Abstract
We studied mitochondrial DNA variation in the European rabbit through the examination of restriction fragment length polymorphism in 526 individuals from 20 locations spread across the Iberian Peninsula. Digestion with eight enzymes of a 1120-bp fragment comprising most of the cytochrome b gene resolved 38 different haplotypes. These haplotypes were distributed in two highly divergent clades, with different but overlapping geographical distributions, and with comparable levels of within-clade variation. The overall phylogeographical pattern suggests a history of long-term regional isolation of two groups of rabbit populations, compatible with the recognition of two subspecies within the Iberian Peninsula, followed by recent contact and admixture. The underlying cause is sought in the alternation of glacial and interglacial periods in the late Pleistocene.
Similar content being viewed by others
Introduction
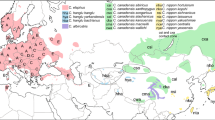
Both wild and domestic European rabbit belong to the single species Oryctolagus cuniculus, which is the sole extant representative of the genus Oryctolagus. Palaeontological data indicate limited morphological diversification of the genus since its appearance at the end of the Miocene (6.5 MYA) (Lopez-Martinez, 1989). Fossil remains from the middle Pliocene led to the recognition of two species, O. lacosti in southern France and north-western Italy and O. laynensis in the Iberian Peninsula. This latter form is thought to be the origin of the extant O. cuniculus (Lopez-Martinez, 1989). Archaeozoological data indicate that the original range of the rabbit was restricted to the Iberian Peninsula and the south of France up to river Loire (Callou, 1995). During the Middle Ages, humans started to have a major impact on rabbit distribution by transporting animals to many other geographical areas and rabbits can nowadays be found almost all over the world (Flux, 1994). Two subspecies are recognized, Oryctolagus cuniculus cuniculus which is present in France and in most areas where they were introduced by humans and O. c. algirus that occupies the Iberian Peninsula, North Africa, Mediterranean and Portuguese Atlantic islands (cf. Ellerman & Morrison-Scott, 1951; Saint-Girons, 1973; Wilson & Reeder, 1992).
The study of restriction fragment length polymorphism (RFLP) of the entire rabbit mtDNA molecule revealed the existence of two very distinct maternal lineages (Ennafaa et al., 1987; Biju-Duval et al., 1991). Lineage A was found in the south-west of the Iberian Peninsula and lineage B in the rest of Europe (including northern Spain) and also in domestic breeds. Further work revealed a total of 22 haplotypes in 12 wild populations from the Iberian Peninsula, southern France and the Mediterranean island of Zembra (Tunisia), and one French domestic stock (Monnerot et al., 1994). A clear subdivision into two groups was also found when nuclear genes were studied (van der Loo et al., 1991, 1999; Ferrand, 1995), with a geographical distribution that largely coincides with that of the two mitochondrial clades A and B. The geographical partitioning of these two groups can be superimposed, with few exceptions, on the distribution of the two subspecies based on classical taxonomy. Thus clade A was tentatively associated with O. c. algirus and clade B with O. c. cuniculus, as well as the corresponding nuclear gene pools (Biju-Duval et al., 1991; van der Loo et al., 1991; Ferrand, 1995; Branco & Ferrand, 1998).
Although the natural range distribution of the rabbit once comprised southern France up to river Loire (Callou, 1995), these populations are not relevant for the understanding of the early steps of rabbit history. As shown by ancient mtDNA, these populations were already less variable than their Iberian counterparts and human intervention has resulted in noticeable changes of their mtDNA genetic composition (Hardy et al., 1995; Loreille et al., 1997). Therefore, French populations (or any others to the north of the Pyrenees) tell us, mostly, the recent history of human-mediated colonization and expansion that occurred from the Middle Ages to the present. Thus, the major clues for the evolutionary history of the rabbit have to be looked for in its very original range, i.e. the Iberian Peninsula, by performing a detailed analysis of rabbit genetic structure and diversity in this area, not yet extensively sampled in previous studies.
