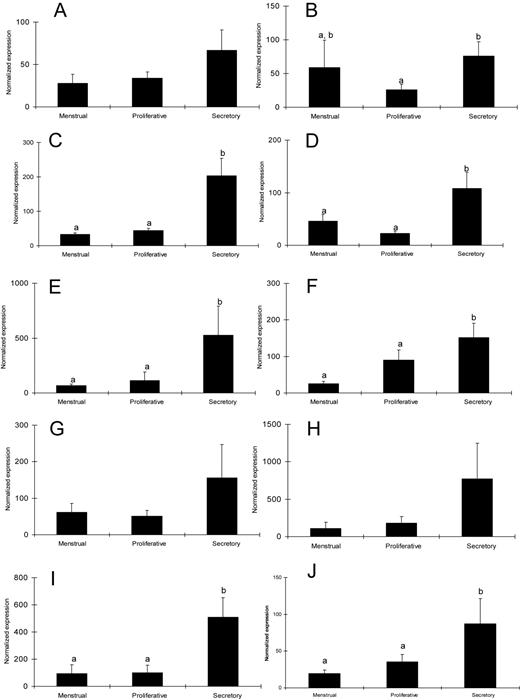
The quantitative expression profiles of TLR1–10 genes during the menstrual cycle in endometrial biopsies are shown in Figure 3 (A–J, respectively). TLRs 3–6, 9 and 10 showed a significantly higher expression of TLR in the secretory phase compared with the proliferative and menstrual phases of the cycle. Only, TLR2 expression was higher in the secretory phase compared with the proliferative phase but similar to menstrual phase. No significant difference was observed in the relative expression of genes for TLR1, 7 and 8 genes during the menstrual cycle.
Mean ± SEM of normalized expression values for Toll-like receptor (TLR) 1–10 genes in endometrial biopsies during different phases of menstrual cycle (A–J, respectively). The level of statistical significance was set at P < 0.05. Different letters denote significant differences.
Discussion
Uterine endometrial epithelial cells are the first layer of uterine defence against pathogens ascending the female reproductive tract. Thus, as detectors of non-self entities, TLRs may be expected to be present in this tissue. In the present investigation, we found the presence of TLR1–10 mRNA in human endometrial biopsies. Our results are in agreement with another report regarding the presence of TLRs 1–6 and 9 in endometrial samples (Young et al., 2004); however, in contrast to that report, we also detected the endometrial expression of TLRs 7, 8 and 10. None of the negative controls taken during our PCRs showed an amplified product (Figure 2), thus confirming the absence of genomic DNA contamination in our test PCR samples. The expression of all TLR molecules except TLR10 has been shown in endometrial cell lines (ECC-1) (Schaefer et al., 2004) and primary uterine epithelial cell cultures (Schaefer et al., 2005). The only significant difference between our investigations and that of Young et al. is that the TLR primers used for the amplification of TLR molecules in our study are different from those used previously. The immunohistochemical staining results of antibodies against TLRs 7–10 and their blocking with its specific peptides confirm the results of our gene expression studies regarding the presence of TLRs 7–10 in endometrial tissue.
Within the TLR family, TLRs 7–9 appear to be phylogenetically closely related to each other (Du et al., 2000) and form a functional subgroup that recognizes viral PAMPs in endosomal or lysosomal compartments (Heil et al., 2004). This is consistent with the fact that viral nucleic acid would be most likely detected by TLRs within an infected cell. For example, ssRNA viruses would reach the endosome through receptor-mediated uptake of a viral particle. There is evidence accumulating that, like TLR3, these TLRs can also respond to ‘self’ nucleic acid, which has been found to be immunostimulatory and may act as a ‘danger signal’ depending on its compartmentalization (Heil et al., 2004). Hence, it may be more correct to think of TLRs 7–9 as detectors of the abnormal localization of nucleic acid rather than as structures or motifs absent from the host (Diebold et al., 2004). It is reported that TLR9 recognizes unmethylated deoxycytidyl-phosphate-deoxyguanosine (CpG) dinucleotides that are common in bacterial and some viral nucleic acids (Hemmi et al., 2000; Bauer et al., 2001). Initially, TLRs 7 and 8 were shown to detect small antiviral compounds known as imidazoquinolines (Hemmi et al., 2002; Jurk et al., 2002). These were guanosine-based antiviral drugs. This indicated that the natural ligands for TLRs 7 and 8 could be viral nucleic acids. It has recently been reported that mouse TLR7 and human TLR8 (but not human TLR7) could recognize synthetic GU-rich ssRNA (Diebold et al., 2004; Lund et al., 2004).
