ABSTRACT
Transfection of cDNA in 3′untranslated region of human nuclear factor for interleukin-6 (NF-IL6 3′UTR) induced tumor suppression in a human hepatoma cell line. cDNA array analysis was used to reveal changes in gene expression profile leading to tumor suppression The results indicate that this suppression was not due to activation of dsRNA-dependent protein kinase, nor to inactivation of oncogenes; rather, all the changes in expression of known genes, induced by NF-IL6 3′UTR cDNA may be ascribed to the suppression of cellular malignancy. Therefore, our results imply that this 3′untranslated region may have played role of a regulator of gene expression profile.
Similar content being viewed by others
INTRODUCTION
Studies on the suppression of malignancy of tumor cells are important for the understanding of the mechanism of tumor suppressors and for the establishment of novel therapies of tumors. Since the last decade of the 20th century, several 3′untranslated regions of eukaryotic mRNAs were found to have in vitro tumor suppressor activity. They caused phenotypic reversion (lowering of tumorigenicity) in cultured tumor cells or malignant cells1, 2, 3, 4, 5, 6. However, the molecular mechanisms of these tumor suppressor effects remain elusive.
Several years ago, a regulatory element with tumor suppressor activity, NF-IL6 3′UTR, was found, which caused tumor suppression in a part of mouse malignant DT cells1, 2. Recently, in order to test its tumor-suppressive effect in different human tumor cell lines, we transfected it into SMMC7721 cells, a human hepatocarcinoma cell line, and resulted in markedly reduced malignancy (reversion) in significant part of the transfected cell clones. Then, we determined the changes in gene expression profile of a typical revertant clone by using the cDNA array analysis, to pave the way for elucidating possible molecular mechanisms of this tumor suppression. In this report, we will present results of this analysis.
MATERIALS AND METHODS
Cell lines and plasmids
SMMC7721, purchased from Cell Bank of Chinese Academy of Sciences, Shanghai, is a typical hepatoma cell line7. p14-6 is a recombinant pcD2 plasmid harboring a middle part of the 3′untranslated region of the human NF-IL6 cDNA4, stored in our lab (GenBank accession number M98056). pcDmr Cl 2 is a plasmid containing the same genes as p14-6, but harbors no insert.
Transfection of cell lines and malignancy tests
Stable transfection was performed by using the Calcium phosphate transfection kit (Invitrogen) according to the instruction manual. The concentration of cytotoxic antibiotic G418 (Sigma) was 2 mg / ml (valence 1 mg/ml) for all cells, including controls. G418R clones appeared in 1-2 week after screening. Cell growth curves determination and soft agar tests were performed according to our previous methods1. Nude mice tumorigenicity tests were performed with the Balb/c nu/nu nude mice, 4 -6 week age (from the Shanghai Center for Experimental Animals, Chinese Academy of Sciences), which were kept in the sterile rooms of Animal house of our Institute. 5× 106 cells per point were injected subcutaneously into the back of animals. Positive tumorigenicity was defined as the appearance of palpable tumor nodules in the injection points one month postinjection.
cDNA array analysis
The cDNA array analysis were done in CASarray Co., Shanghai. The cells were directly lysed in culture bottles with TrizolR (Invitrogen), then subjected to further mRNA purification, cDNA synthesis and hybridizations. The cDNA array membanes used contained 11,557 known genes and ESTs, and every gene had two spots. The analysis was performed twice to guarantee reproducible results, and passed strict quality control.
Semi-quantitative RT-PCR
6 genes, including 3 up-regulated and 3 down-regulated as revealed by cDNA array analysis, were chosen to compare their cDNA amounts by using semi-quantitative RT-PCR. The primers were designed using OLIGO version 6.44 software. For control, cDNA of ribosomal protein gene S29, a well-known housekeeping gene, was used. Reverse transcription of total RNAs from Cl 1, SMMC7721 and pcDmr was done, respectively, using Superscript II reverse transcriptase (Life Technologies and equal amounts (1 μl) of RT mixture were used in a 50 μl PCR reaction containing various primer pairs and FD Taq DNA polymerase (Fuhua Co., Shanghai) under the same conditions. The intensities of bands were measured using Tanon gel electrophoresis analysis software, and the values of their strengths relative to S29 were calculated by dividing their measured band intensities by the average S29 band intensities of RT PCR products of corresponding cell lines. The results were shown in a vertical square graph.
