Abstract
Humans are regularly cited as the main driver of current biodiversity extinction, but the impact of historic volcanic activity is often overlooked. Pre-human evidence of wildlife abundance and diversity are essential for disentangling anthropogenic impacts from natural events. Réunion Island, with its intense and well-documented volcanic activity, endemic biodiversity, long history of isolation and recent human colonization, provides an opportunity to disentangle these processes. We track past demographic changes of a critically endangered seabird, the Mascarene petrel Pseudobulweria aterrima, using genome-wide SNPs. Coalescent modeling suggested that a large ancestral population underwent a substantial population decline in two distinct phases, ca. 125,000 and 37,000 years ago, coinciding with periods of major eruptions of Piton des Neiges. Subsequently, the ancestral population was fragmented into the two known colonies, ca. 1500 years ago, following eruptions of Piton de la Fournaise. In the last century, both colonies declined significantly due to anthropogenic activities, and although the species was initially considered extinct, it was rediscovered in the 1970s. Our findings suggest that the current conservation status of wildlife on volcanic islands should be firstly assessed as a legacy of historic volcanic activity, and thereafter by the increasing anthropogenic impacts, which may ultimately drive species towards extinction.
Similar content being viewed by others
Introduction
Islands support a disproportionate amount of the world’s biodiversity, including an extraordinary level of species endemism, unique functional traits such as flightlessness in birds, and adaptive radiations e.g., Lemuriformes clade in Madagascar1, but they have also experienced extensive biodiversity loss2,3. Human impact has been identified as the primary driver of this biodiversity loss, with many islands’ endemics going extinct following human colonization and the associated habitat fragmentation and introduction of invasive species3,4, including the dodo Raphus cucullatus from Mauritius5,6. Although this narrative assumes that populations of island endemics were abundant before human impact, this assumption is rarely tested as little pre-human colonization information on endemic fauna abundance is available. Natural events, such as historical climatic cycles7,8,9,10 and island-scale volcanic eruptions11,12,13 are also known to impact species distribution and abundance worldwide14. Hence, disentangling anthropogenic impacts from natural events is often limited by the lack of information about historical events that may have led to population collapse before human colonization.
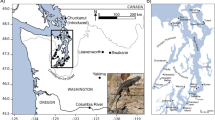
Réunion Island (Western Indian Ocean, WIO), with its intense and well-documented volcanic activity, endemic biodiversity, long history of isolation and comparatively recent human colonization, provides a unique opportunity to disentangle these processes. The island emerged during the late Quaternary (> 2.2 million years ago; Mya) through volcanic activity15, and it was likely colonized by over-sea dispersal from the neighboring islands16. It is composed of two volcanoes: Piton des Neiges, a dormant volcano that has been inactive for about 27 thousand years (kyr)17, and Piton de la Fournaise, one of the most active volcanoes on Earth (Fig. 1a). Human colonization of Réunion Island started around 400 years ago on the west coast, near what became the city of Saint-Paul18.
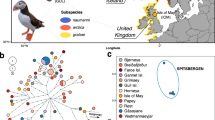
Study area and population genetic structure. (a) Geographic location of the two recently discovered colonies of the Mascarene petrel (Pseudobulweria aterrima) on Réunion Island: Rond Des Chevrons (RDC; yellow star) and Rivière des Remparts (RIR; rose star). The blue triangles represent the location of light-grounded birds (GB). (b) Cluster assignment of Pseudobulweria aterrima individuals to two genetic clusters (K = 2) using 13,855 genome-wide SNPs (n = 87 birds; dataset 1) with Structure. Each single vertical bar represents one individual and each color a distinct genetic cluster. Migrant birds are highlighted by an asterisk in (b). Sampling details can be found in Supplementary Table S1.
High-resolution paleoclimate records (e.g., speleothems, pollen, sediment cores, charcoal and fossils) provide unequivocal archives of pre-human species diversity and distribution, but are often limited or incomplete19,20. In contrast, genomes of extant organisms carry a signature of their evolutionary past, enabling the reconstruction of the demographic history of populations from the genome of their present‐day representatives21,22,23. Such long-term population data can be used to infer the potential drivers of population declines and test alternative evolutionary hypotheses8,9,24,25,26,27. However, even with recent developments of methods for inferring species demographic histories (reviewed in21,28), studies on the endemic flora and fauna of Réunion Island are limited29,30,31, and neglect the possible effects of population structure on demographic inferences32,33,34,35.
To address the lack of long-term population data on the endemic but critically endangered Mascarene petrel Pseudobulweria aterrima (family Procellariidae, Bonaparte 1857) in Réunion Island, we reconstructed its demographic history. The Mascarene petrel is one of the rarest and most threatened seabirds of the world36. The species was previously known from less than 10 specimens collected in Réunion Island during the nineteenth century. Evidence of its continued survival was elusive, and the species was thought to be extinct for more than eight decades, until it was rediscovered in 1970 with a dead grounded bird37. More recently, two breeding colonies have been discovered in Réunion Island: Rond Des Chevrons (hereafter called RDC) and Rivière des Remparts38 (hereafter called RIR; Fig. 1a). This species provides a good study model to investigate the impact of natural versus anthropogenic impacts in the region because (i) the Mascarene petrel has diverged from its congener Fiji Petrel (P. macgillivrayi) from Gau Island (Fiji) and only evolved in Réunion Island39; (ii) as all procellarids, the species is monogamous and highly philopatric, meaning that its population dynamics is strongly impacted by local environmental changes at breeding colonies, and by isolation between colonies40; (iii) the Mascarene petrel is threatened by predation by introduced mammals (rats and cats) at breeding sites and by light-induced mortality (i.e., mortality of fledglings that are attracted to artificial lights)36,41,42. All this suggests that the species is very susceptible to anthropogenic impacts. A genome-wide Single Nucleotide Polymorphism (SNP) dataset was generated with DArTseq for 93 birds from the two recently discovered Mascarene petrel colonies38 and light-grounded birds (Fig. 1a). We first investigated population structure patterns of the Mascarene petrel, and then used two complementary coalescent approaches (i.e., model-free and comparative modeling) to infer changes on effective population size (Ne) and connectivity over time, and investigate how past demographic events were affected by natural versus anthropogenic changes. Specifically, we test whether (i) the Mascarene petrel effective population size was small and fragmented prior to substantial human impact on Réunion Island; (ii) the timing of population declines and fragmentation coincided with major volcanic or climatic events on the island; and (iii) population bottlenecks intensified during the past four decades due to increasing human impact.
Results
Present day population structure in Mascarene petrel
The Discriminant Analysis of Principal Components (DAPC), the Bayesian assignment approach implemented in Structure, and Wright’s F-statistics FST suggested the existence of genetic structure between the two recently discovered breeding colonies of Mascarene petrel. Both DAPC and Structure methods were congruent in inferring K = 2 as the optimal number of genetic clusters (Supplementary Figs. S1a and S2). DAPC clearly separated the birds from RDC and all but six birds from RIR along the first discriminant axis. All grounded birds (attracted by light pollution) were clustered together with the RDC individuals (Supplementary Fig. S1b). When using Structure, assuming K = 2, all the birds from RDC (n = 31) and light-grounded birds (n = 19) were assigned to one of two genetic clusters with an average membership higher than 70%. All but six birds sampled at RIR were grouped in the second cluster with an average membership ranging between 60 and 100% (Fig. 1b; see also Supplementary Fig. S3). Both methods were congruent in inferring the same six migrant birds. Additionally, substantial levels of admixture were observed in both genetic clusters, suggesting the occurrence of historic or contemporary gene flow between the two colonies (Fig. 1b). The Wright’s F-statistics FST supports a significant genetic differentiation among the two colonies, independently of the inclusion (FST = 0.0362; p-value = 0) or exclusion of light-grounded birds (FST = 0.0425; p-value = 0) in the analyses.
Demographic history of Mascarene petrel
The demographic history of the Mascarene petrel was reconstructed using two complementary coalescent approaches. The Stairway Plot v2.043 was firstly used to infer changes on effective population size through time. Since a species' demographic history can be complicated by demographic events other than changes in population size (e.g., population splits or connectivity changes), the composite-likelihood approach implemented in fastsimcoal2 v.2.644 was additionally used to test and compare a set of demographic models incorporating population splits, resizes, and connectivity changes9,29,45,46. The Stairway Plot and fastsimcoal2 are also known to differ on the evolutionary timescales, with the first method being more informative about hundreds of generations, and the latter about the recent past47,48,49,50. The two coalescent methods revealed that Mascarene petrel underwent significant demographic changes during the late Quaternary. Assuming that individuals were part of panmictic populations, the Stairway Plot analyses for each genetic cluster (Supplementary Fig. S4a–c) and for the entire dataset (Fig. 2a) inferred a population maxima ca. 1 Mya, that was preceded by a period of a large and rather constant Ne (> 8 × 105 birds) until ca. 125 kyr. This event was then followed by a continuous population decline until recent times. The analyses suggest a larger Ne for the RDC, even when considering the same number of individuals in the analyses (n = 33 per genetic cluster; Supplementary Fig. S4c). The Stairway Plot runs without closely-related individuals (i.e., parent–offspring and full-sibs; n = 35 for RDC and light-grounded birds, and 19 for RIR) revealed an identical demographic history when using the complete datasets (Supplementary Fig. S4d).
