Abstract
Microbial inoculants are attracting growing interest in agriculture, but their efficacy remains unreliable in relation to their poor survival, partly due to the competition with the soil resident community. We hypothesised that recurrent inoculation could gradually alleviate this competition and improve the survival of the inoculant while increasing its impact on the resident bacterial community. We tested the effectiveness of such strategy with four inoculation sequences of Pseudomonas fluorescens strain B177 in soil microcosms with increasing number and frequency of inoculation, compared to a non-inoculated control. Each sequence was carried out at two inoculation densities (106 and 108 cfu.g soil−1). The four-inoculation sequence induced a higher abundance of P. fluorescens, 2 weeks after the last inoculation. No impact of inoculation sequences was observed on the resident community diversity and composition. Differential abundance analysis identified only 28 out of 576 dominants OTUs affected by the high-density inoculum, whatever the inoculation sequence. Recurrent inoculations induced a strong accumulation of nitrate, not explained by the abundance of nitrifying or nitrate-reducing microorganisms. In summary, inoculant density rather than inoculation pattern matters for inoculation effect on the resident bacterial communities, while recurrent inoculation allowed to slightly enhance the survival of the inoculant and strongly increased soil nitrate content.
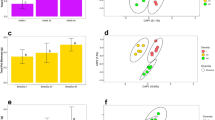
Similar content being viewed by others
Introduction
Synthetic fertilisers and pesticides have played an essential role in achieving contemporary agricultural yields. However, the continuous and excessive application of synthetic fertilizers and pesticides is a growing concern due to their potential adverse effects on human health and ecosystems1,2,3. Microbial inoculants represent promising sustainable alternatives to agrochemical inputs as they offer benefits in terms of crop yield and nutrient use efficiency4, plant tolerance to biotic and abiotic stresses5, resistance to pathogens6 and adaptation to climate change7. This growing interest in microbial inoculants over the last decade is reflected both by an increase in studies on the subject8 and by substantial investments for their development9. Nevertheless, translation of their beneficial effects from controlled conditions to field applications presents many challenges10 including their unreliable survival in soil following application11. Thus, numerous studies have reported a steady decline of inoculants in soil after application with their persistence depending on a variety of biotic and abiotic factors and thus severely impacting their efficacy12,13.
Invasion ecology focuses on the factors that allow invasive species to survive and thrive in a new environment as well on their impact on native ecosystems. As such, an increasing number of studies are adopting an invasion ecology approach to unravel the mechanisms underlying the survival of microbial inoculants. In this context, a microbial inoculant is considered as an alien species introduced into a resident community which has to overcome both abiotic and biotic hurdles before its establishment14. Soil type and diversity are known to affect the invasibility of an environment15. Specifically, a greater diversity of the resident community along with the presence of closely related species can increase the propensity of niche overlap between the soil community and the invader, therefore hampering invasion as suggested by the Darwin’s naturalisation hypothesis16,17,18,19,20. On the contrary, it has also been posited that invaders closely-related to resident species would be more likely to establish due to shared pre-adaptation with resident species21,22.
While diversity and composition of the resident community modulate invasion success, invaders generally affect this resident community in return23,24, with both compositional and functional changes, including shifts in nitrogen cycling25,26,27. In addition, the impact of microbial inoculations on the resident soil microbial communities has often been overlooked despite calls for a more comprehensive assessment of the consequence of microbial inoculation28,29. Mallon et al.30 showed that even transient and unsuccessful bacterial invasion can cause changes in the diversity, structure and niche breadth of the invaded bacterial community, and they specifically observed a shift in the community niche away from the resources used by the invader. According to Darwin’s naturalisation hypothesis, this shift could in turn favour further invasions by the same strain as the competition for resources will be less intense as hypothesised by Mallon et al.30.
In this work, we sought to assess to which extent inoculation frequency is affecting the inoculant survival, as well as its impact on the resident soil microbial community. We tested the hypothesis stated by Mallon et al.30 that recurrent inoculations of the same strain would affect the resident community and enhance the survival and establishment of the inoculant. To test this hypothesis, we inoculated soil microcosms with Pseudomonas fluorescens B177 at two densities with four sequences of inoculation ranging from one to four inputs compared with a non-inoculated control. We then monitored the survival of the inoculated strain and assessed the impact of recurrent inoculations on the composition and diversity of the resident bacterial community as well as on microbial functions. The nitrogen cycle being an essential agrosystemic service in agriculture and a common indicator of soil quality31, we chose it as the model function in this study.
Results
Inoculant survival following single or recurrent inoculations
Pseudomonas fluorescens was not detected in the non-inoculated controls. In 55% of the low inoculum density samples (106 cfu g−1 dry soil), the strain was detected but under the quantification limit while it was always above the quantification limit in the high inoculum density samples (108 cfu g dry soil−1). Therefore, only the high inoculation treatment was included in the quantitative analysis. P. fluorescens abundance was up to 1.3 × 107 and remained above 106 copy g−1 dry soil in all inoculation sequences until the end of the experiment. At week 10, P. fluorescens abundance was significantly higher in Sequence 4 (four inoculations) compared to Sequence 1 (one inoculation) (Tukey’s test, p = 0.03, Fig. 1). However, this difference was not significant anymore at week 14, corresponding to a transient effect of the recurrent inoculations. The abundances of P. fluorescens in Sequences 2 and 3 (two inoculations, 2 or 6 weeks apart) were not significantly different from that in Sequences 1 and 4.
Survival of the inoculant. Abundance of Pseudomonas fluorescens across inoculation sequences (1, 2, 3 and 4) over time (week 4, 10 and 14) for high inoculum density (108 cfu g dry soil−1). Values represent the log copy number per gram of dry soil. Letters above the bars indicate significant differences between sequences for each time block according to Tukey’s test (p = 0.03).