In this work, a comprehensive survey of rabbit mtDNA diversity in the Iberian Peninsula by means of RFLP analysis will be presented. The strong geographical partitioning of mtDNA clades A and B is confirmed by our data, and the occurrence of both maternal lines in several populations is also described. This phylogeographical pattern suggests that two groups of populations were isolated for a certain time and later expanded and overlapped. The possibility that this separation resulted from retreat into glacial refugia during Quaternary ice-ages and that the present mixed populations resulted from secondary contact due to postglacial expansion and recolonization will be discussed. Finally, these two distinct groups will be related with the subspecies algirus and cuniculus and, subsequently, a re-evaluated geographical distribution in the Iberian Peninsula will be suggested.
Materials and methods
Sample collection
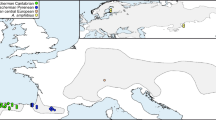
A total of 526 wild rabbits were sampled from 20 locations across the Iberian Peninsula (Fig. 1, sampling sites are named after the closest city or town except for Cabreira, Infantado, Doñana National Park and Las Lomas). Outside the hunting season rabbits were caught by ferreting and blood was collected from the marginal ear vein in EDTA-coated tubes. During the hunting season a portion of liver was taken from freshly killed animals. Samples were stored at −20°C until use.
DNA extraction
Total DNA was obtained from liver or blood by proteinase K digestion, phenol/chloroform extraction and ethanol precipitation using classical techniques. In samples from Las Lomas and Badajoz, mtDNA was obtained from liver purified on CsCl gradients (Ennafaa et al., 1987; Biju-Duval et al., 1991).
Amplification
A 1120-bp sized fragment of mtDNA, including most of the cytochrome b gene, was amplified with Taq Goldstar according to instructions of the manufacturer (Eurogentec). The reaction mixture consisted of 50 μL Eurogentec buffer, 150 μM dNTPs and 1 μM primers. Following 3 min of denaturation at 96°C, PCR was done in a Kontron cycler over 40 cycles of 1 min at 94°C, 1 min at 55°C and 1 min at 72°C. After the last cycle PCR was incubated a further 3 min at 72°C. The primers used were cytb8 (CCCCA TCCAA CATCT CTGCT TGATG AAA; 14242–14269) (Barome et al., 1998) and th3 (GTCTT GTAAG CCAGG AATGG AG; 15340–15361) (Hardy et al., 1994). The numbers indicate the nucleotide positions at the 5′ and 3′ ends, respectively, relative to the rabbit mitochondrial genome (accession number AJ001588, Gissi et al., 1998).
DNA sequencing
The sequence of the 1120-bp fragment for an individual carrying haplotype A2Rba was obtained by direct sequencing using Sequenase PCR Products sequencing kit (USB). This sequence was used as reference in restriction site mapping of lineage A (accession number AJ243096).
Restriction site analysis
Amplified mtDNA was digested with eight restriction enzymes (AluI, BsaI, DdeI, FokI, HaeIII, MseI, TaqI and Tsp509I) following instructions provided by the suppliers. The fragments obtained were separated by electrophoresis on 3% Nusieve agarose gels. The gels were immersed in an ethidium bromide solution 0.5% (w/v) for 10 min and RFLP patterns were visualized under UV light. Profiles are given in Table 1 with all generated fragments ordered by their relative position along the sequence.
Haplotype nomenclature
For simplicity, the 38 observed haplotypes will be referred to by the number given in the first column of Table 2. Sequences corresponding to haplotypes 1 and 19 (accession numbers AJ243197 and AJ243096) were used as references in the restriction site mapping of lineage B and lineage A, respectively. Distinct single endonuclease patterns were identified by a specific letter in order of appearance. Each individual was assigned a multiletter code describing its haplotype for the eight endonucleases (Table 2). Haplotypes described earlier (Ennafaa et al., 1987; Biju-Duval et al., 1991; Monnerot et al., 1994) were retyped and allocated to 12 haplotypes as shown in Table 2.
Data analysis
The MAPSORT subroutine of GCG (Devereux, 1989) was used to locate the site gain or loss for each new endonuclease profile. Haplotype and nucleotide diversity and nucleotide divergence were calculated using the REAP package (McElroy et al., 1991). Maximum parsimony was calculated using the heuristic search option of the PAUP software package (Swofford, 1998). Statistical parsimony was used to choose the best network among 100 networks of equal length. The ‘show patristic distances’ option of PAUP was used to identify the maximum parsimony network closest to the real data (see Templeton et al., 1987, 1992; Templeton & Sing, 1993, for details).