Reconstruction of the demographic history of the Mascarene petrel (Pseudobulweria aterrima) using two complementary coalescent approaches, and major ecological changes on Réunion Island. (a) Demographic history inferred with Stairway Plot, considering the two colonies together (n = 54 unrelated birds). The thick lines correspond to the median values of Ne, and the ribbon represents the 95% confidence interval. The dashed line represents the ancient bottleneck (TbANC1) inferred by model M7 with fastsimcoal2. (b) Illustration of the most realistic model (M7; ancient & recent bottlenecks) revealed by fastsimcoal2. The occurrence of gene flow is exemplified by arrows. (c) Representation of the more pronounced climatic events that took place in Western Indian Ocean: LIG (Last Interglacial; ca. 132–112 kyr; kyr = thousand years), LGM (Last Glacial Maximum; ca. 26.5–19 kyr) and AHP (African Humid Period; ca. 15–5 kyr). The width of bars represents the duration of each event. (d) Review of the major volcanic events that took place on Réunion Island. The width of bars is proportional to the time of activity of Piton des Neiges (brown) and of the activity on the western part of Piton de la Fournaise (yellowish-orange). The arrows represent the origin of Réunion Island (> 2.2 Mya) and Rivière des Remparts valley (ca. 60 kyr). PN4 = Piton des Neiges phase 4. (e) Growth of the human population (green curve) and of production of electricity (dark red) at Réunion Island. Data on human population was compiled from51,52 and INSEE (https://www.insee.fr/fr/statistiques/2522602). Data on electricity production from Electricité de France (EDF). For (a) and (b) analyses were performed considering 2.89 × 10−9 as mutation rate53 and events were scaled considering the generation time of the Barau’s petrel (18.9 years). Population size estimates are given in number of diploid copies. RIR = Rivière des Remparts colony; RDC = Rond Des Chevrons colony; GB = light-grounded birds; Ne = Effective population size; NRIR = Ne of RIR at present time; NRDC = Ne of RDC at present time; N1RIR = Ne of RIR after the fragmentation of the ancestral population into two colonies; N1RDC = Ne of RDC after the fragmentation of the ancestral population into two colonies; NANC2 = Ne of the ancestral population before size changes; Tsplit = Time of the fragmentation of the ancestral population into RIR and RDC; TbRIR = Time when RIR underwent a reduction on population size and connectivity; TbRDC = Time when RDC underwent a reduction on population size and connectivity; TbANC1 = Time when the ancestral population underwent a bottleneck; 2Nm = average number of haploid immigrants entering the population per generation, where 2Nm0 denotes recent gene flow and 2Nm1 ancient gene flow among the two colonies. 2Nm0 = 0.01; 2Nm1 = 0.5.
The likelihood comparison of the seven models (M1–M7) tested with fastsimcoal2 (Supplementary Fig. S5) revealed that models including gene flow (M3, M4, M7) had a better fit than those assuming only population size changes (Fig. 3), suggesting that both events are essential to explain the evolutionary history of the Mascarene petrel. The lowest ΔAIC (difference in Akaike Information Criteria (AIC) to the best demographic model) values were observed for the bottleneck with gene flow model (M3) and the ancient & recent bottlenecks with gene flow model (M7). The model M3 assumed that the fragmentation of the ancestral population into RDC and RIR was followed by a population decline and a reduction in population connectivity between the two colonies. The model M7 parallelized the previous model but included an ancient population decline preceding the fragmentation of the ancestral population into RIR and RDC. Both models exhibited: a similar AIC (AIC = 42,029.1 and 42,031.3; Supplementary Table S2), overlapped on the log10 likelihood distributions across the 100 independent simulations performed per model (Fig. 3), a good fit between the observed and expected 2d-SFS (Supplementary Fig. S6), which combined suggest a similar model fit. Similar to the Stairway Plot, the parameters estimated by M7 suggested a large ancestral population (ca. 4 × 105 birds) that suffered a significant reduction in size ca. 37 kyr (< 1000 birds). The population remained small for many generations, and ca. 1.5 kyr became structured (i.e., split) into RDC and RIR. The colonies remained connected after the population split, but exchanged few migrants (2Nm1 = 0.5, where 2Nm is the average number of haploid immigrants entering the population per generation). This event was followed by a population decline during the past 100 years (i.e., ≤ 4 generations) that reduced the RDC and RIR populations to less than 100 breeding pairs each, and resulted in the almost complete isolation of the colonies (2Nm0 = 0.01; and only from RDC to RIR) (Fig. 2b). Although the 95% confidence intervals (CI) for M7 reflect uncertainty in the parameter estimates (but see 95% CI for M3; Supplementary Table S4), all parameters inferred by the seven models fall within the same orders of magnitude and were consistent with (i) a very large ancestral population (≥ 2.6 × 105 birds); (ii) a population split during the last two millennia; (iii) a population bottleneck for RIR and RDC less than 100 years ago; (iv) and a very low contemporaneous population size (Supplementary Table S3).
Comparison of the seven demographic models tested with fastsimcoal2 for Pseudobulweria aterrima. Boxplots show the log10 Likelihood distributions computed based on the parameters that maximized the likelihood of each model in a total of 100 independent simulations per model. The models were ranked based on the Akaike Information Criteria (AIC). The values below the boxplots denote the ΔLhood (i.e., difference between the observed Likelihood and the maximum expected Likelihood) and ΔAIC (i.e., difference in AIC to the best demographic model) for each model. The best ranked demographic models are highlighted by an asterisk. Models M3 and M7 yielded a similar log10 Likelihood distribution and ΔAIC, suggesting a similar model fit. Analyses were performed using 9897 genome-wide SNPs called for 20 unrelated individuals (10 from each colony). M1 = null model; M2 = bottlenecks without gene flow; M3 = bottlenecks with gene flow; M4 = asymmetric bottleneck; M5 = ancient bottleneck; M6 = two ancient bottlenecks; M7 = ancient and recent bottlenecks; RIR = Rivière des Remparts; RDC = Rond Des Chevrons.
Discussion
In this study, we used a genome-wide SNPs dataset and two complementary modeling approaches to investigate how the past demographic events of the critically endangered Mascarene petrel were affected by natural vs. anthropogenic changes in Réunion Island. Our results reveal that (i) the known Mascarene petrel colonies exhibited a small population size and were isolated long before human colonization; (ii) historical population fluctuations coincided with significant volcanic events at Piton des Neiges and Piton de la Fournaise, unrelated to past climatic events; and (iii) the increasing human impact over the last century exacerbated the population bottlenecks further, driving the species perilously close to extinction. To our knowledge, this study provides the first long-term insights about the main drivers of biodiversity loss in an island endemic, and suggest that the current conservation status of endemic species may be a legacy of historical volcanism, exacerbated by recent human impact.
Fine-scale population structure and historical gene flow
The FST estimates and clustering analyses confirmed that Mascarene petrel colonies constitute two genetically differentiated populations. The physical isolation of breeding populations and philopatry are recognized as one of the main barriers to gene flow in seabirds40,54. The Mascarene petrel colonies are separated by only 19 km straight-line distance, and ecological barriers to gene flow are unlikely as there is no marked terrestrial ecological gradient between them. Fine-scale population genetic structure was also reported for the endemic Barau’s petrel, despite the studied colonies being only 5 km apart55, and in other procellariforms (e.g., Pelecanoides garnotii)49. As for Barau’s petrel, our findings suggest that the most likely mechanism promoting genetic isolation between the two colonies is natal philopatry (i.e., the behavior of returning to the natal colony to breed)56. Although this behavior is still not well understood, the familiarity with the breeding area might increase an individual’s probability of finding suitable nesting habitat and a mate. Dispersal to a new colony could reduce inbreeding and competition, but would bring additional risks such as the time cost of searching an alternative breeding site and the higher potential to encounter unforeseen threats (e.g., predators)56,57,58. It is plausible that philopatry evolved by favoring individuals that returned to their natal site, while those individuals that attempted to find a new breeding site experienced higher mortality and/or lower reproductive success56. Philopatry is often associated with other isolating mechanisms59. Morphometric analyses suggested that birds from RIR and RDC are phenotypically indistinguishable (data not shown), but differences in breeding phenology were observed (i.e., egg laying started in October in RIR and from November to December in RDC)42.
Despite the strong philopatric behavior of the Mascarene petrel, our results indicate historical genomic admixture among colonies and three lines of evidence supports this. First, the three best-ranked demographic models with fastsimcoal2 assumed that populations were more connected in the recent past than in present times. The comparison of the models M2 and M3, which only differ on the migration matrix, confirmed that the evolutionary history of the Mascarene petrel cannot be explained without historical gene flow. Indeed, all models tested with fastsimcoal2 agree that colonies were connected until relatively recently (< 2 kyr). Second, capture-mark-recapture data (n = 42 adult birds for RDC and 37 for RIR) confirmed that birds from a given colony have never been re-sighted as a breeder or prospector at the other colony during the last four and six breeding seasons for RDC and RIR, respectively60, therefore one could reject the hypothesis of recent gene flow. Third, the relatedness analyses confirm the capture-recapture data by showing that birds are not reproducing outside of their known colony (i.e., birds were related only to birds sampled at their own colony; Supplementary Text S1). The absence of first, second- or third-degree relatives outside their breeding colony confirms the absence of gene flow in recent generations.