Diversity and composition of the indigenous bacterial community
Analysis of the alpha diversity of the resident soil bacterial community showed no significant differences between the non-inoculated and inoculated soils (Fig. 2). However, high density inoculations resulted in a significant decrease of the richness, estimated as the number of observed OTUs (Tukey; p < 0.001), at both weeks 10 and 14, whereas no effect was observed on the Faith’s PD and Simpson reciprocal index (Supplementary Fig. 2a,b). While the Simpson reciprocal index remained stable in the non-inoculated treatment, a higher Simpson reciprocal index was observed at week 14 compared to week 10 in the inoculated microcosms (Tukey; p = 0.01, Supplementary Fig. 2c).
Diversity across the inoculation sequences. Number of observed OTUs and Simpson’s Reciprocal index at week 4, 10 and 14 for the four inoculation sequences and NI control. Bright colour stands for high density of inoculation (108 cfu g dry soil−1) and light colour for low density of inoculation (106 cfu g dry soil−1). Differences between groups were tested using ANOVAs followed by Tukey’s honestly significant difference test (p value < 0.05).
We found a significant though weak effect of the inoculation density on the structure of the bacterial community (PERMANOVA nperm = 999, R2 = 0.014, p = 0.017) and no effect of the inoculation sequence (PERMANOVA nperm = 999, R2 = 0.012, p = 0.22) based on both weighted and unweighted UniFrac distances (Supplementary Fig. 3). Thus, the structure of the bacterial community differed significantly between low and high inoculum density without any difference compared to the non-inoculated treatment (PERMANOVA nperm = 999, R2 = 0.043, p = 0.001).
Impact of the inoculation sequence treatments on the relative abundance of autochthonous OTUs
For a deeper insight into the effect of inoculations on the resident bacterial taxa, we used a generalised linear mixed model to identify OTUs exhibiting significant differences in relative abundance between the inoculated treatments and the non-inoculated control (p < 0.05). Only 28 out of the 576 dominant OTUs were significantly affected by P. fluorescens across the different inoculation sequences at week 10. While 9 of the impacted OTUs are shared by all inoculation sequence treatments, sequences 3 and 4 also affected 9 and 13 other OTUs as compared to 2 and 5 other OTUs only affected in the Sequences 1 and 2 (Fig. 3a). The total number of impacted OTUs decreased at week 14 (12 OTUs affected) with once again an increasing number of impacted OTUs from Sequence 1 to 4. Interestingly, a majority of OTUs significantly affected by inoculation exhibited increased relative abundances, especially in the high-density treatments (Fig. 3b).
Number of OTUs affected by each inoculation Sequence. (a) Venn diagrams indicate the number of OTUs whose abundance is significantly different from the NI control for each inoculation sequence at week 10 and week 14 after high inoculation density (108 cfu g dry soil−1). (b) The changes in abundances whether positive or negative compared to the NI control for all inoculation sequences and both high and low inoculation density (108 and 106 cfu g dry soil−1 respectively).
For example, at week 10, 16 OTUs exhibited increases in relative abundance compared to the non-inoculated control whatever the inoculation sequence while, at the same time, 7 OTUs were negatively affected. Most of these impacted OTUs belonged to the Gammaproteobacteria phyla (38.5%) and one of them was assigned to Pseudomonas fluorescens (OTU 9). Interestingly, this OTU 9 exhibited a 99% sequence identity with the inoculated Pseudomonas fluorescens B177 strain. Its relative abundance increases with recurrent inoculations and was strongly and significantly correlated to the abundance of P. fluorescens B177 measured by qPCR (Pearson’s r = 0.88, p < 10–16) (Supplementary Fig. 4).
Recurrent inoculation has a strong influence on the NO3 − pool
Ammonium concentrations remained low at every sampling time (below 3.17 µg NO3− g−1 dry soil) whereas nitrate concentrations exceeded 30 µg NO3− g−1 dry soil at all timepoints. At week 10, we found a significant effect of the inoculation sequence at high inoculation density with an increase of NO3− in Sequences 2, 3 and 4 compared to inoculation sequence 1 and NI control (Fig. 4, p = 10–14). At week 14, significantly higher NO3− levels were still observed in the inoculation sequences 3 and 4 (p = 10–10) but no longer in Sequence 2. When comparing with NO3− concentrations at week 0, the recurrent inoculation of P. fluorescens at high density led to an accumulation of NO3− in the soil that persisted at least for 6 weeks after the last inoculation. In contrast, at low inoculum density, NO3− concentrations were similar between non-inoculated and inoculated treatments.
Nitrate accumulation in soil. Nitrate content (µg NO3− g dry soil−1) at weeks 0, 10 and 14 for the four inoculation sequences and Ni control, after low (light colour) and high (bright colour) inoculation density. Letters above indicate significant differences between inoculation density and sequences according to Tukey’s test (p ≤ 10–10 at both week 10 and 14).
Abundances of functional genes involved in nitrogen cycling
Quantitative real-time PCR assays were performed to quantify the abundances of 16S rRNA, bacterial narG, napA and amoA genes, as well as archaeal amoA genes as molecular markers for the microbial communities involved in NO3− production and consumption. Inoculation density triggered only sporadic increases in the abundance of the narG and napA genes at week 10, however these increases were not significant when compared to the Ni control (Fig. 5). In contrast, the inoculation sequences induced more pronounced abundance changes with similar pattern across genes. At week 10, Sequences 2, 3 and 4 led to increased abundance of AOA, napA and narG as compared to Sequence 1, and also to the Ni control for napA and narG. Sequence 4 also increased AOB abundance at week 10 as compared to Sequence 1. At week 14, Sequences 1 and 4 increased the abundances of narG, napA and AOB as compared to Sequence 3 without significant difference compared to the control. We then investigated whether the increased NO3− levels in the inoculation sequences 2, 3 and 4 were related to changes in the abundance of these N-cycling communities. However, we did not find any significant correlation.