RESULTS AND DISCUSSION
Establishment of the NF-IL6 transfectants of SMMC7721 cell line
The NF-IL6 3′UTR cDNA (plasmid p14-6) was stably transfected into SMMC7721 cells by using standard calcium phosphate procedure. It was found that this hepatoma cell line has a certain endogenous G418 resistance, and thus a higher concentration (2 mg/ml) of G418 was used for screening. This drug concentration led to the reduction in the number of G418R clones; however, all the transfectants contained plasmids, as proved by Southern blots.
Phenotypic characteristics of the NF-IL6 3′UTR transfectants of SMMC7721
By high concentration G418 screening, 14 independent clones were obtained, 6 of which were found to be contaminated by mycoplasma and excluded. The remaining 8 clones were used in experiments.
These p14-6 transfectants had morphology similar to their parent cells. The majority of clones grew slower than SMMC7721 (see Fig 1B-1) and had reduced ability to form colonies in soft agar medium (Tab 1 and Fig 1A; in the figure, only Cl 1 and two controls were shown). In the in vivo tumorigenicity tests, those transfectant clones that formed less colonies in soft agar also appeared to have reduced tumorigenicity (Tab 2). Therefore, these 7 transfectants were revertants. Southern blots with 32P-radiolabeled NF-IL6 3′UTR cDNA as probe showed that the NF-IL6 3′UTR was inserted into different positions of the cellular genome (Fig 2a). Northern blots with the same probe showed that the 3′UTR was highly expressed (Fig 2B). The remaining clone, clone 8, did not revert, possibly due to some cellular reasons. In contrast, the control pcDmr Cl 1-Cl 4 transfectants (4 clones) showed tumorigenicity non-distinctive from SMMC7721 cells (Tab 2), although two cell clones (clones 1 and 3) grew slower than the former (Fig1B-2). Interestingly, the expression of endogenous NF-IL6 gene had no significant changes at both transcriptional (Fig 2B) and translational levels as shown by Western blotting (data not shown).
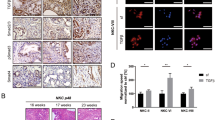
Growth characteristics of cell clones. A, morphology (upper) and soft agar colonies (lower) of SMMC7721 (1), revertant Cl 1 (2) and control pcDmr Cl 2 (3). Bars: 10 μm. B, growth curves of all cell clones. 1, growth curves of 8 NF-IL6 3′UTR transfectants. The signs in the curves are: ▵, Cl 1; iõ iõ, Cl 2; •, Cl 3; ió ió, Cl 4; ið, Cl 5; ▿, Cl 6; ♦, Cl 7; iö, Cl 8. 2, growth curves of control pcDmr clones. The signs in the curves are: ió, pcDmr Cl 1; iõ, pcDmr Cl 2; ▵, pcDmr Cl 3; ▸ ▸, pcDmr Cl 4. In both figures the thicker curves with ▾ signs are growth curves of SMMC7721.
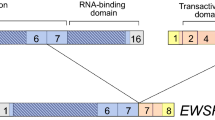
Integration status and expression of NF-IL6 3′UTR. The probe for all hybridizations was 32P-labeled 3′UTR fragment. In both figures, 1-8 stand for transfectants Cl 1-8; S, SMMC7721; I - IV, pcDmr Cl 1-4. A. Southern hybridization of cellular genomic DNAs cut by EcoRI (which had no sites on p14-6 plasmid). B. Northern hybridization of cellular total RNAs. N, endogenous NFIL6 transcript (∼ 2 kb). U, the 3′UTR transcript (∼ 0.5 kb). GAPDH, the same membrane re-probed by GAPDH probe.
Transcriptome analysis of the NF-IL6 3′UTR revertant
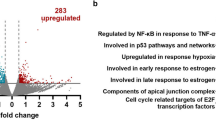
To obtain results as clear as possible, one typical revertant, Cl 1, was chosen for the cDNA array analysis to compare with the SMMC7721 cells and with one of the control transfectant, pcDmr Cl 2. For analysis, the genes that were consistently up- and down-regulated in Cl 1 only, but were differently regulated in SMMC7721 and pcDmr Cl 2, were considered to be induced by NFIL6 3′UTR. By this criterion, the genes, whose different expression was induced by NF-IL6 3′UTR, are shown in Tab 3. For clarity, the average expression ratios between Cl 1 and SMMC7721 cells only are shown. The semi-quantitative RT-PCR results are shown in Fig 3, which are in agreement with cDNA array analysis qualitatively (Tab 3 and Fig 3).
RT-PCR results of 3 up-regulated genes: S59049, AJ133355 and NM_020233, and 3 down-regulated genes: AF012281, M26326 and D31885, in the Cl1, SMMC7721 and pcDmr cell lines, respectively. The ordinate shows ratios of the amplified band strengths of various gene transcripts to the average band strength of the amplified S29., S59049; B, AJ133355; C, NM_020233; D, AF012281; E, M26326; F, D31885.