Clustering methods also highlighted six individuals from RIR with an assignment probability to RDC ≥ 50%. It is plausible that birds moved from RDC to RIR, as the demographic analyses systematically suggested a larger Ne for RDC (= source population) in the recent past. Alternatively, migrants may be a legacy of birds that lost their breeding nests at the time of the ancestral population split, and subsequently moved to the nearest colony to breed (RIR; see below for details). Finally, clustering analyses revealed that all light-grounded birds originated from RDC. These results are not surprising as there is no significant light pollution in Rivière des Remparts canyon, except when fledglings reach the city of Saint-Joseph (Fig. 1a). In contrast, birds from RDC face light pollution while crossing the island towards the sea41. Over the last two decades, a yearly rescue program based on large-scale public awareness led by a local NGO (Society of Ornithological Studies of Réunion, SEOR), has successfully rescued and released 58 Mascarene petrels disoriented by artificial lights (1996–2021)38,41. While the rescue campaign stands out as the most impactful measure to mitigate light-induced mortality, complementary actions such as installing light shields, employing motion sensors to turn lights off, and implementing light restriction during the peak fledging period (recently implemented in Réunion Island) could also be effective in the conservation of the Mascarene petrel61. Altogether, our results suggest that birds from both colonies were part of a larger population that was connected until a recent past, and genetic differentiation of populations must have been a relatively quick evolutionary event, likely exacerbated by high natal philopatry.
Volcanic activity and recent anthropogenic impact on endemic seabird
The coalescent framework revealed two population declines of the Mascarene petrel pre-dating human impact on Réunion Island. According to the Stairway Plot, the Mascarene petrel reached its population maximum ca. 1 Mya and remained large until the Last Interglacial (LIG; 132–112 kyr). Given the recent origin of Réunion Island15, such a large population size may correspond to the spatial and demographic expansion of the species after its founding event on Réunion Island. Both methods implemented in our study identify a very large ancestral population, suggesting that favorable breeding habitat for the Mascarene petrel were abundant. The Mascarene petrel subsequently underwent a drastic population decline that started ca. 125 kyr. This date period does not coincide with any major climatic changes (Fig. 2c). Instead, it corresponds to a period of major eruptions of Piton des Neiges ca. 137–70 kyr62, with massive destructive impacts on terrestrial ecosystems (Fig. 2d). During this period, a large volcanic edifice (Piton des Neiges phase 4; PN4) appeared in the topographic paleo-depressions incised in the central part of Piton des Neiges by erosion during the preceding (ca. 40 kyr) period of eruption quiescence63. The successive volcanic eruptions fed lava flows that progressively accumulated over a thickness > 500 m in the paleo-topographic depressions. Lava flows are known to be particularly destructive at ground level, by burning all vegetation64. In Réunion Island, wildfire propagates in the valley scarps when the lava reaches their base, as observed during the August 2015 eruption of Piton de la Fournaise65. It is therefore likely that the flora and fauna on the scarps of the topographic paleo-depressions filled by the lava flows of PN4 were recurrently impacted by such wildfires and the emitted volcanic gas.
The parameters estimated by the fastsimcoal2 models M3 and M7revealed an identical demographic history for the last two millennia, but M7 allows us to delve further into the past and suggests a significant reduction in population size ca. 37 kyr, which parallels a previously documented population decline of Coffea mauritiana in Réunion Island29, and could be related to the last explosive activity of Piton des Neiges (Fig. 2d). Although confidence intervals reflect uncertainty in the timing of this event, a substantial Ne reduction was also suggested by the Stairway Plot. From ca. 40–27 kyr, Piton des Neiges experienced successive explosive eruptions that fed pyroclastic flows (fast moving fluidized mixture of hot lava blocks, ashes and volcanic gas) on the volcano flanks and the deep valleys17. Moreover, this eruptive period was characterized by a major eruption dated at 37.2 ± 1.5 kyr that spread a 30–40 cm thick fallout deposit over the flanks of Piton des Neiges and the western part of Piton de la Fournaise suggesting a massive island-scale impact on most components of the island terrestrial biodiversity66,67. Thus, the volcanic eruptions of Piton des Neiges may have triggered Mascarene petrel population declines, through a high mortality, breeding failures and, most importantly, the total disappearance of suitable breeding habitat11,13. Volcanic-related population declines and extinctions have also been reported for other birds’ species on other islands. For instance, the volcanic activity at Deception Island resulted in the near-complete local extinction of the gentoo penguin colony on Ardley Island (Antarctic Peninsula)12; the eruption of the Barcena volcano in San Benedicto (Revillagigedo Archipelago) resulted in the local extirpation of the Townsend’s shearwater68, and the recent violent eruption of Kasatochi volcano (central Aleutian archipelago) caused the local extinction of avian biodiversity13. After the latest eruption of Piton des Neiges abruptly ended, the Mascarene petrel appears not to have recovered to previous population levels. Two types of geological events may have limited population recovery. First, the current deep valleys cutting the southern flank of Piton des Neiges started to incise PN4 around 30 kyr69. In Réunion Island, the intense and ongoing erosion results from river incision and valley scarp collapses70,71. Such scarp collapse events could have limited the colony development. Second, the volcanic zone located between the summits of Piton des Neiges and Piton de la Fournaise experienced several eruptions during the past 30 kyr, feeding lava flows that cascaded down the Bras de la Plaine valley that currently hosts the RDC colony72,73. The two ancient population declines for the Mascarene petrel are corroborated by a previous study considering a dataset of 22 light-grounded birds and using microsatellite loci30.
Although the Stairway Plot method does not allow us to infer the population dynamics of the Mascarene petrel during the recent past, the fastsimcoal2 M7 suggests that during the last two millennia (ca. 1.5 kyr), the Mascarene petrel population was fragmented into two genetically distinct clusters found at the two known colonies (RIR and RDC), with reduced levels of post-split gene flow observed among them. This split follows one of the largest flank eruptions of Piton de La Fournaise ca. 1.9 kyr66 (Fig. 2d), starting on the upstream bank of Rivière des Remparts valley (Cratère Commerson). This eruption fed a large lava flow that entirely covered the valley floor down to the ocean, 21 km downstream. Similar to the PN4 period of activity of Piton de Neiges, this major volcanic event may have burned the vegetation on the cliffs hosting the Mascarene petrel nest sites. Consequently, many petrels may have been either fatally injured or displaced. Even if birds had survived this event, the philopatry behavior and nest-site fidelity of petrel species imply that breeders continually return to the same place every year74, even if breeding habitats are no longer available. For instance, the closely related Tahiti petrel (Pseudobulweria rostrata) nests in nickel-rich areas in New Caledonia. It was recently shown that after 10 years of nickel-mining exploration, Tahiti Petrel vocal activity was still recorded at the breeding sites, suggesting that former breeders whose nests sites were destroyed by mining activities were still visiting the area but failed to reproduce75. Likewise, it is plausible that many of the Mascarene petrels that were displaced by the volcanic event at ca. 1.9 kry BP did not move to another breeding site.
Once fragmented, both Mascarene petrel colonies underwent a synchronous population bottleneck during the last century, and the species was even subsequently considered extinct37. This population decline coincided with human-mediated changes in the landscape. Since human settlement ca. 400 years ago, human impact was rapid and substantial. Major bird predators were introduced shortly after human arrival (Rattus rattus in 1675; Rattus norvegicus in 1735, and cats in 1703)5,76. Extensive deforestation for logging, agriculture and cities started 200 years ago and resulted in the destruction of 70% of the native vegetation77. This had a significant impact through forest clearing and the spread of invasive mammals, the latter leading to increased egg and chick predation4,74. More recently, light pollution increased rapidly on the island, with the extension of urban and industrial areas (Fig. 2e), resulting in mortality of petrels and shearwaters, including the Mascarene Petrel36,41. These anthropogenic impacts are not unique to Réunion Island and human-related population collapses/extinctions are well-documented for Procellariform species, including the Hydrobates leucorhous on Grand Colombier Island4, the Pseudobulweria rupinarum on Saint Helena Island74, Puffinus olsoni on the Canary Islands78, and Puffinus boydi on Bermuda79. Current anthropogenic impacts may be compounded by the ongoing loss of nesting habitat via erosion, as illustrated by the recent collapse of Bras de Mahavel70.
