Abstract
The biological toxicity of high levels of breathing gases has been known for centuries, but the mechanism remains elusive. Earlier work mainly focused on the influences of dispersed gas molecules dissolved in water on biomolecules. However, recent studies confirmed the existence of aggregated gas molecules at the water-solid interface. In this paper, we have investigated the binding preference of aggregated gas molecules on proteins with molecular dynamics simulations, using nitrogen (N2) gas and the Src-homology 3 (SH3) domain as the model system. Aggregated N2 molecules were strongly bound by the active sites of the SH3 domain, which could impair the activity of the protein. In contrast, dispersed N2 molecules did not specifically interact with the SH3 domain. These observations extend our understanding of the possible toxicity of aggregates of gas molecules in the function of proteins.
Similar content being viewed by others
Introduction
An increased concentration of breathing gases (N2, O2, Ar, Kr, CO2, Xe, etc.) in body tissues can cause severe biological effects, such as nitrogen narcosis1,2,3,4. Although some mechanisms have been proposed, such as the gas-induced alteration of ion permeability at the cell membrane or the binding of gas to proteins of neurotransmitter receptors5,6,7, the mode of action remains unresolved. Earlier work mainly focused on the influences of dispersed gas molecules dissolved in water on biomolecules. The recent experimental and theoretical studies demonstrated that gas can be aggregated at the water-solid interface in the form of nanobubbles8,9,10,11,12,13,14,15,16,17,18 and nanopancakes19,20 after the great debate21,22,23. Thus, the interaction between aggregated gas molecules, as opposed to dispersed gas molecules and biological macromolecules is an important area of investigation.
The interactions between nanoscale particles and biomolecules have attracted great attention. Previous studies have shown that nanoparticles can adsorb onto proteins24,25,26,27,28,29,30,31,32,33,34,35,36,37,38,39,40,41. If the protein-ligand binding sites are attacked or blocked by the nanoparticles, the biological functions of these proteins could be impaired. Considering that the binding sites and many of the aggregated gases are usually hydrophobic42,43,44,45, we questioned if aggregated gas molecules showed similar toxicity toward proteins as observed with nanoparticles.
In this paper, we applied molecular dynamics simulations to study the adsorption capability of aggregated gas onto the Src-homology 3 (SH3) domain. The SH3 domain was chosen due to its widespread presence in protein families for regulating signaling pathways and for substrate specific-binding46,47,48,49,50,51,52 (Figure 1). Figure 1 was generated using the VASCo PYMOL plug-in53. The functional loss of the SH3 domain would be lethal for the normal functions of living cells. As the gas, N2 was employed due to its abundance in air. We found that aggregated N2 molecules preferred to adhere to the SH3 domain at the ligand-binding sites of the SH3 domain. In contrast, the dispersed N2 molecules showed no specific binding to the SH3 domain.
A typical structure of the SH3 domain bound to its ligand.
The yellow loop represents the corresponding ligand of the SH3 domain. The four key binding sites (8(PHE), 36(TRP), 50(PRO), 53(TYR)) of the SH3 domain are plotted as red sticks. The surface of the SH3 domain is shown in PYMOL-CGOs and colored according to lipophilic potential. The hydrophobic surface is indicated by red and the hydrophilic surface by blue.
Results
In the initial configuration of the system, N2 molecules were placed around the SH3 domain in the water box with box size of Lx = 7.5 nm, Ly = 7.5 nm, Lz = 7.5 nm (Figure 2). Different numbers of N2 molecules were examined in six samples with 100 ns molecular dynamics simulations under the NPT (P = 1 atm, T = 300 K) ensemble.
The initial configuration of the simulation.
N2 molecules were placed around SH3 in the water box with box size of Lx = 7.5 nm, Ly = 7.5 nm, Lz = 7.5 nm. The key residues of the protein domain are noted by the red surfaces. The rest of residues of protein are shown as the cyan semitransparent surface. N2 molecules are shown as blue balls.
