Abstract
The intestinal microbiome plays an important role in mammalian health, disease, and immune function. In light of this function, recent studies have aimed to characterize the microbiomes of various bat species, which are noteworthy for their roles as reservoir hosts for several viruses known to be highly pathogenic in other mammals. Despite ongoing bat microbiome research, its role in immune function and disease, especially the effects of changes in the microbiome on host health, remains nebulous. Here, we describe a novel methodology to investigate the intestinal microbiome of captive Jamaican fruit bats (Artibeus jamaicensis). We observed a high degree of individual variation in addition to sex- and cohort-linked differences. The intestinal microbiome was correlated with intestinal metabolite composition, possibly contributing to differences in immune status. This work provides a basis for future infection and field studies to examine in detail the role of the intestinal microbiome in antiviral immunity.
Similar content being viewed by others
Introduction
Intestinal metabolism and microbiome composition are increasingly implicated in human health outcomes, especially disease course and severity of viral infections1,2,3,4,5,6. A growing body of literature indicates that the intestinal microbiome plays a functional role in many mammals, in part via the synthesis of metabolites that may influence intestinal immunity through responses to infection6,7,8,9,10. Among the most well-established of these associations is the production of various short chain fatty acids (SCFAs), which perform immunoregulatory functions, by bacterial species commonly found in human intestinal microbiomes11,12,13.
Bats are the reservoir hosts of viruses in several families, including Filoviridae, Paramyxoviridae, and Coronaviridae, that can cause severe disease in humans14,15,16,17. There are data indicating that bats, as the only flight-adapted mammals, have developed methods for reducing inflammation incurred during flight. These include reducing inflammatory cytokines such as TNF18, loss of the PYHIN gene family19, reduced activation of the NLRP3 inflammasome20, downregulation of caspase-1-mediated inflammasome activation21, dampened interferon activation as a result of STING mutation22, and increased expression of inflammasome-suppressing ASC223. There is also evidence supporting the constitutive expression of innate immune genes, such as interferon-stimulated genes24,25, to reduce flight-induced oxidative metabolism and DNA damage22,26 in bats.
Flight has also been implicated in the unique signatures of bat intestinal microbiomes27,28,29. Furthermore, the intestinal microbiomes of several bat species are characterized by high abundances of Pseudomonadota and low abundances of Bacteroidetes, a suggested hallmark of dysbiosis in humans30,31. However, transplants of bat intestinal microbiomes to mice infected with H1N1 influenza virus lead to decreased inflammatory responses and increased survival rates32. Thus, microbial signatures that constitute dysregulation in other mammalian species may in fact contribute to bat immune responses to viral infections, potentially contributing to their posited immune tolerance33,34.
The Jamaican fruit bat (Artibeus jamaicensis) harbors diverse coronaviruses in the wild35 and has previously been used as a bat reservoir host model for MERS-CoV infection36. They are susceptible to a broad range of viruses after experimental inoculation, demonstrating Tacaribe virus and rabies virus infections of varying severity depending on inoculation dose and virus strain, respectively, asymptomatic Zika virus infection, and clinically mild bat H18N11 influenza A virus infection37,38,39,40. Despite their use as a reservoir host model and susceptibility to several highly pathogenic viruses, Jamaican fruit bats remain understudied with regards to their intestinal microbiome and immune response to infection, partially due to the difficulty of controlled studies under high containment conditions.
Here, we established novel methods for intestinal microbiome analysis of a captive colony of Jamaican fruit bats under conditions mimicking those of a prototypical infection study in high containment and characterized sex- and cohort-specific differences in the microbiome. We then conducted a metabolomic investigation in relation to differences observed in the intestinal microbiome. This work will provide a foundation for future infection studies to elucidate immune responses and mechanisms of immune tolerance of a model reservoir host species for newly emerging and re-emerging viruses.
Results
Microbiome analysis methods establishment for samples collected in maximum containment
We established novel methods to conduct microbiome analyses of samples obtained from animals under maximum containment (BSL-4) conditions, which can be applied to bats infected with BSL-3 and BSL-4 pathogens. Given the restrictions on sample removal and inactivation from high containment, the methods developed here incorporated acceptable inactivation techniques. We first established protocols for inactivating and extracting bacterial DNA from bat rectal swabs by testing three commercially available methods (Supplementary Fig. 1a,b): TRIzol Reagent (ThermoFisher), the AllPrep DNA/RNA Kit (Qiagen), and the QIAamp Fast DNA Stool Mini Kit (Qiagen). We determined that extraction with TRIzol reagent produced the highest yield of DNA.
Jamaican fruit bat fecal samples and rectal swabs were then compared after TRIzol extraction to establish downstream sampling methodology (Supplementary Fig. 1c). We tested both sample types with a PCR protocol for 16S ribosomal DNA based on manufacturer’s recommendations (Illumina) and modified accordingly41. In this study of Jamaican fruit bats—which produce soft, semi-formed fecal samples—rectal swabs both produced cleaner PCR products than did fecal samples and allowed for accurate longitudinal sampling of individuals.
Microbial community composition of captive Jamaican fruit bats
We first analyzed the captive Jamaican fruit bat intestinal microbiome after acclimation of two cohorts shipped to our facility in October 2021 and May 2022 (N = 20 and N = 30, respectively), characterizing both the bacterial families present and their relative abundances in both cohorts before the onset of longitudinal observation. Following exclusion of unclassified reads and reads classified as chloroplasts, mitochondria, or non-prokaryotes, we found that a small number of families dominated the intestinal microbiome. Of 55 classifiable families, 14 families comprised 97.28% of the intestinal microbiome of October cohort bats (Fig. 1a). Similarly, the baseline microbial community composition of the May cohort bats was also dominated by a small number of bacterial families (Fig. 1a): only 11 families had relative abundances greater than 0.5% across the 30 May cohort bats, with 79 total families categorized as “other”. Dominant families across both cohorts included Moraxellaceae, Enterobacteriaceae, Peptostreptococcaceae, Clostridiaceae, Leuconostocaceae, Mycoplasmataceae, Neisseriaceae, Streptococcaceae, Pseudomonadaceae, and Pasteurellaceae. We also observed individual variation in microbial community composition, regardless of sex and cohort.
Differences between two adult cohorts (October, N = 20 and May, N = 30) of captive Jamaican fruit bats (Artibeus jamaicensis) in intestinal microbial community composition and diversity immediately after relocation between facilities. Samples taken after acclimation. (a) Percent abundance of selected microbial families in the intestinal tract across all bats (pie chart), by cohort (top right), and by individual bat (bottom). Unclassified reads at the family level and reads classified as mitochondria, chloroplasts, or non-prokaryotes were removed. Families with less than 0.5% abundance in either cohort were categorized as “Other” for visualization purposes. (b) Alpha diversity metrics (left to right: Shannon entropy, Simpson’s diversity, Faith’s phylogenetic diversity, and observed features) of the intestinal microbiome by cohort. Median and 95% confidence intervals with individual points overlaid. Significant p-values indicated; Mann–Whitney test (N = 20/30). (c) Principal coordinate analysis of unweighted UniFrac distance showing microbial community composition differences (left) and log10(linear discriminant analysis score) of differentially abundant families (LEfSE, p < 0.05) between cohorts (right).
Microbial community comparison of October and May cohorts
We then compared both the microbial community composition and diversity of the two cohorts, aggregating individuals to the cohort level, with the goal of determining whether they should be analyzed separately. In comparing alpha diversity metrics (Supplementary Table S1) between the two groups, the May cohort had slightly, but not significantly, higher median Shannon entropy and Simpson’s diversity scores than did the October cohort (Fig. 1b). The May cohort samples also had a median Faith’s phylogenetic diversity score of 4.98, as compared to the October cohort median of 3.98 (p = 0.0012, Mann–Whitney test). Taken together, these results indicate greater phylogenetic diversity, but not necessarily richness or evenness, in the intestinal microbiome composition of the May cohort bats as compared to the October cohort bats. Interestingly, this was despite the October cohort bats having a median number of observed features of 764, whereas the May cohort bats only had a median of 317.5 observed features (p = 0.0007, Mann–Whitney test). Thus, though the October cohort samples potentially had a larger total number of species than did the May cohort samples, the May cohort samples likely had greater phylogenetic diversity in those species that were present.
Several bacterial families that were abundant (i.e., percent composition greater than 0.5% of the microbiome) in the October cohort were not abundant in the May cohort. These families included Sphingomonadaceae, Burkholderiaceae, Chitinophagaceae, and Spirochaetaceae. We observed a new family, Carnobacteriaceae, in the May cohort samples. Among the families that remained abundant in both cohorts, there were differences in relative abundance between the two cohorts. Moraxellaceae, Enterobacteriaceae, and Peptostreptococcaceae were relatively more abundant in the October cohort, whereas Clostridiaceae, Mycoplasmataceae, and Streptococcaceae were more enriched in the May cohort.