Among up-regulated genes, first two genes (p53-in duced ribonucleotide reductase small subunit 2 homolog, and regulator of G protein signalling 1) were directly or indirectly related to tumor suppression, and their enhanced expression favored this suppression. Among those genes that were down-regulated, the first gene (PDZ domaincontaining protein 1) is reported to be overexpressed in some epithelial tumors12; therefore, its reduced expression may contribute to the tumor suppression. The functions of other down-regulated genes include resistance to apoptosis (keratin 18), activation of signal transduction (ADP-ribosylation factor 6-like interacting protein), and activation of cell motility (thymosin β4) etc; clearly, their down-regulated expression are unfavorable for tumor growth and metastasis. Therefore, the changes in the expression of above genes, induced by the transfected NF-IL6 3′UTR, all favored to the reversion of cellular phenotype. No oncogene, that was detectably up- or down-regulated, was found by the cDNA array analysis.
We have also performed analysis on expressed proteins of these cell lines by using 2D electrophoresis (will be published elsewhere). The majority of results of 2D electrophoresis were different with cDNA array results. For example, maspin18 and annexin III19 were found up-regulated in 2D analysis. Besides technical reasons, this difference may suggest that the expression of a gene from transcription to final protein product is more complicated than that anticipated by our present knowledge.
Possible mechanism(s) of NF-IL6 3′UTR tumor suppression
It might be thought that some oncogenes were inactivated by the insertion of transfected plasmids, leading to tumor suppression. However, this does not seem to be the case, because no oncogene was found differentially and significantly expressed.
The activation of dsRNA-dependent protein kinase (PKR) by 3′UTR RNA22 was used to explain the tumor suppressor effect of α-tropomyosin; however, this explanation is not satisfactory enough, because, first, some 3′UTRs of mRNA, like the 3′UTR of NF-IL6, do not have stable double stranded structure; second, as the activation of PKR leads to general suppression of protein synthesis, the up-regulation of protein expression (like the maspin and annexin III in our case) is hard to be explained.
To sum up, we are inclined to believe that the transfection of NF-IL6 3′UTR cDNA might possibly play some regulatory role for the expression of a group of genes, either in up- or down-regulated ways. It is well known that the 3′UTRs are sites for interactions with trans-acting factors, and these interactions, as a whole, may be responsible for the maintenance of normal cell life. Therefore, the exogenous 3′UTR might also act as a regulator of gene expression. If such supposition is correct, its molecular mechanism of tumor suppression should be found in the analysis of its regulatory functions.
Abbreviations
- NF-IL6:
-
nuclear factor for interleukin-6 expression
- PKR:
-
double-stranded RNA-dependent protein kinase
- 3′UTR:
-
3′untranslated region
References
Liu DG, Noda M, Wang D, Chen ZZ, Li ZP, Ikawa Y . A cDNA clone with antioncogene activity. Sci China (Ser B) 1991; 7:730–7 [in Chinese]; Sci.China (Ser B) 1992; 35:822–33 [in English].
Liu DG, Li ZP, Akira S, Kishimoto T . Overexpression of a reversion-related protein in the revertant RR cells. Sci China (Ser B) 1995; 35:372–8 [in Chinese]; Sci China (Ser C) 1996; 39:300–09 (in English).
Malumbress M, Perez de Castro I, Santos J, Fernandez Piqueras J, Pellicer A . Hypermethylation of the cell cycle inhibitor p14INK4b 3′untranslated region interferes with its transcription regulation in primary lymphomas. Oncogene 1999; 18:385–96.
Rastinejad F, Conboy MJ, Rando TA, Blau HM . Tumor suppression by RNA from the 3′untranslated region of alphatropomyosin. Cell 1993; 75:1107–17.
Jupe ER, Liu XT, Kiehlbauch JL, McClung JK, Dell'Orco, RT . The 3′untranslated region of prohibitin and cellular immortalization. Exp Cell Res 1996; 224:128–35.
Amara FM, Smith GM, Kuschak TI, Takeuchi TL, Wright JA . A cis-trans interaction at the 3′untranslated region of ribonucleotide reductase mRNA is regulated by TGF-beta 1, TGF-beta 2 and TGF beta 3, Biochem Biophys Res Commun 1996; 228:347–51.
Dong RC, Zhou RH, Lu FD, Tao WZ . Establishment of SMMC7721, a human hepatocarcinoma cell line, and observations on its biological characteristics. Journal of 2nd Military Medical University (Shanghai), 1980; 1:5–9 [in Chinese].