In the first four samples, 330, 310, 290 and 270 nitrogen molecules were placed in the cubic water box, denoted by NS_1, NS_2, NS_3 and NS_4, respectively. The N2 molecules were found to be aggregated and we denoted them as NS. In the other two samples, denoted by DS_1 and DS_2, with 216 and 205 N2 molecules, respectively, the maximum aggregate size was no more than 6 N2 molecules and thus we denoted them as DS. The details of all samples appear in Table 1. The criterion to determine the existence of aggregated N2 molecules followed the Wolde and Frenkel (TWF) theory54. Two N2 molecules were considered as a neighboring pair if their distance was less than rS (the Stillinger radius, in our systems, the rS = 1.5σNN = 0.489 nm)54,55. If a N2 molecule had more than two neighbors, this N2 belonged to the aggregated N2 molecules. More computational details are listed in the Methods section.
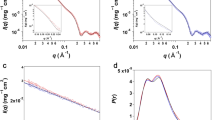
(a) A typical adsorption probability plot of each residue.The X-axis represents the index of residues in the SH3 domain while the Y-axis represents the probability that the minimum distance of the amino acid to any one nitrogen molecule will be less than 0.489 nm in the last 50 ns. The blue rectangles stand for the probabilities of the aggregated N2 system (NS_1) and the red rectangles stand for the probabilities of the dispersed N2 system (DS_1). (b) The exposed area in water of each residue of the SH3 domain.
Representative snapshots of the trajectory for NS_1 and the change of the interface area of this sample are given in Figure 3. (For other samples, see Figure S3 in supplementary information). The interface area was defined as:

(a) Some representative snapshots of one trajectory (NS_1).The key residues of the protein domain are noted by red surfaces. The rest of residues of protein are shown in cyan semi-transparent surface. The N2 are shown as blue balls. (b) The change of interface area of N2 molecules and the SH3 domain corresponding to the trajectory shown in Figure 3a.
The SASASH3, SASANIT and SASASH3+NIT were defined as the solvent accessible surface areas of the SH3 domain, the N2 molecules and the complex of the SH3 domain and the N2 molecules, respectively56. Based on the snapshots and interface area, we could illustrate the whole simulation process. For NS situations, in the first ~5 ns, N2 molecules gathered quickly, corresponding to a decline in the interface area to nearly zero. Then, the aggregated N2 molecules were separated from the SH3 domain for about 25 ns. Next, the interface area rose quickly to about 3 nm2, which was caused by a rapid adsorption of aggregated N2 onto the SH3 domain. This increase in the interface area indicated a strong interaction between aggregated N2 molecules and the hydrophobic surface of the SH3 domain. Then, the aggregated N2 molecules attached the SH3 domain in the rest of the simulation (see the snapshots at t = 35 ns and t = 100 ns). There were some large fluctuations in the curve of the interface area, owing to the quite loose and flexible structure of the aggregated N2 molecules.
Discussion
To further characterize the adsorption between the N2 and the SH3 domain, we calculated the probability of the adsorption of N2 molecules to each amino acid residue in the SH3 domain over the last 50 ns. The distances between the amino acids and N2 molecules were evaluated in every frame (In simulations, we recorded one frame of trajectory every 1 ps). If the minimum distance of one amino acid to any N2 molecule was less than rS (rS = 1.5σNN = 0.489 nm) in one frame, we defined the phenomenon as the adsorption of N2 molecules by this particular amino acid in this specific time frame and the adsorption frequency of this amino acid added 1. The adsorption probability of a specific amino acid was the adsorption frequency of this amino acid divided by the total frame number (50,000 in our simulations).
The typical adsorption probabilities of each residue are illustrated by a histogram (Figure 4a) showing data for NS and DS (Figures of more samples can be found in Figure S2 in supplementary information). A high probability means strong adsorption. Additional details appear in Table 1, which contains the favorable N2-binding residues, defined as those residues with a probability of greater than 0.7 of all samples. In the first four samples (NS_1–NS_4 in Table 1), the SH3 domain had the same favorable binding sites with the aggregated N2 molecules. The amino acid residues 7(LEU), 8(PHE), 35(GLN), 36(TRP), 50(PRO), 52(PRO) and 53(TYR) presented more favorable N2 binding in NS cases. It is known that 8(PHE), 36(TRP), 50(PRO) and 53(TYR) (Figure 1 and Table 1) are the key binding sites of the SH3 domain and its corresponding ligand51. Thus, we can conclude that the binding of the aggregated N2 molecules to the SH3 domain had specificity for the active sites. Figure 1 shows a typical structure of the ligand binding to the SH3 domain. The hydrophobicity of the surface was plotted by lipophilic potential and the surface of the SH3 domain in contact with the ligand was hydrophobic57. Since the aggregated N2 molecules completely covered the ligand-binding sites of the SH3 domain, aggregates of nitrogen might block the key binding sites and hinder the normal process of ligand binding at the SH3 domain. In another words, binding of aggregated N2 molecules might lead to the loss of protein function. Because proteins were the functional units essential for life, our simulation indicated that aggregated gas might be toxic.