Conservation status of Mascarene petrel is not a legacy of climatic changes
Quaternary climatic cycles and subsequent sea-level oscillations have been invoked as one of the major drivers of avian diversification26,53,80, including seabird species7,81,82,83, as they depend on marine ecosystems for foraging and on terrestrial habitats for reproduction. Our study suggests that the demographic dynamics of the Mascarene petrel do not follow the historical climatic events that took place in the WIO (Fig. 2c). Paleoenvironmental records available from Lake Tritrivakely in the Central Highlands of Madagascar84 and Last Interglacial corals on the Seychelles Islands85,86 confirmed that the LIG was characterized by warm temperatures and by higher-than-present sea level. This would be in line with a larger population size for the Mascarene petrel for this period, as its marine distribution seems to be related to high sea surface temperatures. However, the Stairway Plot analyses revealed a large decline ca. 125 kyr that could not be explained by climatic conditions alone, but coincided with volcanic activity of Piton des Neiges (see discussion above). There is multiple evidence that the Last Glacial Maximum (LGM; 26.5–19 kyr) was colder and drier across the WIO, with a substantial decrease in the annual mean temperature (ca. − 4 °C than in the present days), a reduction in rainfall (ca. − 10%), a reduction in sea-level (ca. − 125 m than today), and in the likely reduction of forest habitats84,87. Assuming that the Mascarene petrel population dynamics were influenced by climatic shifts, the population decline would be expected to coincide with the onset of the LGM, but both coalescent approaches agree that the decline pre-dated this period (ca. 125 and 37 kyr). The demographic analyses also suggest that Mascarene petrel never recovered its population size, not even during the African Humid Period (AHP; 15–5 kyr), a period characterized by an abrupt warming and increasing levels of moisture in continental Africa88,89, Madagascar9,90 and Rodrigues19, and that was followed by several megadroughts. The absence of population recovery could, instead, be explained by the persistent reduction in breeding areas for the Mascarene petrel and mortality following intense volcanic activity on Réunion Island, compounded by philopatry and mating behavior (i.e., monogamy) of the species. Altogether our study suggests volcanism as the major force driving the ancient population dynamics of the Mascarene petrel, but we emphasize that molecular chronologies are dependent on appropriate molecular rates8,9,26. The present study used the generation time of the Barau’s petrel for time calibration, but a different generation time estimate may result in older or more recent demographic events. We also highlight that the Stairway Plot method assumes that the observed data stems from a panmictic population43 and deviations from these model assumptions, such as the existence of migration among sub-populations, may bias demographic interpretations27,33,34,91. Moreover, it has also been demonstrated that Ne estimates are sensitive to the number of loci used in the analyses47,50, even though the general trend of population size change over time remains unaffected10. Consequently, the absolute Ne estimates were interpreted with caution.
One of the most striking results of our study is the lower population size and fragmentation prior to human impact on Réunion Island. Both coalescent approaches implemented in this study show that at the time of the first human settlement, the Mascarene petrel had an increased risk of extinction (Ne < 1000 birds), but the cumulative human-impact likely reduced the effective population size further. It is plausible that past volcanism also had catastrophic impacts on other endemic species, especially for those that are restricted to small ranges and/or distributed near the volcanic massifs, such as is the endemic Barau's petrel. Hence, multi-taxa studies are necessary to address the impact of volcanism on Réunion Island and better understand historic and current drivers of biodiversity loss. To conclude, our study suggests that historic volcanic activity may be the primary trigger of the current conservation status of endemic island biodiversity. The past dramatic population fluctuations combined with current anthropogenic impacts ultimately downward species population size to a minima, hampering its recovery from population lows and driving species towards extinction.
Materials and methods
Study species
The Mascarene petrel is the only representative of the Pseudobulweria genus in the Indian Ocean36,38,42. The species is endemic to Réunion Island and was thought to be extinct for more than eight decades, until it was rediscovered in the twentieth century with a dead grounded bird37. Nevertheless, the two breeding colonies investigated in this study (RDC and RIR; Fig. 1a) were only discovered in 2016–201738. The biology of Mascarene petrel remains poorly known as birds spend most of their life at sea and their breeding grounds are restricted to remote and inaccessible cliffs38. Currently, the population size is estimated to be as low as 100 breeding pairs42 and the petrel is listed as Critically Endangered (CR, IUCN 2020). Conservation actions focused on Mascarene petrel effectively started in the last two decades with the implementation of a rescuing program for light-grounded birds41 and predator control at breeding colonies38.
Study area and specimen collection
The mountainous Réunion Island is dominated by high, steep reliefs resulting from successive periods of intense volcanic activity and high erosion rates92. The highest relief reach 3071 m a.s.l. and 2631 m a.s.l. at Piton des Neiges’ and Piton de la Fournaise’s summits, respectively (Fig. 1a). The climate is characterized by a thermal gradient associated with elevation, with an annual mean temperature ranging from 25 °C on the coast to 10 °C at the summits. High mean annual rainfall is recorded on the windward east side of the island (ca. 11,000 mm), whereas the climate is markedly drier on the leeward west side (ca. 500 mm)77. Urban areas are mostly located on the coastal lowlands (< 1300 m a.s.l.), while the native forest is restricted to intermediary and high elevations6,29,77. Mascarene petrel breeding grounds are located in the upper valleys of large rivers, i.e., in the vicinity of past or dormant volcanic areas (Fig. 1a). The first site, RDC (− 21.16°N, 55.51°E), lies in the Bras de la Plaine valley (1250 m above sea level, a.s.l.), and covers an area of about 10,000 m2. The second site, RIR (− 21.29°N, 55.61°E), is located in the Rivière des Remparts valley (650 m a.s.l.) and covers a smaller area (~ 800 m2)38. All samples used in this study come from wild animals. Birds were captured by hand at their nesting burrows between 2016 and 2018. Blood samples (approx. 0.5 ml) were collected by venipuncture of the medial metatarsal or basilic veins. Bird capture, handling and sample collection were approved and carried out in concordance with the principles of the Research center on biology of bird populations (PP 609 and banding authorization 44 of MLC; CRPBO, National Museum of Natural History, Paris) and approved by Réunion Island National Park. All birds were released at their burrow after sampling. All sections of this manuscript followed the ARRIVE guidelines93 for reporting animal research. Blood samples of grounded birds (attracted by light pollution), were collected with the same method between 2008 and 2018 by the SEOR. All samples were stored in 70% ethanol until DNA extraction. Genomic DNA was extracted using the QIAmp Blood and Tissue kit (Qiagen). Collection information for all specimens is given in Supplementary Table S1.
SNP genotyping and filtering
SNP genotyping was carried out by Diversity Arrays Technology (DarT Pty Ltd, Canberra) using the DArTseq™ protocol, a restriction site-associated DNA sequencing method similar to double‐digest restriction‐associated DNA sequencing (ddRAD)24,94. A DArT library was prepared using 20–40 ng/μl of DNA and the restriction enzymes PstI and SphI, following the protocols described by Georges et al. (2018). Sequence data was processed using a proprietary DArT analytical pipeline95,96,97. In the absence of a conspecific reference genome, loci were aligned to the genome assembly of Calonectris borealis39 (family Procellariidae; GCA_013401115.1; Coverage depth = 130×)98. Given the high genomic synteny within birds, the overall genomic structure and gene order is expected to be very similar between members of the same family99. Such synteny has been used for SNP mapping in other seabird studies99,100. The raw SNP data were filtered in-house using the dartR v 2.1.4101 R package (R v4.2.1). Loci for which SNPs has been trimmed from the sequence tag along with the adaptor were filtered out. Genome-wide SNP datasets may contain large numbers of linked loci, which can break assumptions of independence for many analyses24. To control for short-distance linkage disequilibrium, we filtered out multiple-linked SNPs per sequenced tag (i.e., only one SNP per fragmented was retained at random). Knowing also that individuals with a low call rate (i.e., proportion of scored loci for an individual) and high heterozygosity may indicate bad DNA quality or cross-sample contamination102, individuals with more than 20% of missing data103 and/or with a very high individual heterozygosity (≥ 13% for Mascarene petrel; see Supplementary Fig. S7) were removed from our dataset. Any monomorphic loci (i.e., SNP fixed over all individuals) arising as a result of the removal of individuals were also deleted. Further filtering was undertaken for SNP with a call rate (i.e., proportion of scored individuals for a locus) lower than 90% and a genotyping reproducibility (i.e., average repeatability of alleles at a locus across replicates) below 95%. Loci with an average read depth (i.e., counts of sequence tags scored at a locus) lower than 10×25,29,45,46 and higher than 40×, and a minor allele frequency (MAF) lower than 0.01 were also removed104. The SNP dataset was then tested for the presence of sex-linked markers, knowing that if a SNP is present only on the Z chromosome but not on the W chromosome, all females (ZW) will be heterozygous at that locus and all males (ZZ) homozygous. Sex-linked markers were removed from the dataset. Finally, outlier SNPs were identified using two algorithms: PCAdapt105 and OutFLANK106, using a cut-off value of 0.01102. Significant outliers were then excluded as they may represent loci under selection. All thresholds were defined after plotting data 102,104. The filtered dataset contained a total of 13,855 SNP loci and 87 birds (dataset 1). After population clustering analyses, loci that exhibited departures from Hardy–Weinberg equilibrium (HWE) were also excluded (12,784 SNP loci; 87 birds; dataset 2). Information about the number of SNP loci and individuals retained at each step of the quality-filtering analyses are given in Supplementary Table S5.