Functional impact of the inoculation sequences and density on the microbial community. Abundance of ammonia-oxidizers (AOB, AOA) and nitrate reducers (napA and narG) at weeks 0, 10 and 14 for the four inoculation sequences and Ni control, after low (light colour) and high (bright colour) inoculation density. Letters above indicate the significant differences between inoculation sequences (black letters) and a combined variable of inoculation sequence and density (grey letters) according to Tukey’s test (p < 0.005).
Discussion
Bacterial inoculation studies have often focused on the importance of the strain genotype32, root colonisation ability33,34, soil type35 or abiotic perturbations36,37 to identify the determinants of a successful inoculation. However, the resident microbial community also plays a crucial role in the fate of the inoculant38 and in turn, bacterial invasion can induce shifts in the resident microbial community that can influence the success of future inoculations30. Hence, previous inoculations could improve the maintenance capacity of a microbial inoculum in soil. Four strategies of recurrent inoculation were established at two inoculum densities to assess their effectiveness in promoting the survival of a bacterial inoculant and to determine the extent of their impact on the resident bacterial soil community.
Inoculum density rather than recurrent inoculation improved the survival of the inoculant
At low inoculum density, the abundance of P. fluorescens B177 decreased with time to end up close to the detection limit with higher variability, which is inherent to the qPCR methodology at low copy number39. We therefore determined a quantification limit under which detection is possible but not quantification, as recommended in previous studies40,41,42. The high density of the bacterial inoculum used in our experiment exceeded field application by at least 5 orders of magnitude. For instance, commercial indication for P. fluorescens B177 inoculation is around 30 cells/g of soil under wettable powder form at 108 viable cells/g, a common concentration to many other inoculant formulations43,44. Inoculation of P. fluorescens B177 at high density corresponded to 4.4% of the bacterial community (and 0.04% for the low density), which represents a substantial propagule pressure compared with the abundance of other OTUs. Propagule pressure, defined as the product of the number of invaders by the frequency of invasion events, has been shown to enhance the invasion success45 primarily due to a lower probability of stochastic extinction of the invaders46. Our study supports these findings since P. fluorescens abundance remained above 106 copy/g of dry soil until week 14 when inoculated at high density while it declined below the detection limit at low inoculum density. Amor et al.47 demonstrated that a high propagule size favours invasion success when invaders exhibit a negative growth rate while a high propagule frequency is advantageous in the case of positive growth rate. This aligns with our study which shows a decline of P. fluorescens B177 over time, along with a much stronger effect of inoculum density than inoculum frequency. Yet, four recurrent inoculations (Sequence 4) transiently led to higher P. fluorescens abundance at week 10 at high inoculation density. This supports our main hypothesis that recurrent inoculation can improve the survival of the inoculum, but only the highest inoculation frequency was effective, and the benefit was rather limited.
Diversity and composition of the resident bacterial community were weakly affected by recurrent inoculations
As bacterial invasions can affect the resident soil microbial community23, we assessed the impact of recurrent inoculation on the soil bacterial community. Previous studies have shown that even transient or unsuccessful invasion can induce a shift in the resident community structure and niche30,48. We observed a limited impact of the inoculant on the diversity and composition of the soil bacterial community despite the inoculum densities used (108 and 106 cfu g soil−1, representing 4.4 and 0.04% of the community, respectively). The higher inoculum density caused a small but significant decrease in richness as well as an increase of evenness along with changes in the relative abundance of 28 dominant OTUs (Fig. 3). There is no consensus on the effect of inoculation on the diversity of soil microbial community, with some studies reporting a diversity loss49 while others observed an increase in diversity30,50 or no effect at all51. The observed increase in evenness could be explained by the release of nutrients due to the death of the inoculant and favouring the growth of less abundant taxa14. This is supported by the results of the differential abundance analysis showing that most affected OTUs increased in abundance following inoculation (Supplementary Fig. 3). The lower richness in the inoculated treatments could be due to the high proportion of the inoculant representing up to 2.3% of the community at week 10, which can displace numerous rare taxa that will no longer be included in a rarefied data set.
Numerous studies have assessed the impact of single microbial inoculation on the resident soil community24 but less is known about the impact of recurrent inoculations. Despite the fact that the recurrent inoculation also increased the number of invaders, we did not observe any significant effect of the recurrent inoculation sequences on either the composition or the diversity of the resident soil bacterial community. This means that subsequent inoculations in Sequences 2, 3 and 4 did not further affect bacterial diversity beyond that the first inoculation at high density already had. Wang et al.52 also reported that the first inoculation with a consortium of four bacterial strains plays a greater role in influencing the community than any subsequent inoculations. In contrast, some studies reported no impact of inoculation of different strains of P. fluorescens on the soil microbiome at community level53,54. Nevertheless, when looking at the population level, the relative abundance of some OTUs was significantly affected (28 out of 576 dominant OTUs) with more OTUs affected as the number and frequency of inoculations increased. However, this inoculation effect on very few OTUs rather than on the global resident community underscores its remarkable resistance to invasion by Pseudomonas fluorescens B177.
There are two opposing theories concerning the interactions between an invader and closely related species in the resident community. Competition-relatedness hypothesis suggests that invasion is restricted due to overlapping of resource use and shared predators and pathogens55. On the contrary, it has also been postulated that a higher relatedness between invaders and resident population can promote invasion through shared mutualists, facilitation and likelihood to thrive in similar conditions56. Interestingly, in our study, most affected OTUs showed increased relative abundances, suggesting that positive interactions occurred between P. fluorescens and some members of the resident communities which favoured their relative fitness. Notably, among all impacted OTUs, the sharpest increase (i.e., 107-fold increase in relative abundance) was observed for one OTU affiliated to Pseudomonas fluorescens species and having 99% identity with the 16S gene of P. fluorescens B177. Along with the strong correlation between this OTU and the qPCR detection of the inoculant, this strongly suggests that this OTUs corresponds to the inoculated strain. However, due to the methodological limitation related to the sequencing of partial 16S rRNA it is not possible to rule out the possibility that OTU 9 also comprises a few other Pseudomonas fluorescens.