In the last two samples (DS_1 and DS_2) in Table 1, N2 molecules did not aggregate, a completely different outcome that with the NS samples. Another difference was an absence of probabilities greater than 0.7 for the most active binding sites (8, 36, 50 and 53) for DS, which means the adsorption abilities of DS were much weaker than the adsorption abilities of NS at these sites. More importantly, a greater number of N2 molecules did not correspond to the high adsorption probabilities when N2 molecules were dispersed, which could be illustrated by DS_1 and DS_2 with 216 N2 and 205 N2, respectively (shown in Figure S2 in supplementary information). The differences in the probabilities of binding could only be from the existence of the aggregated N2, as shown in Table 1. To further illustrate the difference between DS and NS, we calculated the standard deviation of each residue's adsorption probability to N2 (Table 1). All standard deviations of NS were greater than 0.25, whereas none of those of DS was greater than 0.19. Furthermore, the distribution of probability of DS tended to be well-distributed compared with NS, which indicated the adsorption of dispersed N2 molecules to the SH3 domain had no specific binding sites. This fact implied that the characteristic binding sites were related to the state of N2. Only when the N2 molecules were aggregated could they adsorb well onto the SH3 domain at some specific sites.
Finally, it should be noted that the SH3 domain is a small domain with 57 residues. Each residue in SH3 could be in contact with water and N2 molecules. The calculated exposed area of each residue in water is shown in Figure 4b. Compared with Figure 4a, the residues with the largest exposed areas (15(GLU), 43(GLU) and 57(TYR)) did not correspond to large adsorption probabilities. On the other hand, the exposed area of the key residue 50(PRO) was very small even though its adsorption probability was very large. Considering that the four key residues (8(PHE), 36(TRP), 50(PRO) and 53(TYR)) are all located at the hydrophobic groove (red surface in Figure 1) of the SH3 domain, the ability of each residue to adsorb nitrogen was related to the hydrophobicity of the exposed residues rather than to the exposed areas of these residues.
In general, N2 molecules aggregated and were adsorbed onto the surface of the SH3 domain at several specific binding sites. Considering non-specific adsorptions of dispersed N2 molecules in the system, this binding could be concluded as the hydrophobic interactions between aggregated N2 molecules and the hydrophobic residues in the key binding sites of the SH3 domain. Furthermore, the occupation of the binding sites of the SH3 domain by aggregated N2 molecules would interfere with ligand binding at these same positions, which would poison the protein function. Our study might provide insights into the potentially common phenomenon of the toxicity of aggregated gases in the biological systems and could provide an alternative way to understand the gas effects in anesthesia.
Methods
The molecular dynamics simulations were carried out with the Gromacs 4.5.1 program package58. In simulations, Amber03 force field was applied in protein and water, the nitrogen atoms were modeled as uncharged Lennard-Jones particles with a radius of cross-section of σNN = 3.260 × 10−1 nm and a depth of the potential well of εNN = 2.888 × 10−1 kJ·mol−1 59. The N-N bond lengths of 1.400 × 10−1 nm were maintained by harmonic potentials with spring constants of 1.280 × 106 kJ/mol·nm2 59. The pdb file of the SH3 domain in the simulations was downloaded from RCSB Protein Data Bank50. Simulations were performed under the isothermal-isobaric (NPT, 1 atm and 310 K to mimic in vivo condition) ensemble with periodic boundary conditions applied in three dimensions. A time step of 1 fs was used and data were collected every 1 ps. The particle mesh Ewald method was used to treat the long-range electrostatic interactions with cut-off of 1 nm60.
References
Fenn, W. O. Inert gas narcosis. Ann. NY Acad.Sci. 117, 760–767 (1965).
Miller, K. W. Inert-gas narcosis, high-pressure neurological syndrome and critical volume hypothesis. Science 185, 867–869 (1974).
Fowler, B., Ackles, K. N. & Porlier, G. Effects of inert-gas narcosis on behavior - a critical-review. Undersea Biomedical Research 12, 369–402 (1985).