Population genetic structure and relatedness analyses
The coalescent framework makes several assumptions about the population from which the samples are drawn22. Multiple studies have shown that population structure (i.e., non-random mating)32,33,34,35,107 and relatedness102 between individuals can confound genomic inferences. To minimize the violation of these assumptions, and to examine philopatric characteristics of the species, the population genetic structure of Mascarene petrel and relatedness between the 87 birds that passed the SNP quality filtering (dataset 1) were investigated first. The population structure was examined using three different approaches. First, a dimensionality-reduction clustering was performed with Discriminant Analysis of Principal Components (DAPC), using the Adegenet v2.1.7108 R package. DAPC summarizes the information present on the individuals’ genotypes using principal components analysis prior to a discriminant analysis to maximize differentiation between groups while minimizing variation within groups and makes no assumptions about HWE or linkage disequilibrium109. A K-means clustering algorithm was employed to identify the optimal number of clusters from K = 1 to K = 5. The optimal number of clusters (K) was determined by the lowest Bayesian Information Criterion (BIC) following109. To avoid overfitting of discriminant functions, a-score optimization was used to evaluate the optimal number of principal components to retain in the analysis45. Second, the Bayesian assignment approach implemented in Structure v2.3.4110 was used to assign individuals to a specific number of clusters (K). The number of genetic clusters ranged between 1 and 5, and a total of 10 independent runs were performed for each value of K to check the consistency of results. Analyses were performed assuming an admixture model, correlated allele frequencies, and without population priors, for 10,000 burn‐in and 100,000 replicate runs. Two criteria were used to determine the optimal number of genetic clusters: the log likelihood given K (L(K))110, and the second-order rate of change of mean log likelihood (ΔK) following the Evanno’s method111. Both L(K) and ΔK were calculated using Clumpak112. Third, genetic differentiation between the genetic clusters previously detected by the DAPC and Structure analyses was estimated using Wright’s F-statistics FST113, with and without the light-grounded birds. The analyses were performed with dartR v 2.1.4101 R package. Significance was tested using 10,000 bootstraps.
The genetic relationship between two individuals can be described by the concept of identity-by-descent (IBD; i.e., two alleles are identical if they share a recent common ancestry)114. To account for the potential presence of related individuals in our dataset, the software ngsRelateV2114 implemented in ANGSD framework115 was used to estimate the nine condensed Jacquard coefficients between all pairs of individuals116 (see Supplementary Text S2 for details). These coefficients provide a comprehensive description of the common ancestry between two individuals and were used to infer their familial relationship114,117. Only one individual out of a group of closely related individuals (e.g., parent–offspring or full siblings) was retained in dataset 3 (60 individuals; 12,731 SNPs). See Supplementary Text S1 and Fig. S8 for details about SNP datasets used for each analysis.
Mutation rate and generation time
Currently, few mutation rate and generation time estimates are available for Procellariiformes, hence demographic analyses for the Mascarene petrel were performed using the mutation rate of the Northern fulmar (Fulmarus glacialis; 2.89 × 10−9 substitutions per nucleotide per generation)53. This mutation rate is the only one currently available for Procellariiformes and has been used for similar analyses in other species from the same family, e.g., Puffinus mauretanicus118. Due to the recent rediscovery of the Mascarene petrel, some demographic parameters requiring long-term monitoring of marked birds (especially prebreeding survival rate and age at first breeding) are still missing, precluding the estimation of its generation time (GT). However, the equivalent parameters were available for the endemic Barau's petrel (Pterodroma baraui) and these were used to estimate GT (18.9 years; see Supplementary Text S3 for details about GT estimation for Barau's petrel) by the authors. As Mascarene and Barau's petrel are of similar size and both are exposed to similar environments and anthropogenic threats, the generation time estimated for Barau's petrel was used in this study.
Demographic history of Mascarene petrel
Stairway plot
The genome-wide distribution of allele frequencies of a given set of SNPs in a population (i.e., Site Frequency Spectrum; SFS) has been widely used as a summary statistic for demographic inferences119. The Stairway plot method is a model-free approach that makes use of the SFS generated from population genomic data to estimate a series of population mutation rates (θ = 4Neμ), following a multi-epoch demographic model where epochs coincide with coalescent events119. The genlight SNPs dataset without sites in HWE departure (dataset 2; 87 individuals; 12,784 SNPs) was initially converted to a VCF (Variant Call Format) file using the gl2vcf function implemented in dartR v 2.1.4101 R package. The VCF file was then used to estimate the 1d-SFS (i.e., the SFS for a single population) using the easySFS tool (https://github.com/isaacovercast/easySFS)119,120. Analyses were performed for each genetic cluster (i.e., RDC and light-grounded birds; and RIR) detected by the population structure analyses, to satisfy the assumption of population panmixia22. In the absence of a suitable outgroup to determine the ancestral state of each allele, we considered the minor allele-frequency spectrum (i.e., MAF or folded SFS) for all SFS-related analyses119. MAF selects the least frequent of the two alleles within the dataset and uses that information in the summary SFS. Since the SFS cannot be computed for sites containing missing data, the number of individuals per population was projected down to recover a higher number of loci120 (n = 45 for RDC and light-grounded birds; and n = 33 for RIR). The resulting 1d-SFS was used as input data to generate 199 bootstrap replicates to obtain 95% confidence intervals, using the script provided by Stairway plot v2.0119. Inferences were finally made based on 200 SFS with Stairway plot v2.0 0119. Since this method is sensitive to the number of coalescent events47,50,121, analyses were initially performed considering the entire datasets, and repeated with an equal sample size (n = 33; individuals randomly selected using R) to avoid differences in Ne among populations related to sample size. To evaluate the impact of relatedness on demographic inferences of Mascarene petrel, the Stairway Plot analyses were also repeated considering the dataset without closely-related individuals (n = 35 for RDC and light-grounded birds; and n = 19 for RIR after downsampling). Finally, the analyses were repeated for the entire dataset (n = 54 after downsampling and considering only unrelated individuals). Plots were scaled using 2.89 × 10−9 substitutions per nucleotide per generation53, and 18.9 years as generation time.
Demographic modeling with fastsimcoal2
The composite-likelihood approach implemented in fastsimcoal244 allows the inference of demographic parameters from the joint allele-frequency spectrum (JAFS) of populations using a coalescent simulation framework. A total of seven demographic models were tested for the Mascarene petrel, following a 3-step approach. First, four demographic models assuming size changes after the fragmentation of the ancestral population into the two known colonies, RDC and RIR, were tested. Specifically, the first and simplest model assumed a population split (with population resize) into RDC and RIR colonies (null model, M1). The second model tested the occurrence of a population decline for both colonies after population split and assumed that colonies have been isolated since then (bottlenecks without gene flow, M2). The third model also relied on the occurrence of a population decline for both colonies after population split, but assumed that colonies were exchanging migrants (i.e., 2Nm1 = 0.5, where 2Nm1 is the average number of haploid immigrants entering the population per generation). However, the decrease on Ne was followed by a reduction on population connectivity, with almost no gene flow (2Nm0 = 0.01, where 2Nm0 denoted recent gene flow) and migrants moving exclusively from RDC to RIR colony (= asymmetric gene flow; bottlenecks with gene flow, M3). The fourth model tested the occurrence of a population decline only for RIR, and a constant population size for RDC after the population split. The model also assumed connectivity changes, with the colonies becoming more isolated after the decline of RIR (2Nm1 = 0.5 and 2Nm0 = 0.01) (asymmetric bottleneck, M4). Second, two scenarios assuming size changes before the fragmentation of the ancestral population into RDC and RIR were tested. The fifth model (ancient bottleneck, M5) tested a single population decline before population split, while the sixth model (two ancient bottlenecks, M6) tested the occurrence of two successive declines before population split. Finally, a model including both size changes before and after the population split was built. The seventh model assumed the occurrence of an ancient population decline (i.e., before the population split) and a population bottleneck afterwards for both colonies. Similar to M3 and M4, the model implied connectivity changes, with the two colonies exchanging migrants after the population split (2Nm1 = 0.5), but becoming almost isolated after the declines (asymmetric gene flow; 2Nm0 = 0.01) (ancient & recent bottlenecks, M7). For the models assuming asymmetric gene flow (M3, M4 and M7), RDC was considered to be the source population as this colony covers a larger geographic area, exhibit a larger population size (as suggested by the Stairway Plot analyses for each genetic cluster separately), and also display lower levels of genetic admixture in comparison to RIR. See Supplementary Fig. S5 for an illustration of the seven tested demographic models, and Supplementary Table S6 for parameter tags definition and respective searching ranges. Since it was not computationally feasible to run fastsimcoal244 for the entire dataset, a total of 20 birds (i.e., 10 unrelated individuals from each colony; dataset 4) were selected for the analyses44,122,123 (see Supplementary Table S1 for list of individuals). The easySFS tool119,120 was used to estimate a 2d-SFS (where the two dimensions correspond to RIR and RDC colonies) from a VCF file. Fastsimcoal2 v.2.644 was run using 200,000 coalescent simulations per set of parameters and 40 ECM cycles during parameter estimation from the SFS. 100 independent runs were performed for each demographic model to determine the parameter estimates that maximize the composite-likelihoods44. The best-fitting demographic model was identified (i) on the basis of their Akaike’s information criterion (AIC)124; (ii) on the likelihood distributions obtained based on 100 expected SFS, as an overlap among the distributions between models would indicate no significant difference between the fit of alternative models as differences may be attributed to the variance in the SFS approximation46; and (iii) on the visual inspection of the fit between the expected and observed 2d-SFS119. The nonparametric block-bootstrap approach implemented in46 was used to generate 40 bootstrap replicates. Bootstrap replicates were obtained by dividing the SNPs into 100 blocks, and sampling with replacement 100 blocks for each replicate, to match the original dataset size46. A total of 40 independent fastsimcoal2 runs were performed for each replicate under the two models with the lowest AIC. The parameter estimates with the highest likelihood from each independent run were used to estimate the 95% CI with the Rmisc R package125. The most realistic model was finally scaled considering a generation time of 18.9 years.
