Abstract
During the first steps of HIV infection the Env subunit gp41 is thought to establish contact between the membranes and to be the main driver of fusion. Here we investigated in liquid crystalline membranes the structure and cholesterol recognition of constructs made of a gp41 external region carrying a cholesterol recognition amino acid consensus (CRAC) motif and a hydrophobic membrane anchoring sequence. CD- und ATR-FTIR spectroscopies indicate that the constructs adopt a high degree of helical secondary structure in membrane environments. Furthermore, 15N and 2H solid-state NMR spectra of gp41 polypeptides reconstituted into uniaxially oriented bilayers agree with the CRAC domain being an extension of the transmembrane helix. Upon addition of cholesterol the CRAC NMR spectra remain largely unaffected when being associated with the native gp41 transmembrane sequence but its topology changes when anchored in the membrane by a hydrophobic model sequence. The 2H solid-state NMR spectra of deuterated cholesterol are indicative of a stronger influence of the model sequence on this lipid when compared to the native gp41 sequence. These observations are suggestive of a strong coupling between the transmembrane and the membrane proximal region of gp41 possibly enforced by oligomerization of the transmembrane helical region.
Similar content being viewed by others
Introduction
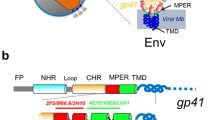
Despite decades of intense research HIV remains a challenging infection that requires life-long treatment with expensive medicines and for which no effective vaccination has become available. During the HIV life cycle the viral and the target cell membranes have to fuse, a process mediated by the viral envelope gycoprotein Env1,2. The envelope protein is a heterodimeric complex of gp120 and gp41, these being the cleavage products of gp160. The gp41 protein forms a trimeric transmembrane unit and three gp120 proteins are exposed at the viral surface3,4,5. Interactions of gp120 with the cell surface receptor CD46 and one of the chemokine receptors CCR5 or CXCR47,8,9 initiates a series of conformational rearrangements during which gp120 is shed, thereby exposing gp41 which adopts an extended pre-fusion state2,10. The primary structure of gp41 is schematically illustrated in Fig. 1.
Schematic illustration of gp41: FP fusion peptide, FPPR fusion peptide proximal region, NHR and CHR N-terminal and C-terminal heptad repeat, respectively, MPER membrane proximal external region including the Cholesterol recognition amino acid consensus (CRAC) motif, TM transmembrane anchor, CT cytoplasmic terminus. The numbering follows the HXB2 subtype of HIV-1 (uniprot P04578).
In the extended prefusion state the hydrophobic amino-terminal fusion peptide anchors in the host cell membrane while the gp41 transmembrane domain, which is 172 amino acid residues further downstream, remains in the viral envelope10,11. The presence of cholesterol in the viral and cellular membranes has been found important during membrane fusion, viral infection and the late phase of the viral cycle being a structural determinant and modulator of fusion and post-fusion events12,13,14.
Cholesterol is abundant in mammalian cells where is it is synthesized in the endoplasmatic reticulum, transported through the Golgi and accumulates at high concentrations in the plasma membrane. Cholesterol is also found in the blood serum in the form of lipoprotein complexes from where it exchanges with the cellular reservoirs (e.g. recent reviews15,16). Cholesterol has profound effects on the physico-chemical properties of lipid membranes where it orders lipids, increases the hydrophobic thickness of membranes, smoothens phase transitions, induces lateral lipid phase separations and membrane curvature strain1. In the presence of cholesterol, the formation of liquid-ordered (Lo) and liquid-disordered phases (Ld) has been observed, which are rich and poor in cholesterol, respectively17,18,19. More recent investigations revealed that the Lo/Ld phase boundaries that occur when cholesterol-rich and cholesterol-poor membrane domains oppose each other is a key element for the functioning of gp41 fusion peptides20,21. Fusion works best when such boundaries occur in both the viral and the cellular membranes22.
Cholesterol has also been shown to interact with proteins either by changing the physico-chemical properties of the membrane or by direct interactions with amino acid side chains1,23,24,25. Thereby, in several high-resolution structures, cholesterol has been found to interact with the protein revealing a number of recurring contacts sites25. First, Ala, Leu and Val commonly interact with the isooctyl tail of cholesterol. Second, the sterol ring interacts with hydrophobic residues where the C18 and C19 methyl groups protrude out of the cholesterol ring and serve as knobs to fit holes or grooves on the protein surfaces predominantly involving branched residues such as Leu and Ile. Third, aromatic residues interact with the saturated parts of the sterol ring by van der Waals and CH–π interactions. Furthermore, π–π interactions of Phe and Tyr with the C5=C6 double bond have been observed. Fourth, the 3β-OH head group of cholesterol takes part in hydrogen bonding networks with water and polar amino acids in particular in loops and turns at the membrane interface25.
Indeed, combinations of the above mentioned amino acids have been found in motifs such as the ‘cholesterol recognition amino acid consensus’ (CRAC) domain (-L/V-X1-5-Y-X1-5-R/K-), the reversed CRAC motif (CARC), the cholesterol consensus motif (CCM), or the sterol sensing domain (SSD)1,23,24,25. However, it should be noted that statistically such combinations of amino acids occur quite frequently and, in most cases do not involve interactions with cholesterol23,25. Furthermore, the cholesterol consensus motif (CCM) has been identified in GPCRs and consists of amino acid arrangements involving several TM helical domains26 whereas the role of the sterol sensing domain (SSD) has been questioned27.