Results
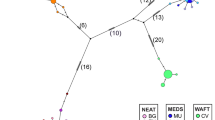
Five hundred and twenty-six wild rabbits from 20 Iberian locations were screened with eight restriction enzymes generating 52 restriction sites which discriminated 38 different haplotypes, of which 26 were new. The 38 haplotypes clustered into the two major lineages A and B, which are concordant with the two main groups previously described (Biju-Duval et al., 1991; Monnerot et al., 1994). Twenty-one haplotypes were of type A and 17 of type B. The phylogenetic relationships were represented by the maximum parsimony network depicted in Fig. 2(a) and this does not differ importantly from that obtained using the distances approach (genetic distances were computed according to Nei & Tajima, 1981; results not shown). Clades A and B were separated by at least 14 mutations (nucleotide divergence 11.9%) and they exhibited similar levels of within-clade polymorphism with nucleotide diversities of 1.1% and 1.4%, respectively. The number of different haplotypes per population averaged 4.6 (range 1–7). Across individual rabbits, 329 carried type A mtDNA and 197 carried type B.
(a) Maximum parsimony network representing the phylogenetic relationships between the 38 mtDNA types. Circle size is proportional to the number of individuals in the total sample. Black points represent potential intermediates. (b) Parsimony networks for populations. Detected haplotypes are represented by grey circles with size proportional to the number of individuals (refer to the previous figure for haplotype number). Unobserved haplotypes and potential intermediates are represented by black points.
Geographical distribution of clades
Globally, the two mtDNA clades A and B show a highly structured geographical pattern of distribution (Fig. 1). Clade A was fixed or almost fixed in all south-western locations whereas clade B was almost the only one present in the north and east. Both clades were approximately equally common in three more or less centrally located populations (Toledo, Bragança and Vila Viçosa).
The high estimated average of nucleotide divergence (4.4%) obtained among populations reflects the level of genetic structuring and, mainly, the coexistence in several populations of haplotypes from clades A and B. Maximum nucleotide diversity ranged from 1.2% in Las Lomas and 1.1% in Peralta for clade A and clade B, respectively, to 7.5% for the mixed population of Bragança (Table 2).
Patterns of haplotype distribution and population diversity
Six out of the 38 haplotypes have a wide geographical distribution. On the other hand, 24 are present only in a single location, of which 10 are in a single individual. These private haplotypes were distributed among 14 of the 20 locations surveyed. The distribution of the 38 haplotypes over the 20 locations is summarized in Fig. 2(b). The observed mtDNA diversity is superimposed over the maximum parsimony network of Fig. 2(a) representing only the haplotypes detected in each location. The observation of Fig. 2(b) reveals a clear trend in the interior vs. exterior position of haplotypes in the network. For clade B, the trend is from coastal Mediterranean locations (Alicante, Tarragona, Lérida) with interior positioned haplotypes, inland to the Ebro valley (Caparroso, Tudela, Peralta), and further westwards to north-eastern Portugal (Bragança) with exterior haplotypes. A similar pattern though less conspicuous, appears to be present for clade A from Las Lomas with interior haplotypes, northwards in the direction of Badajoz and Ciudad Real with haplotypes increasingly positioned towards the exterior. Southern locations proved to be the most variable and are characterized by the widespread occurrence of haplotypes 18, 19 and 25. Lower levels of variation were found along the western coast in a northward direction with one location monomorphic for haplotype 29 which is restricted to this region (Table 2). Inspection of Fig. 2(b) reveals that this particular haplotype characterizes all western locations, and that two of their few private haplotypes are derived from it. The highest level of variability within clade B was observed in the south-eastern coast and in northern Spain. The location of Alicante is characterized by a balanced distribution of five different haplotypes, whereas haplotype 4 dominates the Navarra locations (Caparroso, Peralta and Tudela). The lowest levels of diversity were found in the north-eastern coast, in Lérida and Tarragona, where haplotypes 2 and 14, respectively, predominate. Clades A and B were found to coexist in nine locations, but only in three of them were they equally abundant (Bragança, Toledo and Vila Viçosa). The six remaining locations have either clade A or clade B predominating (Table 2).