We found significant (p = 0.001, PERMANOVA) community composition differences (Supplementary Table S1) in the intestinal microbiome between the two cohorts when using unweighted UniFrac (Fig. 1c, left) distance as well as Bray–Curtis (p = 0.001, PERMANOVA) and weighted UniFrac (p = 0.001, PERMANOVA) distance (Supplementary Fig. 2a,b). We conducted a differential abundance analysis and identified 21 bacterial families that displayed significantly (p < 0.05, LEfSe) different abundance between the cohorts (Fig. 1c, right). Concordant with our findings that the October cohort samples had higher median numbers of species, the October cohort was differentially enriched in 17 of the 21 families while the May cohort samples were relatively more abundant in only four. As expected, the Carnobacteriaceae family was differentially abundant in the May cohort. The other differentially abundant families also aligned with our initial observations of visually apparent abundance differences in microbial community composition in the two cohorts.
Sex and pregnancy status differences in microbial community composition and diversity
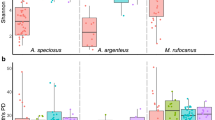
Given the substantial and significant differences between the two cohorts in both microbial community composition and diversity, we chose to analyze the October and May cohorts independently rather than combining them. Considering documented mammalian sex and pregnancy status differences in immunity42,43,44, we analyzed the intestinal microbiota of each cohort by sex and pregnancy status. The intestinal microbiomes of female bats in the October cohort had marginally, but non-significantly, higher median Shannon entropy (female = 6.17; male = 5.97) and Faith’s phylogenetic diversity (female = 4.23; male = 3.52) values than did those of males (Fig. 2a). Female intestinal microbiomes had a median of 1578.5 observed features while male intestinal microbiomes had a median of 397.5 observed features (p = 0.0056, Welch’s t-test); however, female intestinal microbiomes displayed slightly—but not significantly—lower median Simpson’s diversity values than did males’ (female = 0.963; male = 0.975). Thus, female bats in the October cohort had larger average numbers of species in their intestinal microbiomes than male bats, but may have had lower evenness or phylogenetic relatedness, indicating potential dominance of a small number of families.
Intestinal microbial community composition and diversity of October and May cohort bats by sex (male, N = 10 October; N = 9 May and female, N = 10 October; N = 21 May) and pregnancy status (pregnant, N = 2 October; N = 10 May and not pregnant, N = 8 October; N = 11 May). (a) Alpha diversity metrics (left to right: Shannon entropy, Simpson’s diversity, Faith’s phylogenetic diversity, and observed features) of the October (top) and May (bottom) cohort intestinal microbiome by sex. Median and 95% confidence intervals with individual points overlaid. Significant p-values indicated; Mann–Whitney test (N = 10, October; N = 9 (males) and 21 (females), May). (b) Principal coordinate analysis showing unweighted UniFrac distance by sex and pregnancy status in the October cohort (left), and Bray–Curtis distance by sex and pregnancy status in the May cohort (right). Ellipses denote significant (kmeans, p < 0.05) clusters. Points colored by sex and pregnancy status. (c) log10(linear discriminant analysis score) of differentially abundant families (LEfSE, p < 0.05) between male and female bats at intake in the October (left) and May (right) cohorts.
We observed significant sex (p = 0.001, PERMANOVA) and pregnancy status (p = 0.001, PERMANOVA) differences in the microbiome comparing male and female bats in the October cohort when we calculated unweighted UniFrac distance (Fig. 2b). Sex-related community composition differences were also observed via weighted UniFrac distance (p = 0.002, PERMANOVA), but not Bray–Curtis distance (Supplementary Fig. 3a,b). We identified six families that significantly contributed (p < 0.05, LEfSe) to the observed sex differences in community composition in the October cohort (Fig. 2c): Muribaculaceae, Lactobacillaceae, Leptotrichiaceae, Obscuribacteraceae, Sphingomonadaceae, and Pseudomonadaceae. Of these families, four (Muribaculaceae, Lactobacillaceae, Leptotrichiaceae, and Obscuribacteraceae) contributed to less than 0.5% each of the overall microbial community composition in the October cohort. All four of the rare families were differentially abundant in female bats, and both relatively abundant families were more abundant in male bats, further supporting our observation that October cohort female bats likely had a larger number of species overall but lower evenness amongst species than did male bats.
Unlike in the October cohort, we observed no sex differences in microbial diversity via any of Shannon entropy, Simpson’s diversity, Faith’s phylogenetic diversity, or observed features among the May cohort bats (Fig. 2a), though one female bat’s intestinal microbiome (Bat 30) displayed anomalously high diversity across all metrics. However, we did observe differences in microbial community composition, measured by Bray–Curtis distance, by both sex (p = 0.004, PERMANOVA) and pregnancy status (Fig. 2b; p = 0.032, PERMANOVA). This sex-associated community composition difference was present when the data were analyzed via unweighted (p = 0.047, PERMANOVA), but not weighted, UniFrac distance (Supplementary Fig. 4a). We identified three families that significantly contributed (p < 0.05, LEfSe) to observed sex differences after acclimation (Fig. 2c): Neisseriaceae, Chitinophagaceae, and Peptostreptococcaceae. Interestingly, there was no overlap between the two cohorts in those bacteria that contributed to sex differences in microbial community composition. Furthermore, the October cohort had more pronounced sex differences overall, as well as a larger total number of families that significantly contributed to sex differences.
Longitudinal analysis of the bat intestinal microbiome
Next, we investigated changes in the intestinal microbiomes of May cohort bats over a 28-day period (Supplementary Table S2), finding that the beginning (day 0 (D0)) and end of the study period (day 28 (D28)) showed the most substantial differences compared to any of the intermediate timepoints (days 1, 4, 7, and 14 (D1, D4, D7, D14)). Thus, we began by comparing samples taken immediately after acclimation (D0; a subset of May cohort bats, N = 5 females and 4 males) and after 28 days (D28, N = 5 females and 4 males) to assess potential longitudinal changes in microbial diversity and community composition under the repeated isoflurane treatment required to obtain the samples. We also analyzed potential differences in longitudinal changes related to sex, pregnancy status, and housing scheme in our facility.
Bats experienced slight, but non-significant by Wilcoxon matched pairs test, longitudinal increases in both the overall number of species present in their intestinal microbiomes as well as evenness and phylogenetic diversity of their microbial communities (Fig. 3a). Shannon entropy increased from a median of 5.83 at the beginning of the study to a median of 6.45 at the end of the study. Likewise, median Faith’s phylogenetic diversity increased from 4.99 to 5.58 and median observed features increased from 251 to 309. Finally, Simpson’s diversity increased slightly from a median of 0.972 at the beginning of the study to a median of 0.975 by the endpoint.
Intestinal microbial community composition and diversity of May cohort bats at study beginning (D0, N = 9) and end (D28, N = 9). (a) Alpha diversity metrics (left to right: Shannon entropy, Simpson’s diversity, Faith’s phylogenetic diversity, and observed features) of the intestinal microbiome. Median and 95% confidence intervals with individual points overlaid. Gray lines indicate points belonging to the same animal. Significant p-values indicated; Wilcoxon matched pairs test of bats euthanized at study endpoint (N = 9). (b) Principal coordinate analysis showing unweighted UniFrac distance between baseline and endpoint samples. Ellipses denote significant (kmeans, p < 0.05) clusters. Points colored by sampling time point. (c) Bar plot showing MaAsLin2 coefficients for families with corrected p-values < 0.05. Coefficients are equivalent to log2(fold change) over time; negative coefficients indicate decline in relative abundance and positive coefficients indicate increase in relative abundance. p-values corrected using Benjamini–Hochberg procedure shown. (d) Principal coordinate analysis showing Bray–Curtis distance by sex and pregnancy status at endpoint in the May cohort. Ellipses denote significant (kmeans, p < 0.05) clusters. Points colored by sex and pregnancy status.
Microbial community composition, as measured by unweighted UniFrac distance, changed significantly (p = 0.016, PERMANOVA) (Fig. 3b). Neither the weighted UniFrac distance (Supplementary Fig. 5a) nor the Bray–Curtis distance (Supplementary Fig. 5b) captured this trend. To further investigate longitudinal changes in gut microbial composition, we tested the association between day and the relative abundance of families using MaAsLin2. Briefly, the log-transformed relative abundance of all families present in > 10% of the total number of samples were correlated with experimental day using a general linear model adjusting for sex and sample ID. Among 92 families tested, we identified four families that changed significantly by day (corrected p-values < 0.05, Fig. 3c), indicating longitudinal change over the duration of the study. Of these four families, Moraxellaceae and Leuconostocaceae decreased in relative abundance over time, whereas Yersiniaceae and Rhizobiaceae increased in relative abundance over time (Fig. 3c, Supplementary Fig. 7). Overall, the results suggest that time after acclimation influenced bat gut microbial composition to some extent.
We observed no microbiome convergence by cohousing scheme (Supplementary Fig. 6). Upon further inspection, we found that a small number of animals—though not the same animal each time—drove significant changes in relative abundance of different families (Supplementary Fig. 7). This appeared to be especially true for the Sphingobacteriaceae and Bacillaceae families, indicating the potential for a single bat to drive significant results in an entire group.
Observed sex differences were maintained through D28, with significant sex differences in microbial community composition observed via Bray–Curtis distance at D28 (Fig. 3d; p = 0.042, PERMANOVA). This was not observed with weighted and unweighted UniFrac distances (Supplementary Fig. 4b). No pregnancy status differences were observed at D28, likely due to both the small number of bats and the disproportionate number of pregnant females by the end of the study. Interestingly, though we observed statistically significant sex differences by Bray–Curtis distance, we were able to find no individual families that significantly contributed to these sex differences, indicating that these differences were likely due to several small changes in community composition.