Bennett, P. B. & Rostain, J. C. Inert gas narcosis. Bennett and Elliott's Physiology and Medicine of Diving, edited by Brubakk AO and Neumann TS (5th ed.) 300–322 (2003).
Franks, N. P. & Lieb, W. R. Molecular and cellular mechanisms of general anaesthesia (2000).
Trudell, J. R., Koblin, D. D. & Eger, E. I. A molecular description of how noble gases and nitrogen bind to a model site of anesthetic action. Anesthesia and Analgesia 87, 411–418 (1998).
Rostain, J. C. & Balon, N. Recent neurochemical basis of inert gas narcosis and pressure effects. Undersea & Hyperbaric Medicine 33, 197–204 (2006).
Lou, S. T. et al. Nanobubbles on solid surface imaged by atomic force microscopy. Journal of Vacuum Science & Technology B 18, 2573–2575 (2000).
Ishida, N., Inoue, T., Miyahara, M. & Higashitani, K. Nano bubbles on a hydrophobic surface in water observed by tapping-mode atomic force microscopy. Langmuir 16, 6377–6380 (2000).
Koishi, T. et al. Nanoscale hydrophobic interaction and nanobubble nucleation. Phys. Rev. Lett. 93, 4 (2004).
Agrawal, A. et al. Controlling the location and spatial extent of nanobubbles using hydrophobically nanopatterned surfaces. Nano Lett. 5, 1751–1756 (2005).
Zhang, X. H., Khan, A. & Ducker, W. A. A nanoscale gas state. Phys. Rev. Lett. 98, 136101–136104 (2007).
Li, Z. X. et al. Analysis of the gas states at a liquid/solid interface based on interactions at the microscopic level. J. Phys. Chem. B 111, 9325–9329 (2007).
Zhang, L., Chen, H., Li, Z., Fang, H. & Hu, J. Long lifetime of nanobubbles due to high inner density. Science in China Series G-Physics Mechanics & Astronomy 51, 219–224 (2008).
Ducker, W. A. Contact angle and stability of interfacial nanobubbles. Langmuir 25, 8907–8910 (2009).
Craig, V. S. J. Very small bubbles at surfaces-the nanobubble puzzle. Soft Matter 7, 40–48 (2011).
Karpitschka, S. et al. Nonintrusive optical visualization of surface nanobubbles. Phys. Rev. Lett. 109, 066102 (2012).
Ball, P. Nanobubbles are not a superficial matter. ChemPhysChem 13, 2173–2177 (2012).
Zhang, X. H. et al. Detection of novel gaseous states at the highly oriented pyrolytic graphite-water interface. Langmuir 23, 1778–1783 (2007).
Hampton, M. A. & Nguyen, A. V. Accumulation of dissolved gases at hydrophobic surfaces in water and sodium chloride solutions: Implications for coal flotation. Minerals Engineering 22, 786–792 (2009).
Parker, J. L., Claesson, P. M. & Attard, P. Bubbles, cavities and the long-ranged attraction between hydrophobic surfaces. J. Phys. Chem. 98, 8468–8480 (1994).
Doshi, D. A., Watkins, E. B., Israelachvili, J. N. & Majewski, J. Reduced water density at hydrophobic surfaces: Effect of dissolved gases. Proc. Natl. Acad. Sci. U. S. A. 102, 9458–9462 (2005).
Kikuchi, K. et al. Concentration determination of oxygen nanobubbles in electrolyzed water. J. Colloid Interface Sci. 329, 306–309 (2009).
Park, K. H., Chhowalla, M., Iqbal, Z. & Sesti, F. Single-walled carbon nanotubes are a new class of ion channel blockers. J. Biol. Chem. 278, 50212–50216 (2003).
Karajanagi, S. S., Vertegel, A. A., Kane, R. S. & Dordick, J. S. Structure and function of enzymes adsorbed onto single-walled carbon nanotubes. Langmuir 20, 11594–11599 (2004).
Cedervall, T. et al. Detailed identification of plasma proteins adsorbed on copolymer nanoparticles. Angewandte Chemie-International Edition 46, 5754–5756 (2007).
Shen, J.-W., Wu, T., Wang, Q. & Kang, Y. Induced stepwise conformational change of human serum albumin on carbon nanotube surfaces. Biomaterials 29, 3847–3855 (2008).