Discussion
Evidence for only two distinct mtDNA lineages
The fact that all haplotypes found in the original range of the rabbit species cluster within only two clades is a strong evidence that there are no more than two divergent lineages within O. cuniculus. Thus, our data confirm and further extend previous work in which a strong geographical subdivision of the two clades as well as the occurrence of admixed populations were described (Biju-Duval et al., 1991; Monnerot et al., 1994). The detection of both lineages A and B in Badajoz led Monnerot et al. (1994) to suggest the possibility of a contact zone, without, however, excluding the possibility that mixed populations resulted from introductions.
The detailed analysis of the distribution of the two distinct mtDNA clades throughout the Iberian Peninsula revealed a clear geographical pattern whereby south-western populations are fixed or nearly fixed for one lineage and north-eastern populations for the other. Rabbits of both mtDNA clades coexist in several locations in a geographically intermediate region (Fig. 1). The deep genetic differentiation observed, together with the contiguous and marginally overlapping ranges can be explained in a straightforward way by the allopatric origin of two monophyletic groups (Avise, 1989). Indeed, the observed haplotype distribution is in line with the persistence of rabbit populations in two main refugial areas during Pleistocene glaciations, followed by postglacial expansion after the last glacial maximum approximately 20 000 years ago. An alternative hypothesis of single origin would involve a highly biased process of lineage sorting and extinction, which in the present case seems to be much less probable.
Several phylogeographical studies provide evidence for the existence of glacial refugia in the Iberian Peninsula during the Quaternary (for example Cooper et al., 1995; Hewitt, 1996; Dumolin-Lapègue et al., 1997; Comes & Abbot, 1998; Taberlet et al., 1998; Sinclair et al., 1999). Yet, a remarkable feature of the two mtDNA clades observed in rabbits is their deep divergence, suggesting a long-time separation. Using an average rate of mammalian mtDNA divergence, Biju-Duval et al. (1991) estimated a divergence time of 2 MY for clades A and B. This places their origin on a Pliocene/Pleistocene boundary dated by several authors at 1.6–2.0 MYA (Williams et al., 1993), predating the more recent glaciation events.
Our RFLP analysis of cytochrome b yielded an estimated nucleotide divergence of 11.9% between clades A and B, which is in contrast with a previously reported value (4.2%) obtained from the entire mtDNA molecule (Biju-Duval et al., 1991). This may be due to the use of different nucleotide sampling regimes, and accepting the published calibration (Biju-Duval et al., 1991) we calculate a divergence rate of 6% per MY from our data. Within clades, nucleotide diversities (1.1% within clade A and 1.4% within clade B) would then correspond to a most recent common mitochondrial ancestor (MRCMA) at about 183 000 and 233 000 years, for clade A and clade B, respectively. Despite the technical differences, a very similar estimate for MRCMA was made by Biju-Duval et al. (1991) based on the analysis of the southern Spanish population of Las Lomas.
Haplotype phylogeny and population history
The extensive nucleotide diversity observed in both clades in which a one-step evolutionary branching pattern is predominant suggests that a common historical factor – perhaps related with a climatic amelioration and a significant improvement in habitat availability — resulted in increased population sizes and subsequent accumulation of genetic diversity. This indicates that severe population size reductions have been absent, in spite of drastic short-term, high-amplitude climatic oscillations that appear to have occurred during late Pleistocene (see Roy et al., 1996).
A closer look at the haplotype network (Fig. 2a) reveals some differences in the branching patterns of clades A and B; while essentially two groups of haplotypes exist in clade A (common and rare), in clade B haplotypes are more evenly distributed (Table 2) and, clade A displays a branching pattern in which haplotypes are separated by mostly one mutational steps. The more interior haplotypes (18, 19 and 29) are also the ones that are most abundant and have the largest geographical ranges. Other haplotypes seem to have derived from these, occur in low frequencies and have restricted geographical ranges. On the other hand, clade B shows a more linear and scattered branching pattern with several missing nodes. The most central haplotype (14) is not the most abundant and has a scattered geographical distribution. Among the more frequent haplotypes such as haplotypes 2 and 4, large as well as restricted geographical ranges can be found. According to Neigel & Avise (1993) lineage age and lineage range are correlated with older lineages having broader ranges. Therefore, our observations suggest a history of large and stable populations associated with clade A, allowing the diversification and maintenance of many haplotype variants. A history of small or fluctuating and possibly fragmented populations is more in line with the phylogeographical pattern observed for clade B.