Soil nitrate concentration was strongly impacted by the recurrent inoculations
We investigated whether recurrent inoculations had an impact on soil microbial functions, focusing on nitrogen cycling. Significant and persistent effects were observed with inoculation Sequences 3 and 4 at high density which led to increased soil nitrate concentrations at both weeks 10 and 14. Similar finding have been reported in previous studies, where shifts in soil microbial activities such as chitinase, urease or phosphatase were observed in response to microbial inoculation57,58,59. It is unlikely that P. fluorescens B177 had a direct effect on nitrate concentration because its genome does not contain any of the genes required for ammonia-oxidation (amoA) or dissimilatory nitrate-reduction (napA and narG). Since we observed significant differences in nitrate concentrations between Sequences 2 and 3 at high density, this effect was not due to the amount of nutrients released by dead inoculant cells. However, we cannot exclude an indirect effect related to the periodicity of the recurrent release of nutrients by dead inoculant cells. Alternatively, our results also suggest that under recurrent inoculation, P. fluorescens B177 indirectly influenced nitrate concentrations in soil through changes in resident microbial taxa. This is supported by the fact that higher nitrate concentrations were observed in inoculation Sequences 3 and 4, which also had the strongest impact on the relative abundances of resident OTUs. However, we did not observe a significant correlation between any of the genes encoding for nitrate reduction (napA, narG), ammonia oxidation (AOA, AOB) and the nitrate concentrations (Fig. 5). While rare taxa are essential to support some key functions such as methane-production, with their absence leading to a drop in gas production60, in our study, rare taxa were excluded from the differential abundance analysis and were overlooked in the qPCR analysis. Therefore, it is possible that they have played a disproportionate role in the observed nitrate accumulation as this function undergoes fine-tuned regulation.
Conclusion
In this work, we found that despite a high inoculum density of Pseudomonas fluorescens B177, the impact of recurrent inoculation had only a weak impact on the autochthonous microbial community. Inoculation density was the main factor driving the survival of the inoculant as well as the diversity and composition of the soil resident community. Recurrent inoculation led to a transient increase of the abundance of the inoculant compared to single inoculation with only a minor impact on members of the resident community. This shows that an invasion by Pseudomonas fluorescens B177, even at high density, did not induce any important shift in the community, explaining that recurrent inoculation with this strain could not facilitate future invasions by the same strain. Interestingly, recurrent inoculation sequences induced a strong and persistent accumulation of nitrate in soil that could not be attributed to changes in abundances of the targeted nitrogen cycling microbial guilds. Further investigations are needed to understand the mechanisms by which these recurrent inoculation sequences are impacting nitrogen-cycling activity of the resident microbial community.
Methods
Experimental design
Soil was collected from a maize field under conventional agriculture, located in South West of France (64450 Garlède-Mondebat, France). It was characterised as a silty soil (22.2% clay, 66.0% silt and 11.8% sand) with 2.82% organic matter and a pH in water of 5.9. Soil microcosms were prepared by filling 125 mL plasma flasks with 80 g equivalent of dry soil and sieved at 4 mm. The plant growth promoting rhizobacteria Pseudomonas fluorescens strain B177 (produced by GreenCell, France) was used as bacterial inoculant. P. fluorescens B177 was grown at 28 °C in TSB culture medium up to OD of 2. Bacterial cells were then centrifuged at 6000 rpm for 3 min and washed 3 times with a saline solution (0.9% NaCl). The cell pellets were then resuspended to obtain two inoculum densities (106 and 108 cell g of dry soil−1). Four inoculation sequences were applied for each inoculum density, with an increasing number of inoculation events. Sequence 1: one inoculation at day 56 (i.e. week 8), Sequence 2: two inoculations at days 42 and 56 (i.e. weeks 6 and 8), Sequence 3: two inoculations at days 14 and 56 (i.e. weeks 2 and 8), Sequence 4: four inoculations at days 0, 14, 42 and 56 (i.e. weeks 0, 2, 6 and 8). Non-inoculated soils were used as controls. Each treatment (inoculum density * inoculation sequence) was replicated 10 times for a total of 280 microcosms (Supplementary Fig. 1). After each inoculation, the moisture content in soil microcosms was adjusted to 70% of water-holding capacity (WHC). Microcosms were then watered every 2 weeks to maintain a range of 50–70% WHC. The soil microcosms were covered with gauze and incubated at room temperature (ranging from 19 to 21 °C) in the dark during the 14 weeks of experiment.
The soil from ten replicate microcosms per treatment was collected at days 0, 28, 70 and 98 (i.e. weeks 0, 4, 10 and 14) and thoroughly mixed before the following analyses. The sampling at days 70 and 98 corresponds to 2 and 6 weeks after the last inoculation regardless of the inoculation sequence, respectively. Soil sub-samples were stored at – 20 °C until DNA extraction. Soil humidity was also measured on fresh soil sub-samples at each sampling date.
DNA extraction
For each microcosm, about 250 mg equivalent dry soil were weighted and DNA was extracted with the DNeasy 96 PowerSoil Pro kit (Qiagen) according to manufacturer’s instructions. DNA extracts were collected into 100 µL of TE buffer and quantified using the Quant-iT™ PicoGreen® dsDNA Assay Kit (Invitrogen™, France) following the manufacturer's instructions.