Zuo, G. H., Huang, Q., Wei, G. H., Zhou, R. H. & Fang, H. P. Plugging into proteins: Poisoning protein function by a hydrophobic nanoparticle. ACS Nano 4, 7508–7514 (2010).
Zuo, G. H., Gu, W., Fang, H. P. & Zhou, R. H. Carbon nanotube wins the competitive binding over proline-rich motif ligand on sh3 domain. J. Phys. Chem. C 115, 12322–12328 (2011).
Calzolai, L., Franchini, F., Gilliland, D. & Rossi, F. Protein-nanoparticle interaction: Identification of the ubiquitin-gold nanoparticle interaction site. Nano Lett. 10, 3101–3105 (2010).
Brewer, S. H., Glomm, W. R., Johnson, M. C., Knag, M. K. & Franzen, S. Probing bsa binding to citrate-coated gold nanoparticles and surfaces. Langmuir 21, 9303–9307 (2005).
Casanova, D. et al. Counting the number of proteins coupled to single nanoparticles. J. Am. Chem. Soc. 129, 12592–12593 (2007).
De, M., You, C. C., Srivastava, S. & Rotello, V. M. Biomimetic interactions of proteins with functionalized nanoparticles: A thermodynamic study. J. Am. Chem. Soc. 129, 10747–10753 (2007).
Dutta, D. et al. Adsorbed proteins influence the biological activity and molecular targeting of nanomaterials. Toxicol. Sci. 100, 303–315 (2007).
Gessner, A., Lieske, A., Paulke, B. R. & Muller, R. H. Functional groups on polystyrene model nanoparticles: Influence on protein adsorption. J. Biomed. Mater. Res. Part A 65A, 319–326 (2003).
Gref, R. et al. ‘stealth’ corona-core nanoparticles surface modified by polyethylene glycol (peg): Influences of the corona (peg chain length and surface density) and of the core composition on phagocytic uptake and plasma protein adsorption. Colloid Surf. B-Biointerfaces 18, 301–313 (2000).
Hong, R. et al. Control of protein structure and function through surface recognition by tailored nanoparticle scaffolds. J. Am. Chem. Soc. 126, 739–743 (2004).
Kaufman, E. D. et al. Probing protein adsorption onto mercaptoundecanoic acid stabilized gold nanoparticles and surfaces by quartz crystal microbalance and zeta-potential measurements. Langmuir 23, 6053–6062 (2007).
Linse, S. et al. Nucleation of protein fibrillation by nanoparticles. Proc. Natl. Acad. Sci. U. S. A. 104, 8691–8696 (2007).
Lundqvist, M., Sethson, I. & Jonsson, B. H. Protein adsorption onto silica nanoparticles: Conformational changes depend on the particles' curvature and the protein stability. Langmuir 20, 10639–10647 (2004).
Ge, C. C. et al. Binding of blood proteins to carbon nanotubes reduces cytotoxicity. Proc. Natl. Acad. Sci. U. S. A. 108, 16968–16973 (2011).
Privalov, P. L. & Gill, S. J. Stability of protein-structure and hydrophobic interaction. Adv. Protein Chem. 39, 191–234 (1988).
Dill, K. A. Dominant forces in protein folding. Biochemistry 29, 7133–7155 (1990).
Nicholls, A., Sharp, K. A. & Honig, B. Protein folding and association - insights from the interfacial and thermodynamic properties of hydrocarbons. Proteins-Structure Function and Genetics 11, 281–296 (1991).
Zhou, R. H., Huang, X. H., Margulis, C. J. & Berne, B. J. Hydrophobic collapse in multidomain protein folding. Science 305, 1605–1609 (2004).
Koch, C. A., Anderson, D., Moran, M. F., Ellis, C. & Pawson, T. Sh2 and sh3 domains - elements that control interactions of cytoplasmic signaling proteins. Science 252, 668–674 (1991).
Pawson, T. & Gish, G. D. Sh2 and sh3 domains - from structure to function. Cell 71, 359–362 (1992).
Kuriyan, J. & Cowburn, D. Structures of sh2 and sh3 domains. Curr. Opin. Struct. Biol. 3, 828–837 (1993).
Feller, S. M., Ren, R. B., Hanafusa, H. & Baltimore, D. Sh2 and sh3 domains as molecular adhesives - the interactions of crk and abl. Trends in Biochemical Sciences 19, 453–458 (1994).