Historical biogeography
The first evidence for rabbit mtDNA geographical subdivision triggered the suggestion of a glacial refugium in the Mediterranean coast of France (Biju-Duval et al., 1991). More recently, the first northern Iberian populations were surveyed revealing an extensive variation inside clade B thus prompting Monnerot et al. (1994) to suggest that an isolate somewhere in the northern Iberian Peninsula was more plausible than one in southern France. The study of ancient mtDNA revealed that until the Middle Ages, when the natural range extended up to river Loire (Callou, 1995), French populations were characterized by haplotype 2 (clade B). During the Middle Ages rabbits quickly expanded all over France and with them mitochondrial type 1 (also clade B) (Hardy et al., 1995). Present day distribution of mtDNA types confirms the generalized occurrence of haplotypes 1 and 2, being haplotype 1 the most abundant (Mougel, 1997; Monnerot, 1998). Additionally, French populations are characterized by low levels of genetic variability at protein and immunoglobulin loci (van der Loo et al., 1991; Ferrand, 1995; Branco et al., 1999; van der Loo et al., 1999).
The data on genetic differentiation and diversity now available leads us to postulate that the glacial refugia for the rabbit were on the Iberian Peninsula one in the south-west and another one in the Mediterranean coast that probably extended to the north-east. Quaternary glaciations in the Iberian Peninsula appear to have been restricted to the Cordillera Cantábrica and Cordillera Central, leaving the southern region and the Mediterranean coastal area free of ice. A refugial area on the eastern coast of the Iberian Peninsula has already been suggested in phylogeographical studies of the European grasshoper (Chorthippus parallelus) (Cooper et al., 1995) and the Mediterranean ragwort (Senecio gallicus) (Comes & Abbot, 1998). The valley of the river Ebro may have formed a northward extension of the Mediterranean coast refuge. However, due to the proximity of the main glaciation areas of Cordillera Cantábrica, Cordillera Central and Pyrenees, the populations in the north-east refugium may have been subjected to more extreme climatic conditions than those in the south-west, possibly explaining the observed trend for high vs. low levels of genetic structuring that is observed in Fig. 2(b).
The observed phylogeographical pattern is that of two groups of populations with contiguous geographical ranges which overlap to a certain extent. The area of range overlap is a relatively narrow region crossing the Iberian Peninsula in a south-east-north-west direction. We observed a trend for haplotypes interior to the clades to be found at or near the proposed glacial refugia, while haplotypes with exterior positions were more frequently found in the region of overlap. Because a relative temporal scale can be inferred from the hierarchy of haplotype position in a phylogenetic tree whereby most interior haplotypes are the oldest and those with outer positions are younger (Templeton et al., 1995), the described pattern suggests that the south-east-north-west area across the Iberian Peninsula was colonized more recently giving rise to a secondary contact zone between rabbits carrying clades A and B (Fig. 1). This hypothesis is supported by the analysis of DNA extracted from rabbit remains dated from 11 500 to 4500 B.P., as only clade B haplotypes were detected in Central Iberian Peninsula and in the southern foothills of Sierra Nevada (Loreille et al., 1997; Monnerot, unpublished results). Thus, the contact in these areas probably does not exceed 4500 years.
Some clade B haplotypes were observed in the south-west of the Iberian Peninsula but only the most common and widespread B haplotype (2) was detected. This observation may indicate a rapid penetration of rabbits approximately along the route Badajoz – Vila Viçosa – Infantado (Fig. 1). Conversely, a few A haplotypes were found in the north. Their local presence may have natural causes, but introductions by humans are highly probable in this region. Despite a low success rate (Branco et al., 1997), rabbit translocations for hunting purposes are known to have occurred in this region, mostly with animals from the southern regions where they are more abundant.