Intestinal metabolite analysis
Considering the persistent sex differences observed in microbial community composition and the microbiome’s links to intestinal metabolism, we investigated intestinal tract metabolite composition via ileal tissue sampling at the time of euthanasia (D28) for those bats we followed longitudinally (Supplementary Table S1). We were particularly interested in potential sex differences in metabolite composition, as well as the metabolome’s relation to the microbiome.
In concordance with our microbiome data, we observed intestinal metabolite composition differences by sex (Fig. 4a), with male and female bats showing separation along dimension 1 of our sPLSDA analysis. Several metabolites contributed to this difference; asparagine (p = 0.1905, Mann–Whitney test) and arginine (p = 0.1111) were non-significantly enriched and histidine (p = 0.0159) was significantly enriched in males, while the nucleobases adenine (p = 0.0317), adenosine (p = 0.0317), and guanosine (p = 0.0317) were all significantly enriched in females (Fig. 4b, Supplementary Fig. 8).
Metabolomics analysis of May cohort bats (N = 9). Metabolites were detected by mass spectrometry of intestinal samples collected during necropsy. (a) sPLSDA showing bat intestinal metabolomes by sex (female, N = 5 and male, N = 4) based on differences in relative metabolite abundance. Points colored by sex. (b) Loadings of sPLSDA analysis of metabolite differences according to sex. Metabolites greater than or equal to 25% of the maximum absolute loading value are colored according to the direction in which they contribute to intergroup differences and labeled with metabolite name. Metabolites less than 25% of the maximum absolute loading value are shown in gray and unlabeled. (c) Correlation plot showing R values and p-values (shown for significance 0.1 and under) for Pearson correlation between metabolites (columns) and bacterial genera (rows). Positive R values in orange; negative R values in purple. Large p-values in white; small p-values in black. Metabolites organized and labelled by family. Asterisk (*) denotes an abbreviation for the Burkholderia–Caballeronia–Paraburkholderia family.
To identify primary bacterial drivers of the metabolic state, we calculated Pearson correlation coefficients of intestinal metabolite profiles with microbiome genus abundance (Fig. 4c). A cluster of bacterial genera, including Actinomyces, Burkholderia–Caballeronia–Paraburkholderia, Corynebacterium, Pasteurellaceae, Streptococcus, and Mycoplasma correlated with a shared metabolic pattern that was consistent across multiple metabolite families. In general, this group of bacteria was positively correlated with the levels of SCFAs, certain nucleobases and carboxylic acids, pyruvate and lactate, and specific amino acids, such as tryptophan, cystine, and glycine. These bacteria were negatively associated with the levels of flavin and nicotinamide coenzymes, NDP-sugars, and the amino acids aspartate, glutamine, and lysine.
The dominant Actinomyces-associated metabolic pattern was anticorrelated with a cluster of genera including Clostridiaceae, Peptostreptococcaceae, Comamonas, Weissella, Enterobacterales, and Sphingomonadaceae. In contrast to the other cluster of bacteria, these bacteria were correlated with higher levels of glycolytic intermediates, aspartate, and ketone bodies. Clostridiaceae was especially closely associated with increased levels of serine and methionine in the intestine. Finally, independent of any major metabolic patterns observed, Sphingomonadaceae was found to be associated with metabolites related to carbohydrate metabolism, and specifically low levels of metabolites produced as a result of glucosamine metabolism.
Discussion
We investigated community composition and diversity of the intestinal microbiome of two cohorts of captive Jamaican fruit bats, one of which we followed longitudinally over a 28-day period. Through Illumina sequencing and analysis of rectal swabs, we established a novel method for microbiome analysis of samples removed from high and maximum containment. Though many viruses of interest require BSL-4 conditions, such analyses were not previously possible due to limited availability of inactivation methods approved for these conditions45,46,47. The incorporation of acceptable inactivation methods for samples obtained in containment into our methodology will allow for analysis of bat samples during infection with BSL-4 viruses. Furthermore, our use of rectal swabs for longitudinal sampling of each bat facilitates time-course studies during infection and enables the comparison of each bat to its own initial baseline. Together, the ability to longitudinally analyze changes in both the microbiome and metabolome over the course of infection enables detailed studies of their interaction and the potential effects on immunological responses to infection in bats.
Surprisingly given consistency in housing and diet, we observed individual variation in microbial community composition across both cohorts. In humans, the microbiome displays individual variation associated with health-related host phenotypes48. In primates, individual variation in the intestinal microbiome is related to social structures and groups rather than habitat or diet, with adult microbiomes varying more with group affiliation than environment49. Furthermore, work on wild-caught vampire bats (Desmodus rotundus) that were merged into one colony upon capture suggests that social interactions influence the intestinal microbiome even when controlling for other factors50. Wild Jamaican fruit bats form harem groups composed of multiple females and a single male51,52, which may influence their intestinal microbiomes in ways that are difficult to observe.
Importantly, most of these animals harbored the same few dominant bacteria despite variation in their abundances. This aligns with the observation that most mammalian species possess a “core” microbiome, defined as the set of microbial taxa characteristic of that species53. Laboratory mice harbor the same core bacterial families across individuals and even species54, though they may experience less variation due to inbreeding. Future work corroborating these observations in other populations of Jamaican fruit bats is necessary to assess the presence or absence of a species-wide core microbiome.
We observed differences in microbial community composition and diversity between our two cohorts after acclimation, despite previous work suggesting convergence of the microbiome of various wild bat species as a result of captivity55,56. In particular, the May cohort appeared to have a smaller number of total bacterial species but increased phylogenetic diversity between species than did the October cohort. The two cohorts also had phylogenetically distinct microbiomes. Given evidence indicating that functional traits may be conserved in phylogenetically similar bacteria57, the microbiome may be serving different functions in the two cohorts, with the May cohort microbiome potentially having broader functionality. Though reproductive status may drive seasonality of the intestinal microbiome58,59,60, the captive colony from which these cohorts were formed anecdotally experiences no observable reproductive seasonality, ruling out birth pulses as a driver of cohort-level community composition differences.
However, we propose several potential explanations for cohort-level differences in microbial community composition. Despite these cohorts lacking exposure to seasonal changes in their environment, long-standing adaptations to seasonality in fruiting and flowering patterns may have driven seasonality in the microbiome that is retained in captivity, explaining differences between the two cohorts. Diet affects the intestinal microbiome61,62,63; though diets were kept constant between the two cohorts, seasonal differences in sourcing and availability of fruit may also have contributed to our observations. Though there are limited data on the effects of shipment on mammalian intestinal microbiomes, the microbiomes of these two cohorts may have been affected differently by the process of shipment and facility transfer. Though transport conditions were controlled and under strict animal welfare regulations, environmental conditions (temperature, light cycling, etc.) during transit may have varied slightly by season, potentially contributing to differences in the microbiomes of the cohorts upon arrival. Alternatively, data show humanization of the microbiome in captive animals, potentially indicating transfer of intestinal bacteria between animals and their human caretakers64,65,66. Thus, seasonal variation in caretaker microbiomes may have influenced differences between the intestinal microbiomes of our two cohorts. However, all caretakers and investigators were always wearing full personal protective equipment, reducing the likelihood of microbiome transfer. More likely, seasonal variation in the sourcing and availability of fruit, these bats’ primary food source, contributed to differences between the October and May cohorts. Future studies should consider methods such as irradiation of provided food or specific decontamination procedures for care staff to mitigate these risks. Regardless, our observations of individual and cohort variation in community composition highlight the importance of establishing baseline values for both individuals and cohorts in future studies.
Despite these cohort-level differences, several dominant bacterial families in both cohorts, including Enterobacteriaceae, Moraxellaceae, and Streptococcaceae, comprise the microbiota of wild rousette and pteropid bats67,68,69,70, potentially signaling a pattern of dominance of a few families across multiple bat species. Several of these families, including Enterobacteriaceae and Moraxellaceae, belong to the Pseudomonadota phylum, the abundance of which may indicate microbial dysbiosis in humans31,71. However, this microbial signature is thought to contribute to decreased inflammatory responses observed in human centenarians72,73,74,75, providing a potential contributor to bats’ posited immune tolerance.
We demonstrated significant longitudinal changes to microbial diversity and community composition over a 28-day period with multiple anesthetic events. As in the cohort-level differences, the difference in unweighted UniFrac distance at D0 and D28 implies phylogenetic, and thus potentially functional, changes to the intestinal microbiome over this period. This exposure to a new environment likely introduced new microbes to the bats’ intestinal tracts, increasing diversity; similar increases in diversity have been observed in wild bats introduced to a new environment in captivity55. These changes demonstrate the importance of including a longitudinal microbiome control in future studies.
We observed that a small group of bacteria, including Actinomyces and Streptococcus, was correlated with increased levels of SCFAs. While not typically considered primary producers of SCFAs, Actinomyces and Streptococcus are positively associated with systemic SCFA levels in human obesity models76, and Streptococcus can produce acetate, a SCFA, via the Wood-Ljungdahl pathway77. The presence of this association in bats may indicate either that these species are producers of SCFAs, interaction with other species encourage them to produce SCFAs, or that the colonization of these species is encouraged by SCFAs. Various SCFAs have been repeatedly demonstrated to potentially be involved in immunoregulation and gut-brain communication in mouse models78,79,80, and butyrate has been implicated as enhancing antibody responses of murine splenocytes81, indicating the possibility of a similar function in bats.