Data availability
The raw DArTseq genotypes generated by Diversity Arrays Technology (DarT Pty Ltd, Canberra) and analyzed during the current study, the associated specimen metadata, and the genomic datasets (dataset 1–4) used for all the analyses are available on DRYAD (https://doi.org/https://doi.org/10.5061/dryad.s1rn8pkfb). Scripts are available from the corresponding author upon request.
References
Vences, M., Wollenberg, K. C., Vieites, D. R. & Lees, D. C. Madagascar as a model region of species diversification. Trends Ecol. Evol. 24, 456–465 (2009).
Kier, G. et al. A global assessment of endemism and species richness across island and mainland regions. Proc. Natl. Acad. Sci. 106, 9322–9327 (2009).
Russell, J. C. & Kueffer, C. Island Biodiversity in the Anthropocene. Annu. Rev. Environ. Resour. 44, 31–60 (2019).
Duda, M. P. et al. Linking 19th century European settlement to the disruption of a seabird’s natural population dynamics. Proc. Natl. Acad. Sci. USA 117, 32484–32492 (2020).
Mourer-Chauviré, C., Bour, R. & Ribes, S. Recent avian extinctions on Reunion (Mascarene Islands) from the paleontological and historical sources. Bull. Br. Ornithol. Club 126, 40 (2006).
Thébaud, C., Warren, B. H., Strasberg, D. & Cheke, A. Mascarene islands, biology. Encycl. Islands https://doi.org/10.1525/9780520943728-146 (2020).
Trucchi, E. et al. King penguin demography since the last glaciation inferred from genome-wide data. Proc. R. Soc. B Biol. Sci. 281, 20140528 (2014).
Salmona, J., Heller, R., Quéméré, E. & Chikhi, L. Climate change and human colonization triggered habitat loss and fragmentation in Madagascar. Mol. Ecol. 26, 5203–5222 (2017).
Teixeira, H. et al. Past environmental changes affected lemur population dynamics prior to human impact in Madagascar. Commun. Biol. 4, 1084 (2021).
Tiley, G. P. et al. Population genomic structure in Goodman’s mouse lemur reveals long-standing separation of Madagascar’s Central Highlands and eastern rainforests. Mol. Ecol. 31, 4901–4918 (2022).
Finkelstein, M. E. et al. The anatomy of a (potential) disaster: Volcanoes, behavior, and population viability of the short-tailed albatross (Phoebastria albatrus). Biol. Conserv. 143, 321–331 (2010).
Roberts, S. J. et al. Past penguin colony responses to explosive volcanism on the Antarctic Peninsula. Nat. Commun. 8, 14914 (2017).
Sonsthagen, S. A. et al. Legacy or colonization? Posteruption establishment of peregrine falcons (Falco peregrinus) on a volcanically active subarctic island. Ecol. Evol. 7, 107–114 (2017).
Valente, L. M., Etienne, R. S. & Phillimore, A. B. The effects of island ontogeny on species diversity and phylogeny. Proc. R. Soc. B Biol. Sci. 281, 20133227 (2014).
Quidelleur, X., Holt, J. W., Salvany, T. & Bouquerel, H. New K-Ar ages from La Montagne massif, Réunion Island (Indian Ocean), supporting two geomagnetic events in the time period 2.2–2.0 Ma. Geophys. J. Int. 182, 699–710 (2010).
Warren, B. H., Strasberg, D., Bruggemann, J. H., Prys-Jones, R. P. & Thébaud, C. Why does the biota of the Madagascar region have such a strong Asiatic flavour?. Cladistics 26, 526–538 (2010).
Famin, V. et al. Multitechnique geochronology of intrusive and explosive activity on Piton des Neiges Volcano, Réunion Island. Geochem. Geophys. Geosyst. 23, e2021GC010214 (2022).
Maillard, L. Notes sur l’île de la Réunion (Bourbon) vol. 392 (Dentu, 1863).
Li, H. et al. A multimillennial climatic context for the megafaunal extinctions in Madagascar and Mascarene Islands. Sci. Adv. 6, eabb2459 (2020).
Tofanelli, S., Bertoncini, S. & Donati, G. Early human colonization, climate change and megafaunal extinction in madagascar: the contribution of genetics in a framework of reciprocal causations. Front. Ecol. Evol. 9, 1–5 (2022).
Beichman, A. C., Huerta-Sanchez, E. & Lohmueller, K. E. Using genomic data to infer historic population dynamics of nonmodel organisms. Annu. Rev. Ecol. Evol. Syst. 49, 433–456 (2018).
Ho, S. Y. W. & Shapiro, B. Skyline-plot methods for estimating demographic history from nucleotide sequences. Mol. Ecol. Resour. 11, 423–434 (2011).
Mather, N., Traves, S. M. & Ho, S. Y. W. A practical introduction to sequentially Markovian coalescent methods for estimating demographic history from genomic data. Ecol. Evol. 10, 579–589 (2020).
Dorey, J. B. et al. Holocene population expansion of a tropical bee coincides with early human colonization of Fiji rather than climate change. Mol. Ecol. 30, 4005–4022 (2021).
Nadachowska-Brzyska, K., Burri, R., Smeds, L. & Ellegren, H. PSMC analysis of effective population sizes in molecular ecology and its application to black-and-white Ficedula flycatchers. Mol. Ecol. 25, 1058–1072 (2016).
Pujolar, J. M., Dalén, L., Hansen, M. M. & Madsen, J. Demographic inference from whole-genome and RAD sequencing data suggests alternating human impacts on goose populations since the last ice age. Mol. Ecol. 26, 6270–6283 (2017).
Teixeira, H. et al. Impact of model assumptions on demographic inferences: the case study of two sympatric mouse lemurs in northwestern Madagascar. BMC Ecol. Evol. 21, 197 (2021).
Salmona, J., Heller, R., Lascoux, M. & Shafer, A. Inferring demographic history using genomic data. In: Population Genomics 511–537 (Springer, 2017). https://doi.org/10.1007/13836_20
Garot, E., Joët, T., Combes, M. C., Severac, D. & Lashermes, P. Plant population dynamics on oceanic islands during the Late Quaternary climate changes: genetic evidence from a tree species (Coffea mauritiana) in Reunion Island. New Phytol. 224, 974–986 (2019).
Lopez, J. et al. High genetic diversity despite drastic bottleneck in a critically endangered, long-lived seabird, the Mascarene Petrel Pseudobulweria aterrima. Ibis (Lond. 1859) 163, 268–273 (2021).
Salmona, J. et al. Signature of a pre-human population decline in the critically endangered Reunion Island endemic forest bird Coracina newtoni. PLOS ONE 7, e43524 (2012).
Chikhi, L., Sousa, V. C., Luisi, P., Goossens, B. & Beaumont, M. A. The confounding effects of population structure, genetic diversity and the sampling scheme on the detection and quantification of population size changes. Genetics 186, 983–995 (2010).
Heller, R., Chikhi, L. & Siegismund, H. R. The confounding effect of population structure on Bayesian skyline plot inferences of demographic history. PLOS ONE 8, e62992 (2013).
Mazet, O., Rodríguez, W., Grusea, S., Boitard, S. & Chikhi, L. On the importance of being structured: Instantaneous coalescence rates and human evolution-lessons for ancestral population size inference?. Heredity (Edinb). 116, 362–371 (2016).
Orozco-terWengel, P. The devil is in the details: the effect of population structure on demographic inference. Nat. Publ. Gr. https://doi.org/10.1038/hdy.2016.9 (2016).
Le Corre, M., Ollivier, A., Ribes, S. & Jouventin, P. Light-induced mortality of petrels: a 4-year study from Réunion Island (Indian Ocean). Biol. Conserv. 105, 93–102 (2002).
Attie, C., Stahl, J. & Bretagnolle, V. New data on the endangered Mascarene petrel Pseudobulweria aterrima: a third twentieth century specimen and distribution. Waterbird Soc. 20, 406–412 (1997).
Juhasz, C.-C. et al. Discovery of the breeding colonies of a critically endangered and elusive seabird, the Mascarene Petrel (Pseudobulweria aterrima). J. F. Ornithol. 93, 11 (2022).
Gangloff, B. et al. The complete phylogeny of Pseudobulweria, the most endangered seabird genus: systematics, species status and conservation implications. Conserv. Genet. 13, 39–52 (2012).
Lombal, A. J., O’dwyer, J. E., Friesen, V., Woehler, E. J. & Burridge, C. P. Identifying mechanisms of genetic differentiation among populations in vagile species: historical factors dominate genetic differentiation in seabirds. Biol. Rev. 95, 625–651 (2020).
Chevillon, L. et al. 25 years of light-induced petrel groundings in Reunion Island: retrospective analysis and predicted trends. Glob. Ecol. Conserv. 38, e02232 (2022).
Virion, M. C. Plan National d′ Actions en faveur des Petrels endemiques de La Reunion 2020–2029 (2021)
Liu, X. & Fu, Y.-X. Exploring population size changes using SNP frequency spectra. Nat. Genet. 47, 555–559 (2015).