Intraspecific taxonomy
The data on cytoplasmic and nuclear genes (van der Loo et al., 1991; Ferrand, 1995; Branco & Ferrand, 1998; van der Loo et al., 1999) reveal the presence of two distinct and highly diversified evolutionary lineages of the European rabbit and support the recognition of two subspecies in the modern context of an appreciable level of genetic differentiation and historical continuity of barriers to genetic exchange (see Smith et al., 1997; Templeton, 1999). The A and B clades can then be correlated with, respectively, O. c. algirus and O. c. cuniculus. However, the match with traditional taxonomy is not perfect. For example only O. c. algirus was considered present on the Iberian Peninsula as morphological criteria were utilized for the description of rabbit subspecies (Cabrera, 1914). Thus, any future attempts to re-evaluate the taxonomy of O. cuniculus should take the genetic data into account.
Conclusions
We observed (i) a strong discontinuity in the geographical distribution of two distinct clades A and B, that follows a north-west to south-east axis across the Iberian Peninsula (ii) a relatively narrow region along this axis where both clades are present in similar frequencies, and (iii) high levels of within-clade genetic diversity and structuring. We interpret the observed phylogeographical pattern as resulting from a secondary contact of two groups of rabbits after a long-term separation during which a considerable level of diversity accumulated between and within groups. This scenario is compatible with the occurrence of range contractions to two glacial refugia followed by range expansions in response to Quaternary climatic oscillations, resulting in the present day limited range overlap and admixture.
Accession codes
References
Avise, J. C. (1989). Gene trees and organismal histories: a phylogenetic approach to population biology. Evolution, 43: 1192–1208.
Barome, P. -O., Monnerot, M. and Mounolou, J. -C. (1998). Intrageneric phylogeny of Acomys (Rodentia, Muridae) using mitochondrial gene cytochrome b. Mol Phyl Evol, 9: 560–566.
Biju-Duval, C., Ennafaa, H., Dennebouy, N., Monnerot, M., Mignotte, F., Soriguer, R. C. et al (1991). Mitochondrial DNA evolution in Lagomorphs: origin of systematic heteroplasmy and organization of diversity in European rabbits. J Mol Evol, 33: 92–102.
Branco, M. and Ferrand, N. (1998). Genetic polymorphism of rabbit (Oryctolagus cuniculus) tissue acid phosphatases (ACP2 and ACP3). Comp Biochem Physiol, 120B: 405–409.
Branco, M., Ferrand, N., Rocha, J. and Moreno, S. (1997). Gene admixture in wild rabbit populations (Oryctolagus cuniculus): preliminary results of a restocking operation in Doñana National Park (Spain). Gibier Faune Sauvage, 14: 498–499.
Branco, M., Machado, J. C. and Ferrand, N. (1999). Extensive genetic polymorphism of peptidases A, B, C and D, in wild rabbit (Oryctolagus cuniculus) populations from the Iberian Peninsula. Biochem Genet, 37: 237–249.
Cabrera, J. (1914). Fauna Ibérica. Museo Nacional de Ciências Naturales de Madrid, Madrid.
Callou, C. (1995). Modifications de l’aire de répartition du Lapin (Oryctolagus cuniculus) en France et en Espagne, du Pléistocène à l’époque actuelle. État de la question. Anthropozoologica, 21: 95–114.
Comes, H. P. and Abbott, R. J. (1998). The relative importance of historical events and gene flow on the population structure of a Mediterranean ragwort, Senecio gallicus (Asteraceae). Evolution, 52: 355–367.
Cooper, S. J. B., Ibrahim, K. M. and Hewitt, G. M. (1995). Postglacial expansion and genome subdivision in the European grasshopper Chorthippus parallelus. Mol Ecol, 4: 49–60.
Devereux, J. (1989). GCG: Genetic Computer Group sequence analysis software package. Nucl Acids Res, 12: 387–395.
Dumolin-Lapègue, S., Demesure, B., Fineschi, S., Corre, V. L. and Petit, R. J. (1997). Phylogeographic structure of white oaks throughout the European Continent. Genetics, 146: 1475–1487.