Wu, X. D. et al. Structural basis for the specific interaction of lysine-containing proline-rich peptides with the n-terminal sh3 domain of c-crk. Structure 3, 215–226 (1995).
Ball, L. J., Kuhne, R., Schneider-Mergener, J. & Oschkinat, H. Recognition of proline-rich motifs by protein-protein-interaction domains. Angew. Chem.-Int. Edit. Engl. 44, 2852–2869 (2005).
Ahmad, M., Gu, W. & Helms, V. Mechanism of fast peptide recognition by sh3 domains. Angew. Chem.-Int. Edit. Engl. 47, 7626–7630 (2008).
Steinkellner, G., Rader, R., Thallinger, G. G., Kratky, C. & Gruber, K. Vasco: Computation and visualization of annotated protein surface contacts. BMC Bioinformatics 10, 11 (2009).
ten Wolde, P. R. & Frenkel, D. Computer simulation study of gas-liquid nucleation in a lennard-jones system. J. Chem. Phys. 109, 9901–9918 (1998).
Stillinger, F. H. Rigorous basis of frenkel-band theory of association equilibrium. J. Chem. Phys. 38, 1486–1494 (1963).
Nadassy, K., Wodak, S. J. & Janin, J. Structural features of protein-nucleic acid recognition sites. Biochemistry 38, 1999–2017 (1999).
Heiden, W., Moeckel, G. & Brickmann, J. A new approach to analysis and display of local lipophilicity hydrophilicity mapped on molecular-surfaces. Journal of Computer-Aided Molecular Design 7, 503–514 (1993).
Hess, B., Kutzner, C., van der Spoel, D. & Lindahl, E. Gromacs 4: Algorithms for highly efficient, load-balanced and scalable molecular simulation. J. Chem. Theory Comput. 4, 435–447 (2008).
Rappe, A. K., Casewit, C. J., Colwell, K. S., Goddard, W. A. & Skiff, W. M. Uff, a full periodic-table force-field for molecular mechanics and molecular-dynamics simulations. J. Am. Chem. Soc. 114, 10024–10035 (1992).
Essmann, U. et al. A smooth particle mesh ewald method. J. Chem. Phys. 103, 8577–8593 (1995).
Acknowledgements
We thank Jun Hu, Chun-Lei Wang, Wen-Peng Qi, Bo Zhou and Shen Wang for helpful discussion and comments. This work was supported by The Supercomputing Center of Chinese Academy of Sciences and Shanghai Supercomputer Center. MZ and HF acknowledge the funding support from National Science Foundation of China under grant (11290164, 11175230). YG Thanks for the startup funding support from Shanghai Institute of Applied Physics (Y290011011) and “Hundred People Project” from Chinese Academy of Sciences.
Author information
Authors and Affiliations
Contributions
H.P.F. and J.X.C. contributed most of the ideas. H.P.F., Y.G. and M.Z. designed simulations. Y.G. and M.Z. performed most of the numerical simulations. Y.G., M.Z. and G.H.Z. carried out theoretical analysis and H.P.F., Y.G. and M.Z. wrote the paper. All authors discussed the results and commented on the manuscript.
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Electronic supplementary material
Supplementary Information
Supplementary Information
Rights and permissions
This work is licensed under a Creative Commons Attribution-NonCommercial-ShareALike 3.0 Unported License. To view a copy of this license, visit http://creativecommons.org/licenses/by-nc-sa/3.0/
About this article
Cite this article
Zhang, M., Zuo, G., Chen, J. et al. Aggregated Gas Molecules: Toxic to Protein?. Sci Rep 3, 1660 (2013). https://doi.org/10.1038/srep01660
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/srep01660
This article is cited by
-
Initial growth dynamics of 10 nm nanobubbles in the graphene liquid cell
Applied Nanoscience (2021)
-
Frequency-dependent absorbance of broadband terahertz wave in dense plasma sheet
Applied Physics B (2018)
-
A new association state of solutes in nanoconfined aqueous solutions
Science China Physics, Mechanics & Astronomy (2016)
-
Exploring the Effects on Lipid Bilayer Induced by Noble Gases via Molecular Dynamics Simulations
Scientific Reports (2015)
-
Effects on lipid bilayer and nitrogen distribution induced by lateral pressure
Journal of Molecular Modeling (2015)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.