Quantification of the abundances of Pseudomonas fluorescens and functional genes
The abundance of Pseudomonas fluorescens B177 abundance was assessed by real-time quantitative PCR (qPCR) using the SCAR primers BP1F (5′-TGCCTGACGCTGTGATACTT-3′) and BP1R (5′-GCAGAAGAGATGAACCCACC-3′). The limit of quantification was assessed by serial dilutions of P. fluorescens genomic DNA according to Waiblinger et al.40. A relative standard error below 30% throughout the replicates was considered as the threshold to determine the limit of quantification which was determined as 80 copies per reaction, corresponding to an average of 4 × 105 copy/g dry soil in our study. Linearised pGEM-T (pGEM-T Easy Vector, Promega) plasmid containing a clone targeted gene from P. fluorescens B177 strain was quantified by QuantiFluor dsDNA system (Promega). A serial dilution ranging from 102 to 108 copies per reaction was used as standards. qPCR reactions were carried out using a ViiA7 (Life Technologies, USA) in 15 µl reactions containing 7.5 µL of Takyon Low ROX SYBR 2X Master Mix (Eurogentec, France), 0.5 µM of each primer, 250 ng of T4 gene 32 (QBiogene, France) and 2 ng per well of DNA template. The PCR cycling conditions included initial denaturation of 95 °C for 3 min followed by 40 cycles of 95 °C for 15 s, 60 °C for 30 s and 72 °C for 30 s. Two runs were performed for each qPCR assay. Melting curves were run to verify the amplification specificity. The results were then processed using QuantStudio software (v1.7.1, Applied Biosystems).
The abundances of total bacterial and total fungal communities and targeted N-cycle microbial guilds (see below) were estimated by real-time quantitative PCR (qPCR) assays. Total bacterial and fungal communities were quantified using 16S rRNA primers as described by Muyzer et al.61. Abundances of N-cycle microbial guilds were estimated using the amoA gene to quantify bacterial (AOB) and archaeal (AOA) ammonia-oxidizers, and the narG and napA genes to quantify nitrate reducers62. DNA extracts diluted at 1ng/µL (described above) were used as template. Linearised pGEM-T plasmids containing the appropriate targeted gene were used as standards. PCR efficiency for the different assays ranged from 78 to 97%. No template controls resulted in null or negligible values. Inhibition in qPCR assays was tested by mixing control plasmid DNA (pGEM-T Easy Vector, Promega, France) with either soil DNA extracts or water. No inhibition was detected with the amount of DNA used.
16S rRNA amplicons sequencing
Bacterial diversity and composition were analysed via Illumina sequencing of the 16S rRNA amplicons generated from all DNA extracts using a two steps PCR procedure as previously described63. The first step consisted in PCR amplification of the V3-V4 hypervariable region of the bacterial 16S rRNA gene using the following primers: U341F (5′-CCTACGGGRSGCAGCAG-3′) and 805R (5′-GACTACCAGGGTATCTAAT-3′). First step PCRs were performed twice for each sample and pooled. In the second step, 6 µL of pooled 16S rRNA amplicons were used as template for a second PCR with Nextera-indexed sequencing primers to add multiplexing index-sequences. SequalPrep Normalisation kit (Invitrogen) was used to normalize amplicon concentrations prior to pooling. Sequencing was performed on Illumina MiSeq (2 × 250 pb paired-end reads) using the MiSeq reagent kit v2.
Sequence processing and bacterial community composition
Sequences obtained from the 240 samples were processed with an in-house developed Python pipeline (https://forgemia.inra.fr/vasa/illuminametabarcoding). Shortly, 16S rRNA amplicon sequences were assembled using PEAR64 with default settings. Further quality checks were conducted using the scripts from the QIIME pipeline65 and short sequences were removed (< 350 bp). Reference-based and de novo chimera detection as well as OTU clustering were performed using VSEARCH66 and SILVA reference database67. The identity threshold was set at 94% based on replicate sequencing of a bacterial mock community. Representative sequences for each OTU were aligned using INFERNAL and phylogenetic tree was constructed using Infernal (version 1.1.3)68. Taxonomy was assigned using BLAST69 and the SILVA reference database 13267. In total, 8,075,341 sequences were obtained and assigned to 7443 OTUs. α-diversity metrics (observed OTUs, Simpson’s reciprocal index and Faith’s Phylogenetic Diversity) were calculated in QIIME based on rarefied OTU tables (> 10,000 sequences per sample). Bray–Curtis, weighted and unweighted UniFrac dissimilarity matrices70 were also computed to detect variations in the structure of microbial communities.
Mineral nitrogen pool
Soil mineral nitrogen (NH4+ and NO3−) was extracted from 7 g of fresh soil sub-samples with 35 mL of 0.5M K2SO4 solution, then shaken (1 h, 80 rpm at room temperature), filtered and kept frozen until quantification. Colorimetric analysis was performed on Tecan infinite M200 (Life Sciences) according to Anderson and Ingram71.
Data analysis
All statistical analyses were performed with R (version 4.0.1). For each sampling date, differences in the microbial α-diversity, gene abundance and mineral nitrogen pools were tested using ANOVA followed by Tukey HSD test (p < 0.05). Normality and homogeneity of the distribution of residuals were checked and log-transformation were applied when necessary. Permutational multivariate analysis of variance (PermANOVA) was carried out on the unweighted UniFrac distance matrices using adonis2 function implemented in the vegan package72. Pairwise comparisons between treatments were implemented using the pairwise.adonis function from the pairwiseAdonis package (version 0.4). Before comparing the differential abundance of OTUs, low abundance OTUs were filtered, keeping only OTUs with an average relative abundance above 1% across all samples. For each treatment, OTUs present in less than 60% of the samples were also excluded. These filtering steps reduce the zero counts in the OTU table which can increase the number of false positives in the differential abundance analysis. Pairwise comparisons were conducted on the resulting 576 OTUs using the following General Linear Mixed Model (GLMM)73. Considering that the abundance Y of each OTU, in any k replicate of any i treatment, follows a Poisson distribution of parameter ∆ as Y ~ P(∆), we used the following equation:
Here, o represents the offset for each sample calculated as the logarithm of the sample read sum, α is the effect of each treatment and Z is the random sampling effect modelling the data overdispersion. Such model combines a generalised linear model and a mixed model. The generalised linear model allows to infer linear regression from data that does not follow a normal distribution because abundance data typically follow a Poisson distribution. The mixed model includes both fixed effects (treatment effects) and random effects (sampling effects). Each sampling time was analysed separately. The treatment effect was encoded as a factor combining the inoculation densities and the sequences, with nine levels. P values were adjusted using the p.adjust function with the false discovery rate (fdr) method to correct for the high number of OTUs tested. Pairwise comparisons were carried out between Ni control and treatments. Comparisons were considered significant when p < 0.05.