Excoffier, L., Dupanloup, I., Huerta-Sánchez, E., Sousa, V. C. & Foll, M. Robust demographic inference from genomic and SNP data. PLoS Genet. 9, 1003905 (2013).
Bagley, R. K., Sousa, V. C., Niemiller, M. L. & Linnen, C. R. History, geography and host use shape genomewide patterns of genetic variation in the redheaded pine sawfly (Neodiprion lecontei). Mol. Ecol. 26, 1022–1044 (2017).
Meier, J. I. et al. Demographic modelling with whole-genome data reveals parallel origin of similar Pundamilia cichlid species after hybridization. Mol. Ecol. 26, 123–141. https://doi.org/10.1111/mec.13838 (2017).
Lapierre, M., Lambert, A. & Achaz, G. Accuracy of demographic inferences from the site frequency spectrum: The case of the yoruba population. Genetics 206, 139–449 (2017).
Terhorst, J., Kamm, J. A. & Song, Y. S. Robust and scalable inference of population history from hundreds of unphased whole genomes. Nat. Genet. 49, 303–309 (2017).
Cristofari, R. et al. Climate-driven range shifts of the king penguin in a fragmented ecosystem. Nat. Clim. Chang. 8, 245–251 (2018).
Patton, A. H. et al. Contemporary demographic reconstruction methods are robust to genome assembly quality: A case study in tasmanian devils. Mol. Biol. Evol. 36, 2906–2921 (2019).
Lougnon, A. Sous le signe de la tortue: voyages anciens à l’Île Bourbon, 1611–1725 (5. éd.). Sainte-Clotilde, Réunion: Orphie (2006).
Sandron, F. Dynamique de la population réunionnaise (1663–2030). La population réunionnaise—Analyse démographique. IRD Editions, Paris (France) (2007).
Nadachowska-Brzyska, K., Li, C., Smeds, L., Zhang, G. & Ellegren, H. Temporal dynamics of avian populations during pleistocene revealed by whole-genome sequences. Curr. Biol. 25, 1375–1380 (2015).
Friesen, V. L., Burg, T. M. & McCoy, K. D. Mechanisms of population differentiation in seabirds: Invited review. Mol. Ecol. 16, 1765–1785 (2007).
Danckwerts, D. K., Humeau, L., Pinet, P., McQuaid, C. D. & Le Corre, M. Extreme philopatry and genetic diversification at unprecedented scales in a seabird. Sci. Rep. 11, 1–12 (2021).
Pärt, T. Male philopatry confers mating advantage in migratory collared flycatchers. Anim. Behav. 48, 401–409 (1994).
Antaky, C. C., Young, L., Ringma, J. & Price, M. R. Dispersal under the seabird paradox: Probability, life history, or spatial attributes?. Mar. Ornithol. 49, 1–8 (2021).
Coulson, J. C. A review of philopatry in seabirds and comparisons with other waterbird species. J. Waterbird Soc. 39, 229–326 (2016).
Clucas, G. V. et al. Comparative population genomics reveals key barriers to dispersal in Southern Ocean penguins. Mol. Ecol. 27, 4680–4697 (2018).
Saunier, M. et al. Integrating genomic and capture–recapture data to uncover population dynamics of a critically endangered seabird. in prep.
Raine, H., Borg, J. J., Raine, A., Bairner, S. & Cardona, M. B. Light pollution and its effect on Yelkouan Shearwaters in Malta; causes and solutions. BirdLife Malta, Malta (2007).
Salvany, T., Lahitte, P., Nativel, P. & Gillot, P.-Y. Geomorphic evolution of the Piton des Neiges volcano (Réunion Island, Indian Ocean): competition between volcanic construction and erosion since 1.4 Ma. Geomorphology 136, 132–147 (2012).
Gayer, E., Michon, L. & Villeneuve, N. Volcanic island multi-stage construction inferred from a simple geometrical approach: example of Réunion Island. Geomorphology 392, 107900 (2021).
Dale, V. H. et al. Effects of modern volcanic eruptions on vegetation. Volcanoes and the Environment (Cambridge University Press, Cambridge, 2005).
Harris, A. J. L. et al. Effusive crises at Piton de la Fournaise 2014–2015: a review of a multi-national response model. J. Appl. Volcanol. 6, 11 (2017).
Bachèlery, P. & Chevallier, L. Carte volcano-tectonique du Piton de la Fournaise. Doc. Inst. Phys. du Globe Paris (1982).
Dosseto, A., Hannan-Joyner, A., Raines, E., Gayer, E. & Michon, L. Geochemical evolution of soils on Reunion Island. Geochim. Cosmochim. Acta 318, 263–278 (2022).
Martínez-Gómez, J. E. & Jacobsen, J. K. The conservation status of Townsend’s shearwater Puffinus auricularis auricularis. Biol. Conserv. 116, 35–47 (2004).
Sisavath, E. et al. Processes controlling a volcaniclastic turbiditic system during the last climatic cycle: Example of the Cilaos deep-sea fan, offshore La Réunion Island. Sediment. Geol. 281, 180–193 (2012).
Michon, L., Gayer, E., Lucas, A., Bellin, F. & Gougeon, M. The 1965 Mahavel Landslide (Réunion Island, Indian Ocean): Morphology, volumes, flow dynamics, and causes of a rock avalanche in tropical setting. J. Geophys. Res. Earth Surf. 128, e2022JF006944. https://doi.org/10.1029/2022JF006944 (2023).
Bret, L., Fevre, Y., Join, J. L., Robineau, B. & Bachelery, P. Deposits related to degradation processes on Piton des Neiges Volcano (Reunion Island): Overview and geological hazard. J. Volcanol. Geotherm. Res. 123, 25–41 (2003).
Michon, L., Ferrazzini, V., Di Muro, A., Villeneuve, N. & Famin, V. Rift zones and magma plumbing system of Piton de la Fournaise volcano: How do they differ from Hawaii and Etna?. J. Volcanol. Geotherm. Res. 303, 112–129 (2015).
Morandi, A. et al. Pre-historic (<5 kiloyear) explosive activity at Piton de la Fournaise Volcano. In Active Volcanoes of the Southwest Indian Ocean 107–138 (Springer, Berlin 2016).
Welch, A. J., Olson, S. L. & Fleischer, R. C. Phylogenetic relationships of the extinct St Helena petrel, Pterodroma rupinarum Olson, 1975 (Procellariiformes: Procellariidae), based on ancient DNA. Zool. J. Linn. Soc. 170, 494–505 (2014).
Pagenaud, A. et al. Tahiti petrel Pseudobulweria rostrata population decline at a nickel-mining site: A critical need for adapted conservation strategies. Bird Conserv. Int. 32, 246–258 (2022).
Cheke, A. S. Extinct birds of the Mascarenes and Seychelles: A review of the causes of extinction in the light of an important new publication on extinct birds. Phelsuma 21, 4–19 (2013).
Strasberg, D. et al. An assessment of habitat diversity and transformation on La Réunion Island (Mascarene Islands, Indian Ocean) as a basis for identifying broad-scale conservation priorities. Biodivers. Conserv. 14, 3015–3032 (2005).
Rando, J. C. & Alcover, J. A. Evidence for a second western Palaearctic seabird extinction during the last millennium: The Lava Shearwater Puffinus olsoni. Ibis 150, 188–192 (2008).
Olson, S. L. Taxonomic review of the fossil Procellariidae (Aves: Procellariiformes) described from Bermuda by RW Shufeldt. Proc. Biol. Soc. Washingt. (2004).
Kozma, R., Melsted, P., Magnússon, K. P. & Höglund, J. Looking into the past: The reaction of three grouse species to climate change over the last million years using whole genome sequences. Mol. Ecol. 25, 570–580 (2016).
Carrea, C. et al. High vagility facilitates population persistence and expansion prior to the Last Glacial Maximum in an antarctic top predator: The snow petrel (Pagodroma nivea). J. Biogeogr. 46, 442–453 (2019).
Cole, T. L. et al. Receding ice drove parallel expansions in Southern Ocean penguins. Proc. Natl. Acad. Sci. USA 116, 26690–26696 (2019).
Ferrer Obiol, J. et al. Integrating sequence capture and restriction site-associated DNA sequencing to resolve recent radiations of pelagic seabirds. Syst. Biol. 70, 976–996 (2021).
Gasse, F. & Van Campo, E. Late quaternary environmental changes from a pollen and diatom record in the southern tropics (Lake Tritrivakely, Madagascar). Palaeogeogr. Palaeoclimatol. Palaeoecol. 167, 287–308 (2001).
Israelson, C. & Wohlfarth, B. Timing of the last-interglacial high sea level on the Seychelles Islands. Indian Ocean. Quat. Res. 51, 306–316 (1999).
Dutton, A. & Lambeck, K. Ice volume and sea level during the last interglacial. Science 337, 216–219 (2012).
Gasse, F. & Van Campo, E. A 40 000-yr pollen and diatom record from Lake Tritrivakely, Madagascar, in the southern tropics. Quat. Res. 49, 299–311 (1998).
Demenocal, P. et al. Abrupt onset and termination of the African Humid Period: rapid climate responses to gradual insolation forcing. Quat. Sci. Rev. 19, 347–361 (2000).