We also found associations between bacteria and tryptophan levels in our data. Streptococcus, which was associated with increased tryptophan, can produce serotonin from tryptophan82. Clostridiaceae, which was associated with decreased tryptophan, can decarboxylate tryptophan83. These data indicate a potential role of these bacteria in regulating tryptophan levels. In humans, tryptophan is associated with intestinal immunity, with alterations being linked to disease and the intestinal microbiome84. Thus, concordant with previous literature describing humans and mice85,86,87, the intestinal microbiome of Jamaican fruit bats may drive metabolome changes that affect intestinal immunity and immunoregulation.
We observed sex differences in microbial community composition measured by unweighted UniFrac, but not Bray–Curtis distance, in both cohorts, indicating that male and female bat microbiomes may have substantial overlap in species present, but that non-overlapping species are phylogenetically distant. This result highlights the need for more work investigating the roles of microbes not shared by male and female bats in order to further elucidate sex differences.
Regardless, these sex differences align with observed differences in microbial diversity and community composition based on reproductive stage and pregnancy status in lesser long-nosed bats (Leptonycteris yerbabuenae)60. Prior work suggests that when sex differences in the intestinal microbiome are present, female bats have more diverse microbiomes than males59,60,88, though our observations on this relationship were equivocal. We also observed the maintenance of the initially observed sex differences over the study period. However, the factors contributing to these sex differences changed between the study’s onset and its end, indicating that male and female microbiomes may react differently to stimuli. This finding is consistent with observations in mice indicating that aging leads to sex-specific remodeling of the intestinal microbiome89 and that diet- and stress-induced cecal and fecal microbiome changes are sex-dependent90.
As with our observations in the microbiome, we found sex differences in the intestinal metabolome. The metabolomes of various bodily fluids and tissues have been shown to respond to stimuli and disease states in a sex-dependent manner across a range of species91,92,93, suggesting that sex differences in the metabolome are persistent and may be linked to health status. Thus, further work on sex differences in the intestinal microbiomes and metabolomes of Jamaican fruit bats will be useful for better understanding sex differences in various health states, including immune responses to viral infections. To our knowledge, this study is one of the first to link microbial community composition to function via metabolites, thus providing a foundation for further research building on the functional role of the bat intestinal microbiome.
Limitations of this study included the fact that we only followed nine bats longitudinally, and of five longitudinally studied females, four were pregnant. These bats were not age-matched; as age may affect microbiome composition, especially during infection studies34,94,95, this could contribute to the noise in our data, as could the fact that the pregnancies observed were not synchronized. Finally, captive bats may not be representative of a population of wild bats. Though experiments in the lab will never fully recapitulate natural phenomena, these experimental studies provide a controlled setup to longitudinally assess immune responses.
The human microbiome has been explored as a predictor for immune-related disease outcomes, including response to rheumatoid arthritis treatment and mortality in allogeneic hematopoietic-cell transplantation96,97. Specific metabolites already reliably predict certain conditions or disease states in humans, including prediabetes and diabetes mellitus98. Few studies in bats have combined microbiome analysis with metabolomics; we hope this study provides a foundation for further investigation of the functional role of the bat intestinal microbiome, allowing for prediction of the results of changes in the microbiome. Given our initial work on associations between bacteria, metabolites linked to intestinal immunity and immunoregulation, and health states (i.e., reproductive status) in this study, intestinal microbial composition could eventually be used to infer disease states and outcomes.
We recognize the logistical challenges inherent in longitudinally sampling and conducting infection studies in bats, especially wild bats. However, accounting for individual variation, potential seasonality, and life course differences in the microbiome will allow for more robust microbiome studies in both captive and wild bats. The methodology described here establishes a foundation for future infection studies conducting immunological assessments of the intestinal microbiomes of Jamaican fruit and other bats, including with viruses that must be handled under high containment conditions. Finally, the integration of metabolomics with microbiome data will allow for more detailed studies of the functional role of the microbiome and the effects of microbiome changes on immunity with the intestinal metabolome as a mediator, potentially leading to the bat intestinal microbiome as an effective predictive tool for health and disease states.
Materials and methods
Ethics statement
All animal experiments were conducted in an AAALAC International-accredited facility and were approved by the Rocky Mountain Laboratories (RML) Animal Institutional Care and Use Committee following the guidelines put forth in the Guide for the Care and Use of Laboratory Animals 8th edition, the Animal Welfare Act, United States Department of Agriculture and the United States Public Health Service Policy on the Humane Care and Use of Laboratory Animals. The study is reported in accordance with ARRIVE guidelines.
Biosafety
All experiments were conducted as if under BSL-3 or BSL-4 conditions as approved by the Institutional Biosafety Committee (IBC). For the removal of specimens from high containment areas, inactivation of all samples was performed according to IBC-approved standard operating procedures.
Animals
Mixed-sex healthy young adult to adult Jamaican fruit bats were transferred from Colorado State University (CSU)’s closed colony to RML (approx. weight range = 32.3 g to 55.5 g upon intake at RML). At CSU, a colony of several hundred bats was housed in an open mixed-sex free-flight facility99. No previous procedures were performed on any animal, and animals were not genetically modified or immunosuppressed. The temperature range was kept between 22 and 26 °C, with humidity between 30 and 75%. The enclosure’s floors, walls, and equipment were sanitized biweekly. Due to (1) no prior availability of microbiome data on which to base a power analysis for this species and (2) the observational and descriptive nature of this study, no power calculation was performed to determine group sizes. For similar reasons, alongside the lack of an explicit treatment group, each animal’s own baseline is used as its reference point, and no negative control group was included. Animals were randomly selected for enrollment from the full colony at CSU. After transport, animals were randomly housed in new cage groups and kept separated by sex. For all longitudinal sampling, the subset of sampled animals was randomly assigned prior to study start. No animals were excluded after enrollment. Pentobarbital sodium and phenytoin sodium after isoflurane inhalation was used for euthanasia of the October cohort; bilateral thoracotomy after isoflurane inhalation was used for euthanasia of the May cohort.
A first shipment (October 2021) consisted of 20 bats, 10 male and 10 female, of which two were pregnant. The second shipment (May 2022) consisted of 30 bats, 9 male and 21 female, of which 10 were pregnant (Supplementary Table 1). Physical exams were performed on all bats at intake, and bats were cohoused in our facility in sex-separated groups of up to five bats per cage. Cages were cleaned with water and cage pans disinfected daily with 5% Micro-Chem Plus Detergent Disinfectant (NCL). Bats were provided a diet of various fresh, non-citrus fruits supplemented with 15 g of Mazuri Softbill Diet (Mazuri) per 10 bats daily. Fruit was provided ad libitum and replaced twice daily. Temperature and humidity were kept constant and in the same range as the CSU facility. Bats were allowed a minimum of 5 days to acclimate to the facility prior to study onset. All cages were housed within the same room and provided the same diet to avoid environmental confounding. Any environmental differences that may have occurred between October and May, as well as seasonal availability and sourcing of fresh fruit, were not controlled for beyond what was described above. Animal care staff and experimenters were aware that bats belonged either to the October or May cohort. Sequencing and downstream analysis were performed blinded until results were allocated to the metadata.
Cross-sectional microbiome sample collection
Rectal swabs and fecal samples were obtained from 20 bats in the October cohort, and rectal swabs were obtained from 30 bats in the May cohort after a 5- or 6-day acclimation period but within a week of their arriving at our facility. Swabs and fecal samples were either frozen immediately after collection or transferred to TRIzol (ThermoFisher) and stored at − 80 °C. All swabs were collected using pre-wetted Puritan 6″ Sterile Mini-Tip Polyester Swabs with Ultrafine Polyester Handles (Puritan) under the influence of 5% isoflurane anesthesia.
A subset of samples was used to establish the methods for microbiome sampling and analysis of samples coming from maximum containment (BSL-4) conditions. Due to limited availability of bat samples, these methods were first tested and optimized using feces collected from cages of an in-house breeding colony of multimammate rats (Mastomys natalensis) maintained at our facility100. Fecal samples were stored at – 80 °C until use. Nucleic acid extraction was performed using one of TRIzol Reagent (ThermoFisher), the AllPrep DNA/RNA Kit (Qiagen), or the QIAamp Fast DNA Stool Mini Kit (Qiagen) per manufacturer’s instructions. The concentration of DNA was quantified with the NanoDrop 8000 (Thermo Scientific) before use in downstream applications.
Metabolome and longitudinal microbiome sample collection
Swabs were collected from a subset of animals (N = 9; female, N = 5 and male, N = 4; pregnant, N = 4 and not pregnant, N = 1) in the May cohort for longitudinal microbiome analysis. Rectal swabs were taken from these bats on each of days 1, 4, 7, 14, and 28. On day 28, all nine animals were euthanized, necropsied, and a 1 cm sample of ileal tissue was collected from each bat for metabolome analysis.
Nucleic acid extraction for cross-sectional and longitudinal microbiome analysis of rectal swab samples
Swabs were immediately transferred to TRIzol (Invitrogen) and frozen at − 80 °C until use. DNA was isolated using TRIzol reagent per manufacturer’s instructions with a final resuspension volume of 0.5 mL.