Data availability
16S rRNA raw sequences were deposited at the NCBI under the accession number PRJNA1033835. Data that support the findings of this study is also available from the corresponding author upon reasonable request.
References
Sebilo, M., Mayer, B., Nicolardot, B., Pinay, G. & Mariotti, A. Long-term fate of nitrate fertilizer in agricultural soils. Proc. Natl. Acad. Sci. 110, 18185–18189 (2013).
Liess, M. et al. Pesticides are the dominant stressors for vulnerable insects in lowland streams. Water Res. 201, 117262 (2021).
Schulz, R., Bub, S., Petschick, L. L., Stehle, S. & Wolfram, J. Applied pesticide toxicity shifts toward plants and invertebrates, even in GM crops. Science 372, 81–84 (2021).
Schütz, L. et al. Improving crop yield and nutrient use efficiency via biofertilization—a global meta-analysis. Front. Plant Sci. 8, 2204 (2018).
Porter, S. S. et al. Beneficial microbes ameliorate abiotic and biotic sources of stress on plants. Funct. Ecol. 34, 2075–2086 (2020).
O’Brien, P. A. Biological control of plant diseases. Austral. Plant Pathol. 46, 293–304 (2017).
Liu, X., LeRoux, X. & Salles, J. F. The legacy of microbial inoculants in agroecosystems and potential for tackling climate change challenges. iScience 25, 103821 (2022).
Santos, M. S., Nogueira, M. A. & Hungria, M. Microbial inoculants: Reviewing the past, discussing the present and previewing an outstanding future for the use of beneficial bacteria in agriculture. AMB Express 9, 205 (2019).
Waltz, E. A new crop of microbe startups raises big bucks, takes on the establishment. Nat. Biotechnol. 35, 1120–1122 (2017).
Kaminsky, L. M., Trexler, R. V., Malik, R. J., Hockett, K. L. & Bell, T. H. The inherent conflicts in developing soil microbial inoculants. Trends Biotechnol. https://doi.org/10.1016/j.tibtech.2018.11.011 (2019).
Parnell, J. J. et al. From the lab to the farm: An industrial perspective of plant beneficial microorganisms. Front. Plant Sci. 7, 25 (2016).
Fischer, S. E., Jofré, E. C., Cordero, P. V., GutiérrezMañero, F. J. & Mori, G. B. Survival of native Pseudomonas in soil and wheat rhizosphere and antagonist activity against plant pathogenic fungi. Antonie van Leeuwenhoek 97, 241–251 (2010).
Sharma, R. et al. Survival, efficacy and rhizospheric effects of bacterial inoculants on Cajanus cajan. Agric. Ecosyst. Environ. 240, 244–252 (2017).
Mallon, C. A., van Elsas, J. D. & Salles, J. F. Microbial invasions: The process, patterns, and mechanisms. Trends Microbiol. 23, 719–729 (2015).
Schreiter, S. et al. Soil type-dependent effects of a potential biocontrol inoculant on indigenous bacterial communities in the rhizosphere of field-grown lettuce. FEMS Microbiol. Ecol. 90, 718–730 (2014).
Darwin, C. & Kebler, L. On the Origin of Species by Means of Natural Selection, or The Preservation of Favoured Races in the Struggle for Life (J Murray, 1859).
van Elsas, J. D. et al. Microbial diversity determines the invasion of soil by a bacterial pathogen. Proc. Natl. Acad. Sci. 109, 1159–1164 (2012).
De Roy, K. et al. Environmental conditions and community evenness determine the outcome of biological invasion. Nat. Commun. 4, 1383 (2013).
Eisenhauer, N., Schulz, W., Scheu, S. & Jousset, A. Niche dimensionality links biodiversity and invasibility of microbial communities. Funct. Ecol. 27, 282–288 (2013).
Vivant, A.-L., Garmyn, D., Maron, P.-A., Nowak, V. & Piveteau, P. Microbial diversity and structure are drivers of the biological barrier effect against listeria monocytogenes in soil. PLoS One 8, e76991 (2013).
Spor, A. et al. Habitat disturbances modulate the barrier effect of resident soil microbiota on Listeria monocytogenes invasion success. Front. Microbiol. 11, 25 (2020).
Gravuer, K. & Scow, K. M. Invader-resident relatedness and soil management history shape patterns of invasion of compost microbial populations into agricultural soils. Appl. Soil Ecol. 158, 103795 (2021).
Mawarda, P. C., Le Roux, X., Dirk van Elsas, J. & Salles, J. F. Deliberate introduction of invisible invaders: A critical appraisal of the impact of microbial inoculants on soil microbial communities. Soil Biol. Biochem. 148, 107874 (2020).
Vuolo, F., Novello, G., Bona, E., Gorrasi, S. & Gamalero, E. Impact of plant-beneficial bacterial inocula on the resident bacteriome: Current knowledge and future perspectives. Microorganisms 10, 2462 (2022).