Xu XR, Huang J, Xu ZG, Qian BZ, Zhu ZD, Yan Q, Cai T, Zhang X, Xiao HS, Qu J, Liu F, Huang QH, Cheng ZH, Li NG, Du JJ, Hu W, Shen KT, Lu G, Fu G, Zhong M, Xu SH, Gu WY, Huang WT, Zhao XT, Hu GL, Gu JR, Chen Z, Han ZG . Insight into hepatocellular carcinogenesis at transcriptome level by comparing gene expression profiles of hepatocellular carcinoma with those of corresponding noncancerous liver. Proc Natl Acad Sci USA 2001; 8:15089–94.
Tanaka H, Arakawa H, Yamaguchi T, Shiraishi K, Fukuda S, Matsui K, Takei Y, Nakamura Y . A ribonucleotide reductase gene involved in a p53-dependent cell-cycle checkpoint for DNA damage. Nature 2000; 404:42–9.
Druey KM, Blumer KJ, Kang VH, Kehr JH . Inhibition of Gprotein-mediated MAP kinase activation by a new mammalian gene family. Nature 1996; 404:742–6.
Hedvat CV, Irving SG . The isolation and characterization of MINOR, a novel mitogen-inducible nuclear orphan receptor. Mol Endocrinol 1995; 9:1692–70.
Kocher O, Comella N, Tognazzi K, Brown LF . Identification and partial characterization of PDZK1: a novel protein containing PDZ interaction domains. Lab Invest 1998; 78:117–25.
Sorom AJ, Nyberg SL, Gores GJ . Keratin, fas, and cryptogenic liver failure. Liver Transpl 2002; 8:1195–97.
Houldsworth J, Attardi G . Two distinct genes for ADP/ATP translocase are expressed at the mRNA level in adult human liver. Proc Natl Acad Sci USA 1988; 85:377–81.
Tang Y, Katuri V, Dillner A, Mishra B, Deng CX, Mishra L . Disruption of transforming growth factor-β signaling in ELF β-spectrin-deficient mice. Science 2003; 299:574–7.
Huff T, Muller CS, Otto AM, Netzker R, Hannappel E . beta-Thymosins, small acidic peptides with multiple functions, Int. J. Biochem. Cell Biol 2001; 3:205–20.
Argraves WS, Suzuki S, Arai H, Thompson K, Pierschbacher MD, Ruoslahti E . Amino acid sequence of the human fibronectin receptor. J Cell Biol 1987; 105:1183–90.
Maass N, Hojo T, Zhang M, Sager R, Jonat W, Nagasaki K . Maspin-a novel protease inhibitor with tumor-suppressing activity in breast cancer. Acta Oncol 2001; 39:931–4.
Niimi S, Oshizawa T, Yamaguchi T, Harashima M, Seki T, Ariga T, Kawanishi T, Hayakawa T . Specific expression of annexin III in rat-small-hepatocytes, Biochem. Biophys. Res Commun 2003; 300:770–4.
Frosch BA, Berquin I, Emmert-Buck MR, Moin K, Sloane BF . Molecular regulation, membrane association and secretion of tumor cathepsin B. APMIS 1999; 107:28–37.
Strack S, Kini S, Ebner FF, Wadzinski BE, Colbran RJ . Differential cellular and subcellular localization of protein phosphatase 1 isoforms in brain. J Comp Neurol 1999; 413:373–84.
Davis S, Watson JC . In vitro activation of the interferon-induced, double-stranded RNA-dependent protein kinase PKR by RNA from the 3′ untranslated regions of human alpha-tropomyosin. Proc Natl Acad Sci USA 1996; 93:508–13.
Denecke B, Meyerdierks A, Bottger EC . RGS1 is expressed in monocytes and acts as a GTPase-activating protein for G-protein-coupled chemoattractant receptors. J Biol Chem 1999; 274:26860–8.
Acknowledgements
We thank Feng LIU and Jin Mei ZHANG for routine keeping of nude mice, Ying Zi GE for part of routine cell culture, and Gen Jun XU for encouragements. This work was supported by the National Natural Science Foundation of China grant (No. 39970172), and by the Creation Foundation from Shanghai Intsitutes for Biological Sciences.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
LIU, D., JIANG, Q., WEI, Y. et al. Gene expression profile favoring phenotypic reversion: a clue for mechanism of tumor suppression by NF-IL6 3′UTR. Cell Res 13, 509–514 (2003). https://doi.org/10.1038/sj.cr.7290195
Received:
Revised:
Accepted:
Issue Date:
DOI: https://doi.org/10.1038/sj.cr.7290195