Nucleic acid amplification
The V3 and V4 regions of the 16S rRNA gene were amplified via amplicon PCR using 12.5 μL 2× KAPA HiFi HotStart ReadyMix (Roche) and 5 μL each of established primers101 at 1 μM each with 2.5 μL of extracted DNA. Cycling conditions were as follows: 95 °C for 3 min, followed by 35 cycles of 95 °C for 30 s, 55 °C for 30 s, and 72 °C for 30 s. PCR cycling conditions were based on manufacturer’s instructions (Illumina) and modified for bat samples. Samples were then held at 72 °C for 5 min followed by 4 °C indefinitely. PCR products were run on 1% agarose gels at 120 V for 25 min to verify successful amplification of the desired region before sequencing and library preparation.
Illumina sequencing of 16S rRNA gene
Each sample was indexed and cleaned for library preparation per manufacturer’s instructions (Illumina). Twenty-five microliters of supernatant from each sample was collected for sequencing. Samples were fragment-sized using either a BioAnalyzer DNA 1000 chip (Agilent, October cohort) or a TapeStation 4200 (Agilent, May cohort) and quantitated using KAPA Library Quant Kit (Illumina) Universal qPCR Mix (KAPA Biosystems). Samples were diluted and multiplexed into a single pool using equal volumes. For the May cohort samples, an initial nano flow cell on a MiSeq Sequencing System (Illumina), which sequenced 150 cycles in each read direction for a total of 300 cycles each, was run to normalize the sample pool. Finally, a titration with 7% PhiX was used to cluster one V3 flow cell on a MiSeq Sequencing System (Illumina), which was sequenced for 300 cycles in each read direction for a total of 600 cycles each.
Microbiome data analysis
The Quantitative Insights into Microbial Ecology 2 (QIIME2) tool (version 2021.7; open-source software)102 pipeline was used to trim and quality filter sequences. Amplicon sequence variants (ASVs) were determined using Deblur103 and clustered into de novo operational taxonomic units (OTUs) at the 98% similarity level using VSEARCH104. Rarefaction curves to assess species richness and comparability of the samples were also generated through these pipelines. Chimeras were removed, and taxonomy was assigned to OTUs using a classifier built with Scikit-learn105 and verified against the Greengenes106 16S rRNA gene database. A phylogenetic tree for diversity analyses was generated with the FastTree pipeline107. QIIME2 was used to process the dataset. As QIIME2 uses rarefaction as a built-in normalization method (to remove bias due to variable sequencing depths), downstream analyses—including ordination and differential abundance analysis—were performed on normalized data.
Metabolite and lipid sample preparation
Liquid chromatography-mass spectrometry (LCMS) grade solvents were used for all LCMS methods. Tissue samples were immersed directly in 0.4 mL of methanol and shredded at 30Hz for 10 min using a tissue shredder and one stainless steel bead (Qiagen, 5 mm) per sample. Supernatant was then irradiated at 2 mRad for sample removal from high containment. Then, 0.4 mL of water and 0.4 mL of chloroform were added to each sample. Samples were shaken for 30 min at 4 °C and centrifuged at 16,000×g for 20 min to establish layering. Four hundred microliters of the top (aqueous) layer was collected. The aqueous layer was diluted 5× in 50% methanol in water for LCMS injection. A subaliquot of the aqueous layer was taken for O-benzylhydroxylamine derivatization of carboxylic acids and SCFA analysis.
Short chain fatty acid derivatization
Samples were derivatized with O-benzylhydroxylamine (O-BHA) according to previously established protocols with modifications108,109. A reaction buffer consisting of 1 M pyridine and 0.5 M hydrochloric acid in water was prepared fresh. A volume of 35 µL of the aqueous metabolite extract was sub-aliquoted. Ten microliters of 1 M O-BHA in reaction buffer and 10 µL of 1 M 1-ethyl-3-(3-dimethylaminopropyl)carbodiimide in reaction buffer were added to the sample. Samples were shaken at room temperature for 2 h. The reaction was quenched with 50 µL of 0.1% formic acid for 10 min. Derivatized carboxylic acid compounds were extracted via the addition of 400 µL ethyl acetate. Following mixing and centrifugation at 16,000×g for 5 min at 4 °C to induce layering, the upper (organic) layer was collected and dried under vacuum. Samples were resuspended in 300 µL of water for LCMS injection.
Liquid chromatography–mass spectrometry
Tributylamine and all synthetic molecular references were purchased from Millipore Sigma. Methanol, LCMS grade water, isopropanol, and acetic acid were purchased through Fisher Scientific. All samples were separated using a Sciex ExionLC™ AC system and measured using a Sciex 5500 QTRAP® mass spectrometer. Aqueous metabolites were analyzed using a previously established ion pairing method with modification110. Quality control samples were injected after every 10 injections. Samples were separated on a Waters™ Atlantis T3 column (100 Å, 3 µm, 3 mm × 100 mm) and eluted using a binary gradient from 5 mM tributylamine, 5 mM acetic acid in 2% isopropanol, 5% methanol, 93% water (v/v) to 100% isopropanol over 15 min. Two distinct MRM pairs in negative mode were used for each metabolite. Derivatized short chain fatty acid samples were separated with a Waters™ Atlantis dC18 column (100 Å, 3 µm, 3 mm × 100 mm) and eluted using a 6-min gradient from 5 to 80% B with buffer A as 0.1% formic acid in water and B as 0.1% formic acid in methanol. Short chain fatty acids and central metabolic carboxylic acids were detected using MRMs from previously established methods, and identity was confirmed by comparison to derivatized standards108,109. All signals were integrated using MultiQuant® Software 3.0.3. Signals with greater than 50% missing values were discarded, and remaining missing values were replaced with the lowest registered signal value. All signals with a QC coefficient of variance greater than 30% were discarded. Metabolites with multiple MRMs were quantified with the higher intensity MRM. Filtered datasets were total sum normalized prior to analysis. Short chain fatty acid datasets were stitched to their corresponding polar metabolite dataset via common signals for lactate.
Data analysis
Percent abundance of microbial families in each sample was calculated and visualized in GraphPad Prism (version 9.3.1). Alpha diversity (observed features, Shannon entropy, Simpson’s diversity, and Faith’s phylogenetic diversity) of each sample was calculated in QIIME2. Alpha diversity was visualized, and statistical significance (Mann–Whitney tests for comparison of males to females and October to May cohorts, and Wilcoxon matched pairs tests for longitudinal analysis) calculated, using GraphPad Prism. Beta diversity (Bray–Curtis distance, and weighted and unweighted UniFrac distance) was calculated and visualized in R (version 4.1.3)111 through the RStudio interface (version 2022.02.1)112 using the ecodist113, factoextra114, mia115, and ggplot2116 packages. PERMANOVAs to assess statistical significance of microbial community composition differences were also performed in R using the vegan package117. LEfSe analysis118 was performed in Galaxy and visualized in GraphPad Prism to assess differential abundance across groups. Microbiome Multivariable Associations with Linear Models (MaAsLin2, version 1.12.0) was used to test associations between experimental days and microbial families in R 4.2.3119. Briefly, the unrarefied OTU table was collapsed to the family level, normalized to relative abundance, and log transformed. Families present in > 10% of the total number of samples were selected for the association test. General linear model (LM) was applied with including sex and sample ID as fixed effect and random effects, respectively. p-values were corrected using the Benjamini–Hochberg procedure. Single and multi-variate analysis on metabolomics data was performed in MarkerView® Software 1.3.1 or using the mixOmics R package120 and loadings and variates visualized in GraphPad Prism. Abundances of specific metabolites were visualized and analyzed in GraphPad Prism. Correlations between metabolites and bacterial genera were calculated and visualized in GraphPad Prism.
Data availability
Data to be deposited in Figshare under https://doi.org/10.6084/m9.figshare.24917568. The sequencing data are accessible on NCBI’s Sequence Read Archive under Accession Number SRP440894.
References
Yeo, J.-M. et al. Lactobacillus fermentum CJL-112 protects mice against influenza virus infection by activating T-helper 1 and eliciting a protective immune response. Int. Immunopharmacol. 18, 50–54. https://doi.org/10.1016/j.intimp.2013.10.020 (2014).
Reinold, J. et al. A pro-inflammatory gut microbiome characterizes SARS-CoV-2 infected patients and a reduction in the connectivity of an anti-inflammatory bacterial network associates with severe COVID-19. Front. Cell Infect. Microbiol. 11, 747816. https://doi.org/10.3389/fcimb.2021.747816 (2021).
Sarkar, A. et al. The gut microbiome as a biomarker of differential susceptibility to SARS-CoV-2. Trends Mol. Med. 27, 1115–1134. https://doi.org/10.1016/j.molmed.2021.09.009 (2021).
Yeoh, Y. K. et al. Gut microbiota composition reflects disease severity and dysfunctional immune responses in patients with COVID-19. Gut 70, 698–706. https://doi.org/10.1136/gutjnl-2020-323020 (2021).
Song, J. A. et al. Oral intake of Lactobacillus rhamnosus M21 enhances the survival rate of mice lethally infected with influenza virus. J. Microbiol. Immunol. Infect. 49, 16–23. https://doi.org/10.1016/j.jmii.2014.07.011 (2016).