Ellerman, J. R. and Morrison-Scott, T. C. S. (1951). Checklist of the Palaearctic and Indian Mammals 1758–1946. Trustees of the British Museum, London.
Ennafaa, H., Monnerot, M., Gaaied, A. and Mounolou, J. C. (1987). Rabbit mitochondrial DNA: preliminary comparison between some domestic and wild animals. Génét Sél Évol, 19: 279–288.
Ferrand, N. (1995). Variação genética de proteínas em populações de coelho (Oryctolagus cuniculus). Análise da diferenciação subespecífica, subestruturação, expansão geográfica e domesticação. PhD Thesis, Faculdade de Ciências, Universidade do Porto, Porto.
Flux, J. E. C. (1994). World distribution. In: Thompson, H. V. and King, C. M. (eds) The European Rabbit, pp. 8–21. Oxford Science Publications, Oxford.
Gissi, C., Gullberg, A. and Arnason, U. (1998). The complete mitochondrial DNA sequence of the rabbit, Oryctolagus cuniculus. Genomics, 50: 161–169.
Hardy, C., Casane, D., Vigne, J. D., Callou, C., Dennebouy, N., Mounolou, J. C. et al (1994). Ancient DNA from Bronze Age bones of European rabbit (Oryctolagus cuniculus). Experientia, 50: 564–570.
Hardy, C., Callou, C., Vigne, J. D., Casane, D., Dennebouy, N., Mounolou, J. C. et al (1995). Rabbit mitochondrial DNA diversity from prehistoric to modern times. J Mol Evol, 40: 227–237.
Hewitt, G. M. (1996). Some genetic consequences of ice ages, and their role in divergence and speciation. Biol J Linn Soc, 58: 247–276.
van Der Loo, W., Ferrand, N. and Soriguer, R. M. (1991). Estimation of gene diversity at the b locus of the constant region of the immunoglobulin light chain in natural populations of European rabbit (Oryctolagus cuniculus) in Portugal, Andalusia and on the Azorean islands. Genetics, 127: 789–799.
van Der Loo, W., Mougel, F., Sánchez, M. S., Bouton, C., Castien, E., Fonseca, A. et al (1999). Cytonuclear disequilibria in wild populations of rabbit (Oryctolagus cuniculus L.) suggest unequal allele turnover rates at the b locus (IGKC1). Immunogenetics, 49: 629–643, 10.1007/s002510050659.
Lopez-Martinez, N. (1989) Revision sistematica y biostratigrafica de los lagomorphos (Mammalia) del neogeno y cuaternario de España. Memorias del Museo Paleontologico de la Universidad de Zaragoza. Disputacion General de Aragon.
Loreille, O., Mounolou, J. -C. and Monnerot, M. (1997). Historical expansion of rabbits and ancient DNA. C R Soc Biol, 191: 537–544.
Mc Elroy, D., Moran, P., Bermingham, E. and Kornfield, J. (1991) : The restriction enzyme analysis package, version 4.0. University of Maine, Orono, ME.
Monnerot, M. (1998) Diversity of Oryctolagus cuniculus populations. Euro-American Mammal Congress, Santiago de Compostela.
Monnerot, M., Vigne, J. D., Biju-Duval, C., Casane, D., Callou, C., Hardy, C. et al (1994). Rabbit and Man. Genetic and historic approach. Génét Sél Évol, 26(Suppl. 1),167–182.
Mougel, F. (1997) Variation de trois types de marqueurs génétiques dans l′ évolution de l’espèce Oryctolagus cuniculus: aspects moléculaire et relations avec la biologie et la structure des populations. PhD Thesis, Université de Paris Sud, Orsay.
Nei, M. and Tajima, F. (1981). DNA polymorphism detectable by restriction endonucleases. Genetics, 97: 145–163.
Neigel, J. E. and Avise, J. C. (1993). Application of a random walk model to geographic distributions of animal mitochondrial DNA variation. Genetics, 135: 1209–1220.
Roy, K., Valentine, J. W., Jablonski, D. and Kidwell, S. M. (1996). Scales of climatic variability and time averaging in Pleistocene biotas: implications for ecology and evolution. Trends Ecol Evol, 11: 458–462.
Saint-Girons, M. C. (1973). Les mammifères de France et du Benelux. Doin Editions, Paris.