Di Salvo, L. P., Cellucci, G. C., Carlino, M. E. & García de Salamone, I. E. Plant growth-promoting rhizobacteria inoculation and nitrogen fertilization increase maize (Zea mays L.) grain yield and modified rhizosphere microbial communities. Appl. Soil Ecol. 126, 113–120 (2018).
Ke, X. et al. Effect of inoculation with nitrogen-fixing bacterium Pseudomonas stutzeri A1501 on maize plant growth and the microbiome indigenous to the rhizosphere. System. Appl. Microbiol. 42, 248–260 (2019).
Renoud, S. et al. Effect of inoculation level on the impact of the PGPR Azospirillum lipoferum CRT1 on selected microbial functional groups in the rhizosphere of field maize. Microorganisms 10, 325 (2022).
Hart, M. M., Antunes, P. M., Chaudhary, V. B. & Abbott, L. K. Fungal inoculants in the field: Is the reward greater than the risk?. Funct. Ecol. 32, 126–135 (2018).
Jack, C. N., Petipas, R. H., Cheeke, T. E., Rowland, J. L. & Friesen, M. L. Microbial inoculants: Silver bullet or microbial Jurassic park?. Trends Microbiol. 29, 299–308 (2021).
Mallon, C. A. et al. The impact of failure: Unsuccessful bacterial invasions steer the soil microbial community away from the invader’s niche. ISME J. 12, 728–741 (2018).
Bedolla-Rivera, H. I., Negrete-Rodríguez, M. L. X., Gámez-Vázquez, F. P., Álvarez-Bernal, D. & Conde-Barajas, E. Analyzing the impact of intensive agriculture on soil quality: A systematic review and global meta-analysis of quality indexes. Agronomy 13, 2166 (2023).
Raaijmakers, J. M. & Weller, D. M. Exploiting genotypic diversity of 2,4-diacetylphloroglucinol-producing Pseudomonas spp.: Characterization of superior root-colonizing P. fluorescens strain Q8r1–96. Appl. Environ. Microbiol. 67, 2545–2554 (2001).
Howie, W. J. Effects of soil matric potential and cell motility on wheat root colonization by fluorescent pseudomonads suppressive to take-all. Phytopathology 77, 286 (1987).
Buddrus-Schiemann, K., Schmid, M., Schreiner, K., Welzl, G. & Hartmann, A. Root colonization by Pseudomonas sp. DSMZ 13134 and impact on the indigenous rhizosphere bacterial community of barley. Microb. Ecol. 60, 381–393 (2010).
Elsas, J. D., Dijkstra, A. F., Govaert, J. M. & Veen, J. A. Survival of Pseudomonas fluorescens and Bacillus subtilis introduced into two soils of different texture in field microplots. FEMS Microbiol. Lett. 38, 151–160 (1986).
Mallon, C. A. et al. Resource pulses can alleviate the biodiversity–invasion relationship in soil microbial communities. Ecology 96, 915–926 (2015).
Araujo, M. A. V., Mendonça-Hagler, L. C., Hagler, A. N. & van Elsas, J. D. Survival of genetically modified Pseudomonas fluorescens introduced into subtropical soil microcosms. FEMS Microbiol. Ecol. 13, 205–216 (1994).
Mawarda, P. C., Mallon, C. A., Le Roux, X., van Elsas, J. D. & Salles, J. F. Interactions between Bacterial inoculants and native soil bacterial community: The case of spore-forming Bacillus spp.. FEMS Microbiol. Ecol. 98, 127 (2022).
Taylor, S. C. et al. The ultimate qPCR experiment: Producing publication quality, reproducible data the first time. Trends Biotechnol. 37, 761–774 (2019).
Waiblinger, H.-U., Graf, N., Broll, H., Grohmann, L. & Pietsch, K. Evaluation of real-time PCR results at the limit of detection. J. Verbr. Lebensm. 6, 411–417 (2011).
Forootan, A. et al. Methods to determine limit of detection and limit of quantification in quantitative real-time PCR (qPCR). Biomol. Detect. Quantif. 12, 1–6 (2017).
Nutz, S., Döll, K. & Karlovsky, P. Determination of the LOQ in real-time PCR by receiver operating characteristic curve analysis: Application to qPCR assays for Fusarium verticillioides and F. proliferatum. Anal. Bioanal. Chem. 401, 717–726 (2011).
Berninger, T., González López, Ó., Bejarano, A., Preininger, C. & Sessitsch, A. Maintenance and assessment of cell viability in formulation of non-sporulating bacterial inoculants. Microbial Biotechnol. 11, 277–301 (2018).
Nerek, E., Sokołowska, B., Nerek, E. & Sokołowska, B. Pseudomonas spp. in biological plant protection and growth promotion. AIMSES 9, 493–504 (2022).
Ketola, T., Saarinen, K. & Lindström, L. Propagule pressure increase and phylogenetic diversity decrease community’s susceptibility to invasion. BMC Ecol. 17, 15 (2017).
Jones, M. L., Ramoneda, J., Rivett, D. W. & Bell, T. Biotic resistance shapes the influence of propagule pressure on invasion success in bacterial communities. Ecology 98, 1743–1749 (2017).
Wittmann, M. J., Metzler, D., Gabriel, W. & Jeschke, J. M. Decomposing propagule pressure: The effects of propagule size and propagule frequency on invasion success. Oikos 123, 441–450 (2014).
Amor, D. R., Ratzke, C. & Gore, J. Transient invaders can induce shifts between alternative stable states of microbial communities. Sci. Adv. 6, eaay8676 (2020).
Jiménez, J. A., Novinscak, A. & Filion, M. Inoculation with the plant-growth-promoting rhizobacterium Pseudomonas fluorescens LBUM677 impacts the rhizosphere microbiome of three oilseed crops. Front. Microbiol. 11, 25 (2020).
Albright, M. B. N., Sevanto, S., Gallegos-Graves, L. V. & Dunbar, J. Biotic interactions are more important than propagule pressure in microbial community invasions. mBio 11, 02089–20 (2020).