Yang, X. L. et al. The intestinal microbiome primes host innate immunity against enteric virus systemic infection through type I interferon. mBio 12, 21. https://doi.org/10.1128/mBio.00366-21 (2021).
Heintz-Buschart, A. & Wilmes, P. Human gut microbiome: Function matters. Trends Microbiol. 26, 563–574. https://doi.org/10.1016/j.tim.2017.11.002 (2018).
Zheng, D. P., Liwinski, T. & Elinav, E. Interaction between microbiota and immunity in health and disease. Cell Res. 30, 492–506. https://doi.org/10.1038/s41422-020-0332-7 (2020).
Neu, J. & Pammi, M. Necrotizing enterocolitis: The intestinal microbiome, metabolome and inflammatory mediators. Semin. Fetal Neonatal Med. 23, 400–405. https://doi.org/10.1016/j.siny.2018.08.001 (2018).
Sun, S. et al. Altered intestinal microbiome and metabolome correspond to the clinical outcome of sepsis. Crit. Care 27, 127. https://doi.org/10.1186/s13054-023-04412-x (2023).
Morrison, D. J. & Preston, T. Formation of short chain fatty acids by the gut microbiota and their impact on human metabolism. Gut Microbes 7, 189–200. https://doi.org/10.1080/19490976.2015.1134082 (2016).
Luhrs, H. et al. Butyrate inhibits NF-kappaB activation in lamina propria macrophages of patients with ulcerative colitis. Scand. J. Gastroenterol. 37, 458–466. https://doi.org/10.1080/003655202317316105 (2002).
Arpaia, N. et al. Metabolites produced by commensal bacteria promote peripheral regulatory T-cell generation. Nature 504, 451–455. https://doi.org/10.1038/nature12726 (2013).
Amman, B. R., Swanepoel, R., Nichol, S. T. & Towner, J. S. Ecology of filoviruses. Curr. Top. Microbiol. Immunol. 411, 23–61. https://doi.org/10.1007/82_2017_10 (2017).
Zhou, P. et al. A pneumonia outbreak associated with a new coronavirus of probable bat origin. Nature 579, 270–273. https://doi.org/10.1038/s41586-020-2012-7 (2020).
Amman, B. R. et al. A recently discovered pathogenic paramyxovirus, sosuga virus, is present in Rousettus aegyptiacus fruit bats at multiple locations in Uganda. J. Wildl. Dis. 51, 774–779. https://doi.org/10.7589/2015-02-044 (2015).
Guth, S. et al. Bats host the most virulent—But not the most dangerous—Zoonotic viruses. Proc. Natl. Acad. Sci. 119, e2113628119. https://doi.org/10.1073/pnas.2113628119 (2022).
Banerjee, A., Rapin, N., Bollinger, T. & Misra, V. Lack of inflammatory gene expression in bats: A unique role for a transcription repressor. Sci. Rep. 7, 2232. https://doi.org/10.1038/s41598-017-01513-w (2017).
Ahn, M., Cui, J., Irving, A. T. & Wang, L.-F. Unique loss of the PYHIN gene family in bats amongst mammals: Implications for inflammasome sensing. Sci. Rep. 6, 21722. https://doi.org/10.1038/srep21722 (2016).
Ahn, M. et al. Dampened NLRP3-mediated inflammation in bats and implications for a special viral reservoir host. Nat. Microbiol. 4, 789–799. https://doi.org/10.1038/s41564-019-0371-3 (2019).
Goh, G. et al. Complementary regulation of caspase-1 and IL-1β reveals additional mechanisms of dampened inflammation in bats. Proc. Natl. Acad. Sci. 117, 28939–28949. https://doi.org/10.1073/pnas.2003352117 (2020).
Xie, J. et al. Dampened STING-dependent interferon activation in bats. Cell Host Microbe 23, 297–301. https://doi.org/10.1016/j.chom.2018.01.006 (2018).
Ahn, M. et al. Bat ASC2 suppresses inflammasomes and ameliorates inflammatory diseases. Cell 186, 2144–2159. https://doi.org/10.1016/j.cell.2023.03.036 (2023).
Banerjee, A. et al. Novel insights into immune systems of bats. Front. Immunol. 11, 26. https://doi.org/10.3389/fimmu.2020.00026 (2020).
Zhou, P. et al. Contraction of the type I IFN locus and unusual constitutive expression of IFN-b1 in bats. Proc. Natl. Acad. Sci. 113, 2696–2701. https://doi.org/10.1073/pnas.1518240113 (2016).
Shen, Y. Y. et al. Adaptive evolution of energy metabolism genes and the origin of flight in bats. Proc. Natl. Acad. Sci. U.S.A. 107, 8666–8671. https://doi.org/10.1073/pnas.0912613107 (2010).
Song, S. J. et al. Comparative analyses of vertebrate gut microbiomes reveal convergence between birds and bats. mBio 11, 19. https://doi.org/10.1128/mBio.02901-19 (2020).
Caviedes-Vidal, E. et al. The digestive adaptation of flying vertebrates: High intestinal paracellular absorption compensates for smaller guts. Proc. Natl. Acad. Sci. U.S.A. 104, 19132–19137. https://doi.org/10.1073/pnas.0703159104 (2007).
Jones, D. N., Ravelomanantsoa, N. A. F., Yeoman, C. J., Plowright, R. K. & Brook, C. E. Do gastrointestinal microbiomes play a role in bats’ unique viral hosting capacity? Trends Microbiol. 30, 632–642. https://doi.org/10.1016/j.tim.2021.12.009 (2022).
Luo, J., Liang, S. & Jin, F. Gut microbiota in antiviral strategy from bats to humans: A missing link in COVID-19. Sci. China Life Sci. 64, 942–956. https://doi.org/10.1007/s11427-020-1847-7 (2021).
Shin, N. R., Whon, T. W. & Bae, J. W. Proteobacteria: Microbial signature of dysbiosis in gut microbiota. Trends Biotechnol. 33, 496–503. https://doi.org/10.1016/j.tibtech.2015.06.011 (2015).
Liu, B. et al. The gut microbiota of bats confers tolerance to influenza virus (H1N1) infection in mice. Transbound Emerg. Dis. https://doi.org/10.1111/tbed.14478 (2022).
Ingala, M. R., Becker, D. J., BakHolm, J., Kristiansen, K. & Simmons, N. B. Habitat fragmentation is associated with dietary shifts and microbiota variability in common vampire bats. Ecol. Evol. 9, 6508–6523. https://doi.org/10.1002/ece3.5228 (2019).
Wasimuddin, et al. Astrovirus infections induce age-dependent dysbiosis in gut microbiomes of bats. ISME J. 12, 2883–2893. https://doi.org/10.1038/s41396-018-0239-1 (2018).
Moreira-Soto, A. et al. Neotropical bats from Costa Rica harbour diverse coronaviruses. Zoonoses Public Health 62, 501–505. https://doi.org/10.1111/zph.12181 (2015).
Munster, V. J. et al. Replication and shedding of MERS-CoV in Jamaican fruit bats (Artibeus jamaicensis). Sci. Rep. 6, 21878. https://doi.org/10.1038/srep21878 (2016).
Cogswell-Hawkinson, A. et al. Tacaribe virus causes fatal infection of an ostensible reservoir host, the Jamaican fruit bat. J. Virol. 86, 5791–5799. https://doi.org/10.1128/JVI.00201-12 (2012).
Reid, J. E. & Jackson, A. C. Experimental rabies virus infection in Artibeus jamaicensis bats with CVS-24 variants. J. Neurovirol. 7, 511–517. https://doi.org/10.1080/135502801753248097 (2001).
Ciminski, K. et al. Bat influenza viruses transmit among bats but are poorly adapted to non-bat species. Nat. Microbiol. 4, 2298–2309. https://doi.org/10.1038/s41564-019-0556-9 (2019).
Malmlov, A. et al. Experimental Zika virus infection of Jamaican fruit bats (Artibeus jamaicensis) and possible entry of virus into brain via activated microglial cells. PLoS Negl. Trop. Dis. 13, e0007071. https://doi.org/10.1371/journal.pntd.0007071 (2019).
Illumina. 16S Metagenomic Sequencing Library Preparation. https://support.illumina.com/documents/documentation/chemistry_documentation/16s/16s-metagenomic-library-prep-guide-15044223-b.pdf.
Klein, S. L. & Flanagan, K. L. Sex differences in immune responses. Nat. Rev. Immunol. 16, 626–638. https://doi.org/10.1038/nri.2016.90 (2016).
Takahashi, T. & Iwasaki, A. Sex differences in immune responses. Science 371, 347–348. https://doi.org/10.1126/science.abe7199 (2021).
Cornish, E. F., Filipovic, I., Asenius, F., Williams, D. J. & McDonnell, T. Innate immune responses to acute viral infection during pregnancy. Front. Immunol. 11, 572567. https://doi.org/10.3389/fimmu.2020.572567 (2020).
Haddock, E. & Feldmann, F. Validating the inactivation effectiveness of chemicals on Ebola virus. Methods Mol. Biol. 1628, 251–257. https://doi.org/10.1007/978-1-4939-7116-9_20 (2017).