Previously, we have reported the localization of TLRs 1–6 in various sections of the female reproductive tract using immunohistochemistry (Fazeli et al., 2005). Here, we provide further information regarding the localization of TLRs 7–10 in the endometrial tissue. For nearly all TLR molecules studied in the present investigation, the staining was not limited to epithelial cells and glands. It was also present in the stroma of the endometrium. No difference was found between proliferative- and secretory-stage endometrium staining with antibodies for different TLR molecules except slight increase in the stromal staining of TLR9. However, immunohistochemical staining is a qualitative technique and as such is not ideal for quantitative analysis. The specificity of staining for each TLR molecule was verified by blocking the staining using specific peptides for the respective antibody. Several other studies using immunohistochemistry have demonstrated the presence of TLR7 [tonsils (Mansson et al., 2006)] and TLR9 [liver (Martin-Armas, 2006), conjunctiva (Bonini et al., 2005) and gut (Rumio et al., 2004)] in different human tissues.
The endometrial environment is under the control of sex hormones during the menstrual cycle. The sex hormones not only regulate the anatomical and histological characteristics of endometrium (Beier and Beier-Hellwig, 1998; Classen-Linke et al., 1998) but are involved in the influx and localization of immune cells in the endometrium (Spornitz, 1992; Yeaman et al., 1997; von Rango, et al., 2001). For example, uterine natural killer (uNK) cells are found in the human uterus in large numbers spread throughout the endometrium with increasing numbers as the menstrual cycle progresses (Hunt, 1994; Givan et al., 1997). uNK cells mediate interferon-gamma production in the endometrium and are believed to be involved in the development of spiral arteries during early pregnancy and the control of trophoblast invasion. A recent article has shown that uNK cells express TLRs 2–4 (Eriksson et al., 2006) and that their response to TLR agonists is dependent on other cells within the endometrium. TLRs may therefore play a role in implantation other than the control of pathogens. Defensins, or cationic peptides, represent an important component of innate immune system at mucosal surfaces, including the female reproductive tract (Gallo et al., 2006; Lehrer and Ganz, 2002). Several of these broad-spectrum natural antimicrobial peptides are expressed in urogenital tissues, and their expression seems to be regulated by cycle-associated changes in sex hormones. For example, human intestinal defensin-5 (HD-5) mRNA is expressed in the vagina, ectocervix and variably in the endocervix, endometrium and Fallopian tube (Quayle et al., 1998). The endometrial expression of HD-5 mRNA has been reported to be higher during the early secretory phase of the cycle. The secreted HD-5 peptide in cervicovaginal lavage was also highest during the secretory phase of the menstrual cycle. The levels of other antimicrobials such as lactoferrin and lysozyme are also affected by oestradiol concentrations and the stage of the oestrous cycle (Cohen et al., 1984; Walmer et al., 1992). The adaptive immune system is also influenced by the altering levels of sex hormones during the menstrual cycle. Antigen presentation has been shown to be suppressed in response to increasing concentrations of oestrogen. Further to this, receptors for oestrogen have been found on both CD8 and CD4 T cells, so it is likely that their actions are also modified by changing concentrations of oestrogen. The actions of cytotoxic T cells appear to be down-regulated during days 14–28 of the menstrual cycle, when concentrations of progesterone and oestrogen are high (Beagley and Gockel, 2003). IgA and IgG have been shown to alter during the menstrual cycle, with the highest total levels of immunoglobulin occurring during menses, when oestrogen and progesterone are low and lowest at ovulation, when oestrogen is high and progesterone is low. However, responses vary in different parts of the reproductive tract with rising oestradiol increasing IgA in uterine secretions but suppressing levels in cervical mucus (Beagley and Gockel, 2003).
The present investigation clearly demonstrates alterations in relative expression of TLR2–6, 9 and 10 genes in the endometrial tissue during the menstrual cycle. Although these TLR molecules are expressed throughout the cycle, it seems the lowest amount of these genes is expressed during menstrual and proliferative stages of the cycle. The oestrogen levels are higher at the proliferative phase of the cycle compared with the secretory phase. At the same time, progesterone level is relatively higher at the secretory phase compared with the proliferative phase of the cycle. This may indicate an inhibitory effect of the oestrogen and/or a supporting influence of progesterone on the expression of TLR molecules in the endometrium. Further research should be focused towards understanding the regulation of expression of TLR molecules by sex hormones.
In agreement with our findings, Jorgenson et al. (2005) have recently demonstrated the cycle-dependent expression of TLR3 in primary endometrial epithelial tissue. Although Lesmeister et al. (2005) showed that in vitro treatment of endometrial epithelial cell lines with 17beta-oestradiol did not affect TLR3 mRNA or protein expression, treatment with 17beta-oestradiol did suppress cytokine and chemokine production resulting from TLR3 stimulation with poly I:C, suggesting that 17beta-oestradiol modulates TLR3 function.
In conclusion, we report the presence and localization of TLRs 7–10 for the first time in human endometrial biopsies. Our investigations indicate that the expression of TLRs 2–6, 9 and 10 is altered in the endometrium during the menstrual cycle. Further investigations should be directed towards understanding the underlying mechanism leading to changes in TLR expression in endometrium during the menstrual cycle as well as the significance of these cycle-dependent changes in mediating innate and adaptive immune responses in the female reproductive tract.
Acknowledgements
We thank Dr E. Sostaic for her critical discussion during the course of the experiments and preparation of the manuscript and Dr M. Aarabi for statistical support.