Sinclair, W. T., Morman, J. D. and Ennos, R. (1999). The postglacial history of Scots pine (Pinus sylvestris L.) in western Europe: evidence from mitochondrial DNA variation. Mol Ecol, 8: 83–88.
Smith, H. M., Chiszar, D. and Montanucci, R. R. (1997). Subspecies and classification. Herpetol Rev, 28: 13–16.
Swofford, D. L. (1998). PAUP*: Phylogenetic analysis using parsimony (*and other methods), version 4 beta. Sinauer Associates, Sunderland, MA.
Taberlet, P., Fumagalli, L., Wust-Saucy, A. G. and Cosson, J. F. (1998). Comparative phylogeography and postglacial colonization routes in Europe. Mol Ecol, 7: 453–464.
Templeton, A. R. (1999). Human Races: a genetic and evolutionary perspective. Am Anthropologist, 100: 632–650.
Templeton, A. R. and Sing, C. F. (1993). A cladistic analysis of phenotypic associations with haplotypes inferred from restriction endonuclease mapping. IV. Nested analyses with cladogram uncertainty and recombination. Genetics, 134: 659–669.
Templeton, A. R., Boerwinkle, E. and Sing, C. F. (1987). A cladistic analysis of phenotypic associations with haplotypes inferred from restriction endonuclease mapping. I. Basic theory and an analysis of alcohol dehydrogenase activity in Drosophila. Genetics, 117: 343–351.
Templeton, A. R., Crandall, K. A. and Sing, C. F. (1992). A cladistic analysis of phenotypic associations with haplotypes inferred from restriction endonuclease mapping and DNA sequence data. III. Cladogram estimation. Genetics, 132: 619–633.
Templeton, A. R., Routman, E. and Phillips, C. A. (1995). Separating population structure from population history: a cladistic analysis of the geographical distribution of mitochondrial DNA haplotypes in the tiger salamander, Ambystoma tigrinum. Genetics, 140: 767–782.
Williams, M. A. J., Dunkerley, D. L., de Deckker, P., Kershaw, A. P. and Stokes, T. (1993). Quaternary Environments. Edward Arnold, New York.
Wilson, D. E. and Reeder, D. (1992). Mammal Species of the World. A Taxonomic and Geographic Reference. Smithsonian Institute Press, Washington.
Acknowledgements
Financial support was obtained from the French Ministery of Research and Technology in the program of Systematics and Biodiversity (ACCSV7, granted to MM), from JNICT (project STRDA/C/CEN/471/92), from FCT (PRAXIS XXI BD/5007/95 PhD grant to MB, and FMRH/BSAB/50/98 sabbatical grant to NF) and from Direcção Geral das Florestas. We thank P.C. Alves, J.L. Rosa and A. Loureiro for their help in the Portuguese sampling campaigns and also S. Moreno, R. Villafuerte, E. Castien, I. Rodriguez, J. Arques, V. Peiró, J. Ruiz-Olmo, R. Soriguer and M. Sanchez for their help in some Spanish sampling campaigns. We thank Pim Arntzen, Tristão Branco, Paul Sunnucks and Alan Templeton for comments and suggestions.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Branco, M., Ferrand, N. & Monnerot, M. Phylogeography of the European rabbit (Oryctolagus cuniculus) in the Iberian Peninsula inferred from RFLP analysis of the cytochrome b gene. Heredity 85, 307–317 (2000). https://doi.org/10.1046/j.1365-2540.2000.00756.x
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1046/j.1365-2540.2000.00756.x
Keywords
This article is cited by
-
An overview of the lagomorph immune system and its genetic diversity
Immunogenetics (2016)
-
Proteomic study of the brackish water mussel Mytilopsis leucophaeata
Zoological Studies (2015)
-
Genetic diversity comparison of the DQA gene in European rabbit (Oryctolagus cuniculus) populations
Immunogenetics (2015)
-
Low persistence in nature of captive reared rabbits after restocking operations
European Journal of Wildlife Research (2015)
-
The use of faecal markers for the delimitation of the European rabbit’s social territories (Oryctolagus cuniculus L.)
acta ethologica (2013)