Tierney, J. E. & DeMenocal, P. B. Abrupt shifts in Horn of Africa hydroclimate since the last glacial maximum. Science 342, 843–846 (2013).
Wang, L. et al. The African humid period, rapid climate change events, the timing of human colonization, and megafaunal extinctions in Madagascar during the Holocene: Evidence from a 2m Anjohibe Cave stalagmite. Quat. Sci. Rev. 210, 136–153 (2019).
Mazet, O., Rodríguez, W. & Chikhi, L. Demographic inference using genetic data from a single individual: separating population size variation from population structure. Theor. Popul. Biol. 104, 46–58 (2015).
Gayer, E., Michon, L., Louvat, P. & Gaillardet, J. Storm-induced precipitation variability control of long-term erosion. Earth Planet. Sci. Lett. 517, 61–70 (2019).
du Sert, N. P. et al. Reporting animal research: Explanation and elaboration for the arrive guidelines 2.0. PLoS Biol. 18, 3000411 (2020).
Georges, A. et al. Genomewide SNP markers breathe new life into phylogeography and species delimitation for the problematic short-necked turtles (Chelidae: Emydura) of eastern Australia. Mol. Ecol. 27, 5195–5213 (2018).
Sansaloni, C. et al. Diversity Arrays Technology (DArT) and next-generation sequencing combined: genome-wide, high throughput, highly informative genotyping for molecular breeding of Eucalyptus. BMC Proc. 5, 1–2 (2011).
Kilian, A. et al. Diversity arrays technology: A generic genome profiling technology on open platforms. Data Prod. Anal. Popul. Genomics Methods Protoc 67–89 (2012).
Jaccoud, D., Peng, K., Feinstein, D. & Kilian, A. Diversity arrays: a solid state technology for sequence information independent genotyping. Nucleic Acids Res. 29, 25 (2001).
Feng, S. et al. Dense sampling of bird diversity increases power of comparative genomics. Nature 587, 252–257 (2020).
Rexer-Huber, K. et al. Genomics detects population structure within and between ocean basins in a circumpolar seabird: The white-chinned petrel. Mol. Ecol. 28, 4552–4572 (2019).
Galla, S. J. et al. Reference genomes from distantly related species can be used for discovery of single nucleotide polymorphisms to inform conservation management. Genes 10, 9 (2019).
Mijangos, J. L., Gruber, B., Berry, O., Pacioni, C. & Georges, A. dartR v2: An accessible genetic analysis platform for conservation, ecology and agriculture. Methods Ecol. Evol. 13, 2150–2158 (2022).
Devloo-Delva, F. et al. Accounting for kin sampling reveals genetic connectivity in Tasmanian and New Zealand school sharks Galeorhinus galeus. Ecol. Evol. 9, 4465–4472 (2019).
Ewart, K. M. et al. Museum specimens provide reliable SNP data for population genomic analysis of a widely distributed but threatened cockatoo species. Mol. Ecol. Resour. 19, 1578–1592 (2019).
Feutry, P. et al. One panel to rule them all: DArTcap genotyping for population structure, historical demography, and kinship analyses, and its application to a threatened shark. Mol. Ecol. Resour. 20, 1470–1485 (2020).
Luu, K., Bazin, E. & Blum, M. G. B. pcadapt: An R package to perform genome scans for selection based on principal component analysis. Mol. Ecol. Resour. 17, 67–77 (2017).
Whitlock, M. C., Lotterhos, K. E. & Bronstein, J. L. Reliable detection of loci responsible for local adaptation: Inference of a null model through trimming the distribution of FST. Am. Nat. 186, S24–S36 (2015).
Parreira, B., Quéméré, E., Vanpé, C., Carvalho, I. & Chikhi, L. Genetic consequences of social structure in the golden-crowned sifaka. Heredity https://doi.org/10.1038/s41437-020-0345-5 (2020).
Jombart, T. & Ahmed, I. adegenet 1.3–1: New tools for the analysis of genome-wide SNP data. Bioinformatics 27, 3070–3071 (2011).
Jombart, T., Devillard, S. & Balloux, F. Discriminant analysis of principal components: A new method for the analysis of genetically structured populations. BMC Genet. 11, 94 (2010).
Pritchard, J. K., Stephens, M. & Donnelly, P. Inference of population structure using multilocus genotype data. Genetics 155, 945–959 (2000).
Evanno, G., Regnaut, S. & Goudet, J. Detecting the number of clusters of individuals using the software STRUCTURE: A simulation study. Mol. Ecol. 14, 2611–2620 (2005).
Kopelman, N. M., Mayzel, J., Jakobsson, M., Rosenberg, N. A. & Mayrose, I. Clumpak: A program for identifying clustering modes and packaging population structure inferences across K. Mol. Ecol. Resour. 15, 1179–1191 (2015).
Weir, B. S. & Cockerham, C. C. Estimating F-statistics for the analysis of population structure. Oxf. Univ. Press 38, 1358–1370 (1984).
Hanghøj, K., Moltke, I., Andersen, P. A., Manica, A. & Korneliussen, T. S. Fast and accurate relatedness estimation from high-throughput sequencing data in the presence of inbreeding. Gigascience 8, 1–9 (2019).
Korneliussen, T. S., Albrechtsen, A. & Nielsen, R. ANGSD: Analysis of next generation sequencing data. BMC Bioinform. 15, 1–13 (2014).
Jacquard, A. The genetic structure of populations Vol. 5 (Springer Science & Business Media, Berlin, 2012).
Blouin, M. S. DNA-based methods for pedigree reconstruction and kinship analysis in natural populations. Trends Ecol. Evol. 18, 503–511 (2003).
Cuevas-Caballé, C. et al. The first genome of the balearic shearwater (Puffinus mauretanicus) provides a valuable resource for conservation genomics and sheds light on adaptation to a pelagic lifestyle. Genome Biol. Evol. 14, 1–14 (2022).
Gutenkunst, R. N., Hernandez, R. D., Williamson, S. H. & Bustamante, C. D. Inferring the joint demographic history of multiple populations from multidimensional SNP frequency data. PLOS Genet. 5, e1000695 (2009).
Coffman, A. J., Hsieh, P. H., Gravel, S. & Gutenkunst, R. N. Computationally efficient composite likelihood statistics for demographic inference. Mol. Biol. Evol. 33, 591–593 (2015).
Foote, A. D. et al. Genome-culture coevolution promotes rapid divergence of killer whale ecotypes. Nat. Commun. 7, 11693 (2016).
Beichman, A. C., Phung, T. N. & Lohmueller, K. E. Comparison of single genome and allele frequency data reveals discordant demographic histories. G Genes Genomes Genet 7, 3605–3620 (2017).
Excoffier, L. & Foll, M. fastsimcoal: A continuous-time coalescent simulator of genomic diversity under arbitrarily complex evolutionary scenarios. Bioinformatics 27, 1332–1334 (2011).
Akaike, H. A new look at the statistical model identification. IEEE Trans. Automat. Contr. 19, 716–723 (1974).
Harrell, F. E. Jr. Package ‘hmisc’. CRAN2018 2019, 235–236 (2019).
Acknowledgements
We thank the Parc national de La Réunion for the permission to conduct the field work at RIR and RDC colonies. Field work was only possible thanks to the assistance and support of Jerome Dubos, Yahaia Soulaimana-Mattoir, Julie Tourmetz, Martin Riethmuller and Patxi Souharce. We also thank everyone who has participated in the SEOR rescue campaign of Mascarene petrel. We thank Jade Lopez for performing preliminary analyses and Félix Pellerin for his help with computational issues. We are very grateful to Andrej Killian and the Diversity Arrays Technology platform for library preparation and sequencing. The authors are very grateful to the genotoul bioinformatics platform Toulouse Occitanie (Bioinfo Genotoul, https://doi.org/10.15454/1.5572369328961167E12) for providing help, computing and storage resources for the genomic analyses. This study was supported by FEDER Smac (2020-2022, N° RE0022954), and the LIFE + Petrels (grant number: LIFE13 BIO/FR/000075) co-led by Le Parc National de La Réunion, l’Université de La Réunion, SEOR and l’Office National de la Chasse et de la Faune Sauvage, with financial support from the European Union, La Direction de l’Environnement l’Aménagement et du Logement (DEAL) and Le Conseil Départemental de La Réunion. The study was additionally funded by the Bertarelli Foundation as part of the Bertarelli Programme in Marine Science (Project 822916).
Author information
Authors and Affiliations
Contributions
H.T. and L.H. designed and conceived the study. M.L.C., P.P. and F.X.C. collected the samples for genomic analyses. L.H. carried out the laboratory work. H.T. processed the DArTseq data and performed the genomic analyses, under the supervision of L.H. N.N. provided the pipeline for the SNPs filtering. Results were discussed with M.N., M.L.C, A.J., L.M and L.H., who contributed to the intellectual direction of the study. H.T. wrote the first draft of the manuscript. All authors read and approved the final manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Teixeira, H., Le Corre, M., Michon, L. et al. Past volcanic activity predisposes an endemic threatened seabird to negative anthropogenic impacts. Sci Rep 14, 1960 (2024). https://doi.org/10.1038/s41598-024-52556-9
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-024-52556-9
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.