Haddock, E., Feldmann, F. & Feldmann, H. Effective chemical inactivation of Ebola virus. Emerg. Infect. Dis. 22, 1292–1294. https://doi.org/10.3201/eid2207.160233 (2016).
Haddock, E., Feldmann, F., Shupert, W. L. & Feldmann, H. Inactivation of SARS-CoV-2 laboratory specimens. Am. J. Trop. Med. Hyg. 104, 2195–2198. https://doi.org/10.4269/ajtmh.21-0229 (2021).
Chen, L. et al. The long-term genetic stability and individual specificity of the human gut microbiome. Cell 184, 2302–2315. https://doi.org/10.1016/j.cell.2021.03.024 (2021).
Rudolph, K. et al. Drivers of gut microbiome variation within and between groups of a wild Malagasy primate. Microbiome 10, 28. https://doi.org/10.1186/s40168-021-01223-6 (2022).
Yarlagadda, K., Razik, I., Malhi, R. S. & Carter, G. G. Social convergence of gut microbiomes in vampire bats. Biol. Lett. 17, 20210389. https://doi.org/10.1098/rsbl.2021.0389 (2021).
Ortega, J. & Arita, H. T. Structure and social dynamics of harem groups in Artibeus jamaicensis (Chiroptera: Phyllostomidae). J. Mammal. 80, 1173–1185. https://doi.org/10.2307/1383168 (1999).
Kunz, T. H., August, P. V. & Burnett, C. D. Harem social-organization in cave roosting Artibeus jamaicensis (Chiroptera, Phyllostomidae). Biotropica 15, 133–138. https://doi.org/10.2307/2387958 (1983).
Neu, A. T., Allen, E. E. & Roy, K. Defining and quantifying the core microbiome: Challenges and prospects. Proc. Natl. Acad. Sci. U.S.A. 118, 51. https://doi.org/10.1073/pnas.2104429118 (2021).
Yang, J. & Chun, J. Taxonomic composition and variation in the gut microbiota of laboratory mice. Mamm. Genome 32, 297–310. https://doi.org/10.1007/s00335-021-09871-7 (2021).
Edenborough, K. M. et al. Microbiomes in the insectivorous bat species Mops condylurus rapidly converge in captivity. PLoS ONE 15, e0223629. https://doi.org/10.1371/journal.pone.0223629 (2020).
Xiao, Y. et al. Captivity causes taxonomic and functional convergence of gut microbial communities in bats. PeerJ 7, e6844. https://doi.org/10.7717/peerj.6844 (2019).
Martiny, A. C., Treseder, K. & Pusch, G. Phylogenetic conservatism of functional traits in microorganisms. ISME J. 7, 830–838. https://doi.org/10.1038/ismej.2012.160 (2013).
Dietrich, M., Kearney, T., Seamark, E. C. J., Paweska, J. T. & Markotter, W. Synchronized shift of oral, faecal and urinary microbiotas in bats and natural infection dynamics during seasonal reproduction. R. Soc. Open Sci. 5, 180041. https://doi.org/10.1098/rsos.180041 (2018).
Gaona, O., Cerqueda-Garcia, D., Moya, A., Neri-Barrios, X. & Falcon, L. I. Geographical separation and physiology drive differentiation of microbial communities of two discrete populations of the bat Leptonycteris yerbabuenae. Microbiologyopen 9, 1113–1127. https://doi.org/10.1002/mbo3.1022 (2020).
Gaona, O., Gomez-Acata, E. S., Cerqueda-Garcia, D., Neri-Barrios, C. X. & Falcon, L. I. Fecal microbiota of different reproductive stages of the central population of the lesser-long nosed bat, Leptonycteris yerbabuenae. PLoS ONE 14, e0219982. https://doi.org/10.1371/journal.pone.0219982 (2019).
Gong, L., Liu, B., Wu, H., Feng, J. & Jiang, T. Seasonal dietary shifts alter the gut microbiota of avivorous bats: Implication for adaptation to energy harvest and nutritional utilization. mSphere 6, e0046721. https://doi.org/10.1128/mSphere.00467-21 (2021).
Li, J. et al. Fecal bacteriome and mycobiome in bats with diverse diets in South China. Curr. Microbiol. 75, 1352–1361. https://doi.org/10.1007/s00284-018-1530-0 (2018).
Zepeda Mendoza, M. L. et al. Hologenomic adaptations underlying the evolution of sanguivory in the common vampire bat. Nat. Ecol. Evol. 2, 659–668. https://doi.org/10.1038/s41559-018-0476-8 (2018).
Houtz, J. L., Sanders, J. G., Denice, A. & Moeller, A. H. Predictable and host-species specific humanization of the gut microbiota in captive primates. Mol. Ecol. 30, 3677–3687. https://doi.org/10.1111/mec.15994 (2021).
Trevelline, B. K. & Moeller, A. H. Robustness of mammalian gut microbiota to humanization in captivity. Front. Ecol. Evol. 9, 5089. https://doi.org/10.3389/fevo.2021.785089 (2022).
Clayton, J. B. et al. Captivity humanizes the primate microbiome. Proc. Natl. Acad. Sci. U.S.A. 113, 10376–10381. https://doi.org/10.1073/pnas.1521835113 (2016).
Banskar, S., Bhute, S. S., Suryavanshi, M. V., Punekar, S. & Shouche, Y. S. Microbiome analysis reveals the abundance of bacterial pathogens in Rousettus leschenaultii guano. Sci. Rep. 6, 36948. https://doi.org/10.1038/srep36948 (2016).
Dietrich, M. & Markotter, W. Studying the microbiota of bats: Accuracy of direct and indirect samplings. Ecol. Evol. 9, 1730–1735. https://doi.org/10.1002/ece3.4842 (2019).
Alwin Prem Anand, A. & Sripathi, K. Digestion of cellulose and xylan by symbiotic bacteria in the intestine of the Indian flying fox (Pteropus giganteus). Compar. Biochem. Physiol. A Mol. Integr. Physiol. 139, 65–69. https://doi.org/10.1016/j.cbpb.2004.07.006 (2004).
Henry, R. et al. What’s the risk? Identifying potential human pathogens within grey-headed flying foxes faeces. PLoS ONE 13, e0191301. https://doi.org/10.1371/journal.pone.0191301 (2018).
Rinninella, E. et al. What is the healthy gut microbiota composition? A changing ecosystem across age, environment, diet, and diseases. Microorganisms 7, 14. https://doi.org/10.3390/microorganisms7010014 (2019).
Franceschi, C. et al. Inflammaging and anti-inflammaging: A systemic perspective on aging and longevity emerged from studies in humans. Mech. Ageing Dev. 128, 92–105. https://doi.org/10.1016/j.mad.2006.11.016 (2007).
Odamaki, T. et al. Age-related changes in gut microbiota composition from newborn to centenarian: A cross-sectional study. BMC Microbiol. 16, 90. https://doi.org/10.1186/s12866-016-0708-5 (2016).
Biagi, E. et al. The gut microbiota of centenarians: Signatures of longevity in the gut microbiota profile. Mech. Ageing Dev. 165, 180–184. https://doi.org/10.1016/j.mad.2016.12.013 (2017).
Santoro, A. et al. Gut microbiota changes in the extreme decades of human life: A focus on centenarians. Cell Mol. Life Sci. 75, 129–148. https://doi.org/10.1007/s00018-017-2674-y (2018).
Goffredo, M. et al. Role of gut microbiota and short chain fatty acids in modulating energy harvest and fat partitioning in youth. J. Clin. Endocrinol. Metab. 101, 4367–4376. https://doi.org/10.1210/jc.2016-1797 (2016).
Koh, A., De Vadder, F., Kovatcheva-Datchary, P. & Backhed, F. From dietary fiber to host physiology: Short-chain fatty acids as key bacterial metabolites. Cell 165, 1332–1345. https://doi.org/10.1016/j.cell.2016.05.041 (2016).
Silva, Y. P., Bernardi, A. & Frozza, R. L. The role of short-chain fatty acids from gut microbiota in gut–brain communication. Front. Endocrinol. 11, 25. https://doi.org/10.3389/fendo.2020.00025 (2020).
Yang, W. et al. Intestinal microbiota-derived short-chain fatty acids regulation of immune cell IL-22 production and gut immunity. Nat. Commun. 11, 4457. https://doi.org/10.1038/s41467-020-18262-6 (2020).
Zhang, W. et al. Flos Abelmoschus manihot extract attenuates DSS-induced colitis by regulating gut microbiota and Th17/Treg balance. Biomed. Pharmacother. 117, 109162. https://doi.org/10.1016/j.biopha.2019.109162 (2019).
Kishiro, Y., Ueda, K., Fujiwara, M. & Yamamoto, I. Butyrate enhances the in vitro anti-SRBC (sheep red blood cell) antibody responses in murine splenocytes. Jpn. J. Pharmacol. 66, 369–376. https://doi.org/10.1254/jjp.66.369 (1994).
Ozogul, F., Kuley, E., Ozogul, Y. & Ozogul, I. The function of lactic acid bacteria on biogenic amines production by food-borne pathogens in arginine decarboxylase broth. Food Sci. Technol. Res. 18, 795–804 (2012).
Williams, B. B. et al. Discovery and characterization of gut microbiota decarboxylases that can produce the neurotransmitter tryptamine. Cell Host Microbe 16, 495–503. https://doi.org/10.1016/j.chom.2014.09.001 (2014).