Cornell, C. et al. Do bioinoculants affect resident microbial communities? A meta-analysis. Front. Agron. 3, 25 (2021).
Wang, Z. et al. Succession of the resident soil microbial community in response to periodic inoculations. Appl. Environ. Microbiol. 87, e00046-e121 (2021).
Cipriano, M. A. P. et al. Lettuce and rhizosphere microbiome responses to growth promoting Pseudomonas species under field conditions. FEMS Microbiol. Ecol. 92, fiw197 (2016).
Roquigny, R. et al. Deciphering the rhizosphere and geocaulosphere microbiomes of potato following inoculation with the biocontrol agent Pseudomonas fluorescens strain LBUM223. Phytobiomes J. 2, 92–99 (2018).
Cahill, J. F., Kembel, S. W., Lamb, E. G. & Keddy, P. A. Does phylogenetic relatedness influence the strength of competition among vascular plants?. Perspect. Plant Ecol. Evol. Syst. 10, 41–50 (2008).
Traveset, A. & Richardson, D. M. Plant invasions: The role of biotic interactions—an overview. Plant Invas. Role Biotic Interact. 20, 1–25. https://doi.org/10.1079/9781789242171.0001 (2020).
Mar Vázquez, M., César, S., Azcón, R. & Barea, J. M. Interactions between arbuscular mycorrhizal fungi and other microbial inoculants (Azospirillum, Pseudomonas, Trichoderma) and their effects on microbial population and enzyme activities in the rhizosphere of maize plants. Appl. Soil Ecol. 15, 261–272 (2000).
Nassal, D. et al. Effects of phosphorus-mobilizing bacteria on tomato growth and soil microbial activity. Plant Soil 427, 17–37 (2018).
Wu, F. et al. Acinetobacter calcoaceticus CSY-P13 mitigates stress of ferulic and p-hydroxybenzoic acids in cucumber by affecting antioxidant enzyme activity and soil bacterial community. Front. Microbiol. 9, 25 (2018).
Sierocinski, P., Soria Pascual, J., Padfield, D., Salter, M. & Buckling, A. The impact of propagule pressure on whole community invasions in biomethane-producing communities. iScience 24, 102659 (2021).
Muyzer, G., de Waal, E. C. & Uitterlinden, A. G. Profiling of complex microbial populations by denaturing gradient gel electrophoresis analysis of polymerase chain reaction-amplified genes coding for 16S rRNA. Appl. Environ. Microbiol. 59, 695–700 (1993).
Bru, D. et al. Determinants of the distribution of nitrogen-cycling microbial communities at the landscape scale. ISME J. 5, 532–542 (2011).
Berry, D., Ben Mahfoudh, K., Wagner, M. & Loy, A. Barcoded primers used in multiplex amplicon pyrosequencing bias amplification. Appl. Environ. Microbiol. 77, 7846–7849 (2011).
Zhang, J., Kobert, K., Flouri, T. & Stamatakis, A. PEAR: A fast and accurate Illumina Paired-End reAd mergeR. Bioinformatics 30, 614–620 (2014).
Caporaso, J. G. et al. QIIME allows analysis of high-throughput community sequencing data. Nat. Methods 7, 335–336 (2010).
Rognes, T., Flouri, T., Nichols, B., Quince, C. & Mahé, F. VSEARCH: A versatile open source tool for metagenomics. PeerJ 4, e2584 (2016).
Quast, C. et al. The SILVA ribosomal RNA gene database project: Improved data processing and web-based tools. Nucleic Acids Res. 41, D590–D596 (2013).
Nawrocki, E. P. & Eddy, S. R. Infernal 1.1: 100-fold faster RNA homology searches. Bioinformatics 29, 2933–2935 (2013).
Altschul, S. F., Gish, W., Miller, W., Myers, E. W. & Lipman, D. J. Basic local alignment search tool. J. Mol. Biol. 215, 403–410 (1990).
Lozupone, C., Lladser, M. E., Knights, D., Stombaugh, J. & Knight, R. UniFrac: An effective distance metric for microbial community comparison. ISME J. 5, 169–172 (2011).
Anderson, J. M. & Ingram, J. Tropical Soil Biology and Fertility: A Hand Book of Methods, XF2006286592 (1989).
Dixon, P. VEGAN, a package of R functions for community ecology. J. Veg. Sci. 14, 927–930 (2003).
Huet, S. et al. Experimental community coalescence sheds light on microbial interactions in soil and restores impaired functions. Microbiome 11, 42 (2023).
Acknowledgements
The authors thank Ahmed Taibi, researcher at GreenCell, for its inputs during the reviewing process.
Funding
This work was funded by the Agence Nationale de la Recherche – project number: ANR-20-CE02-0014.
Author information
Authors and Affiliations
Contributions
M.P., X.L.R., L.P. and A.S. designed the experiment, A.D.Z. provided the inoculant, M.P., M.C.B., N.R. conducted the experiment and sampling, M.P., D.B. and M.C.B. analysed the samples, M.P., A.S. and A.M. conducted the bioinformatic and statistical analysis, M.P. wrote the manuscript with the help of L.P., A.S. and X.L. All authors approved the submitted version.
Corresponding author
Ethics declarations
Competing interests
A.Z.D is R&D manager at the GreenCell company, which markets the Pseudomonas fluorescens B177 strain protected under BUDAPEST treaty at Pasteur Institute (CNCM I-4015), as a component of the ‘CERES’ product. This experiment, the analyses and the manuscript strictly follow the ethical standards.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Papin, M., Philippot, L., Breuil, M.C. et al. Survival of a microbial inoculant in soil after recurrent inoculations. Sci Rep 14, 4177 (2024). https://doi.org/10.1038/s41598-024-54069-x
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-024-54069-x
Keywords
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.