Agus, A., Planchais, J. & Sokol, H. Gut microbiota regulation of tryptophan metabolism in health and disease. Cell Host Microbe 23, 716–724. https://doi.org/10.1016/j.chom.2018.05.003 (2018).
Strazar, M. et al. Gut microbiome-mediated metabolism effects on immunity in rural and urban African populations. Nat. Commun. 12, 4845. https://doi.org/10.1038/s41467-021-25213-2 (2021).
Fan, Y. et al. Multi-omics analysis reveals aberrant gut-metabolome-immune network in Schizophrenia. Front. Immunol. 13, 812293. https://doi.org/10.3389/fimmu.2022.812293 (2022).
Wu, G. et al. Integrated microbiome and metabolome analysis reveals the potential therapeutic mechanism of Qing–Fei–Pai–Du decoction in mice with coronavirus-induced pneumonia. Front. Cell Infect. Microbiol. 12, 950983. https://doi.org/10.3389/fcimb.2022.950983 (2022).
Phillips, C. D. et al. Microbiome analysis among bats describes influences of host phylogeny, life history, physiology and geography. Mol. Ecol. 21, 2617–2627. https://doi.org/10.1111/j.1365-294X.2012.05568.x (2012).
Ma, J. et al. Gut microbiota remodeling reverses aging-associated inflammation and dysregulation of systemic bile acid homeostasis in mice sex-specifically. Gut Microbes 11, 1450–1474. https://doi.org/10.1080/19490976.2020.1763770 (2020).
Lyte, J. M., Koester, L. R., Daniels, K. M. & Lyte, M. Distinct cecal and fecal microbiome responses to stress are accompanied by sex- and diet-dependent changes in behavior and gut serotonin. Front. Neurosci. 16, 827343. https://doi.org/10.3389/fnins.2022.827343 (2022).
Chi, L. et al. Nicotine alters the gut microbiome and metabolites of gut-brain interactions in a sex-specific manner. Chem. Res. Toxicol. 30, 2110–2119. https://doi.org/10.1021/acs.chemrestox.7b00162 (2017).
Zhang, D. et al. Sex-dependent changes in the microbiota profile, serum metabolism, and hormone levels of growing pigs after dietary supplementation with Lactobacillus. Appl. Microbiol. Biotechnol. 105, 4775–4789. https://doi.org/10.1007/s00253-021-11310-1 (2021).
Tso, V. K. et al. Metabolomic profiles are gender, disease and time specific in the interleukin-10 gene-deficient mouse model of inflammatory bowel disease. PLoS ONE 8, e67654. https://doi.org/10.1371/journal.pone.0067654 (2013).
Wu, C. S. et al. Age-dependent remodeling of gut microbiome and host serum metabolome in mice. Aging (Albany N.Y.) 13, 6330–6345. https://doi.org/10.18632/aging.202525 (2021).
Yin, Z. et al. Changes in the gut microbiota during Asian particolored bat (Vespertilio sinensis) development. PeerJ 8, e9003. https://doi.org/10.7717/peerj.9003 (2020).
Peled, J. U. et al. Microbiota as predictor of mortality in allogeneic hematopoietic-cell transplantation. N. Engl. J. Med. 382, 822–834. https://doi.org/10.1056/NEJMoa1900623 (2020).
Yan, H. et al. Pharmacomicrobiology of methotrexate in rheumatoid arthritis: Gut microbiome as predictor of therapeutic response. Front. Immunol. 12, 789334. https://doi.org/10.3389/fimmu.2021.789334 (2021).
Arneth, B., Arneth, R. & Shams, M. Metabolomics of type 1 and type 2 diabetes. Int. J. Mol. Sci. 20, 467. https://doi.org/10.3390/ijms20102467 (2019).
Schountz, T. Immunology of bats and their viruses: Challenges and opportunities. Viruses 6, 4880–4901. https://doi.org/10.3390/v6124880 (2014).
Safronetz, D. et al. Establishment of a genetically confirmed breeding colony of Mastomys natalensis from wild-caught founders from West Africa. Viruses 13, 590. https://doi.org/10.3390/v13040590 (2021).
Klindworth, A. et al. Evaluation of general 16S ribosomal RNA gene PCR primers for classical and next-generation sequencing-based diversity studies. Nucleic Acids Res. 41, e1. https://doi.org/10.1093/nar/gks808 (2013).
Bolyen, E. et al. Reproducible, interactive, scalable and extensible microbiome data science using QIIME 2. Nat. Biotechnol. 37, 852–857. https://doi.org/10.1038/s41587-019-0209-9 (2019).
Amir, A. et al. Deblur rapidly resolves single-nucleotide community sequence patterns. mSystems 2, 16. https://doi.org/10.1128/mSystems.00191-16 (2017).
Rognes, T., Flouri, T., Nichols, B., Quince, C. & Mahe, F. VSEARCH: A versatile open source tool for metagenomics. PeerJ 4, e2584. https://doi.org/10.7717/peerj.2584 (2016).
Pedregosa, F. et al. Scikit-learn: Machine learning in python. J. Mach. Learn. Res. 12, 2825–2830 (2011).
DeSantis, T. Z. et al. Greengenes, a chimera-checked 16S rRNA gene database and workbench compatible with ARB. Appl. Environ. Microbiol. 72, 5069–5072. https://doi.org/10.1128/AEM.03006-05 (2006).
Price, M. N., Dehal, P. S. & Arkin, A. P. FastTree 2—Approximately maximum-likelihood trees for large alignments. PLoS ONE 5, e9490. https://doi.org/10.1371/journal.pone.0009490 (2010).
Zeng, M. & Cao, H. Fast quantification of short chain fatty acids and ketone bodies by liquid chromatography–tandem mass spectrometry after facile derivatization coupled with liquid-liquid extraction. J. Chromatogr. B Anal. Technol. Biomed. Life Sci. 1083, 137–145. https://doi.org/10.1016/j.jchromb.2018.02.040 (2018).
Jaochico, A., Sangaraju, D. & Shahidi-Latham, S. K. A rapid derivatization based LC–MS/MS method for quantitation of short chain fatty acids in human plasma and urine. Bioanalysis 11, 741–753. https://doi.org/10.4155/bio-2018-0241 (2019).
McCloskey, D., Gangoiti, J. A., Palsson, B. O. & Feist, A. M. A pH and solvent optimized reverse-phase ion-paring-LC-MS/MS method that leverages multiple scan-types for targeted absolute quantification of intracellular metabolites. Metabolomics 11, 1338–1350. https://doi.org/10.1007/s11306-015-0790-y (2015).
R: A Language and Environment for Statistical Computing v. 4.1.3 (R Foundation for Statistical Computing, 2022).
RStudio: Integrated Development for R v. 2022.02.1 (PBC, 2020).
Goslee, S. C. & Urban, D. L. The ecodist package for dissimilarity-based analysis of ecological data. J. Stat. Softw. 22, 1–19. https://doi.org/10.18637/jss.v022.i07 (2007).
factoextra: Extract and Visualize the Result of Multivariate Data Analyses v. 1.0.7 (2020).
mia: Microbiome Analysis (2022).
Wickham, H. ggplot2: Elegant Graphics for Data Analysis (Springer, 2009).
vegan: Community Ecology Package v. 2.6-2 (2022).
Segata, N. et al. Metagenomic biomarker discovery and explanation. Genome Biol. 12, R60. https://doi.org/10.1186/gb-2011-12-6-r60 (2011).
Mallick, H. et al. Multivariable association discovery in population-scale meta-omics studies. PLoS Comput. Biol. 17, e1009442. https://doi.org/10.1371/journal.pcbi.1009442 (2021).
Rohart, F., Gautier, B., Singh, A. & Le Cao, K. A. mixOmics: An R package for omics feature selection and multiple data integration. PLoS Comput. Biol. 13, e1005752. https://doi.org/10.1371/journal.pcbi.1005752 (2017).
Acknowledgements
The authors would like to thank the RMVB and CSU animal care staff for caring for the animals. They would like to thank the RTB for assisting with sample clean-up and sequencing and the OOC for their support.
.
Funding
This work was supported by the Intramural Research Program of the National Institute of Allergy and Infectious Diseases (NIAID), National Institutes of Health (1ZIAAI0011179-01).
Author information
Authors and Affiliations
Contributions
Conceptualization, V.J.M. and J.R.P.; methodology, J.C.R., J.L., K.C., B.S., and J.R.P.; data curation, J.C.R., A.S., M.G.H., E.B., Y.Z., and B.S.; writing—original draft preparation, J.C.R., A.S., M.G.H., and B.S.; writing—review and editing, J.C.R., V.J.M., and J.R.P.; visualization, J.C.R., Y.Z., and B.S.; resources, L.V.K., C.M.B., and T.S.; supervision, V.J.M. and J.R.P. All authors have read and agreed to the published version of the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Riopelle, J.C., Shamsaddini, A., Holbrook, M.G. et al. Sex differences and individual variability in the captive Jamaican fruit bat (Artibeus jamaicensis) intestinal microbiome and metabolome. Sci Rep 14, 3381 (2024). https://doi.org/10.1038/s41598-024-53645-5
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-024-53645-5
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.