The membrane proximal external region (MPER) of gp41 is highly conserved among HIV and SIV strains28. The MPER, the fusion peptide and the heptad repeat sequences (Fig. 1) are all considered essential during the fusion of viral and cellular membranes by destabilizing the bilayer packing during fusion1,29,30,31,32. A CRAC motif with the sequence LWYIK1,23,24,25,33 is part of the membrane proximal region. Mutagenesis of CRAC resulted in a reduced infection rate and a slowing of the viral life cycle34. Notably, epitopes for the 2F5, 4E10, 10E8 and LN01 broadly neutralizing antibodies against HIV include locations of the MPER and knowing more about its structure bears considerable promise for the rational design of an AIDS vaccine35,36,37,38,39,40.
Whereas most studies are indicative of helical conformations of MPER in membrane environments considerable discrepancies exist about the details of how the domain interacts with lipid bilayers or how the associated transmembrane domain is organized41,42,43,44. For example, while the structure of the TMD has been evaluated from NMR data in DMPC/DHPC bicelles assuming a trimeric structure45,46, a more extensive analysis using NMR relaxation data, RDCs, PRE, EPR and ultracentrifugation data reveals that in such environments the TMD is a monomeric continuous helix extending into the MPER43. When a construct including a more extended MPER domain has been investigated the MPER amino-termini of three subunits form a hydrophobic core44. As a result the N- and C-terminal helices of MPER are connected by a 90° turn while the C-terminal helix adopts a kinked arrangement relative to the TMD. In yet another X-ray structure MPER has been found to form a continuous helix with the C-terminal heptad repeat sequence47. Notably, in the presence of 2F5 antibodies β-turn structures have also been observed for the MPER48,49.
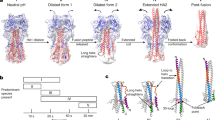
The fusion events that follow the initial contacts of gp120 with the cellular receptors occur via transient structural rearrangements that remain largely speculative and result in a post-fusion structure where the fusion peptide and the transmembrane domain, although far removed in the primary sequence, are in close physical proximity50. In order to bring the membranes into contact the extended prefusion arrangement has to transform to the compact postfusion bundle of helical hairpins made of six helices in antiparallel arrangement11,47,51. The exposed pre-hairpin conformation is of particular interest because it is accessible to the humoral immune response during a relatively long lag phase of HIV infection52,53 and has become the target of antiviral fusion inhibitor drugs and vaccines54,55,56.
In order to understand the mechanism of viral entry into the host cells it is important to understand in molecular detail the structural changes of gp41 during receptor recognition, membrane fusion and cellular entry. Although high-resolution structural investigations promise to be excellent starting points to develop broadly neutralizing vaccines the conformational and topological details of the MPER domain remain mysterious. Structural data and propositions include an amphipathic helix localized in the membrane interface at an alignment parallel to the surface29, a helix-kink-helix structure where the helices are at about 45° relative to the membrane normal44 or a continuous extension of the transmembrane helix43. Notably, the most recent data from a plethora of MAS solid-state NMR distance and water accessibility measurements were obtained from bilayer samples and seem in accordance with a predictive theoretical model29 where a trimeric helix-kink-helix structure is obtained for a MPER-TMD construct57. When the data are used as constraints in evaluating structural templates from analogous viral fusion proteins the CRAC sequence constitutes part of the kinked non-helical region57. The diverging data suggest that the MPER region exhibits a high degree of conformational plasticity which may be of importance to optimize its interactions with membranes during the different stages of the viral life cycle.
Therefore, in this paper we have investigated polypeptide sequences encompassing the MPER connected to transmembrane helical membrane anchors and their interactions with cholesterol in liquid crystalline phospholipid bilayers by CD, FTIR and solid-state NMR spectroscopies. Whereas CD and FTIR spectroscopies reveal the secondary structure composition of polypeptides, solid-state NMR of polypeptides that are labelled with 15N and 2H at selected sites are sensitive indicators of interactions within membranes and provide direct information about the alignment of the labelled sites relative to the membrane normal58. In particular, we tested the structural and topological arrangement of the MPER, its structural coupling with the transmembrane helical anchor and if interactions of cholesterol with the CRAC domain of gp41 result in structural and/or topological changes.
Materials and methods
Organic solvents were from Sigma-Aldrich (St Quentin Fallavier France) and of 99% purity. Cholesterol, POPC and POPS were from Avanti Polar Lipids (Birmingham, AL).
Peptide synthesis
The peptides were prepared by solid-phase peptide synthesis using the standard cycles of a Millipore 9050 automatic peptide synthesizer (Millipore, Darmstadt, Germany) and Fmoc chemistry. Fmoc-protected amino acids from NovaBiochem (Merck Millipore, Darmstadt, Germany) were used at four-fold excess, the Tentagel-R-RAM resin was from Rapp Polymere (Tübingen, Germany). Cleavage of the peptide was performed in 28 mL trifluoroacetic acid (TFA), 1.5 mL water and 0.3 mL triethylsilane for 4 h. After solubilization and ether precipitation the TFA counter ions were exchanged in 4% acetic acid. After purification by semipreparative HPLC using an acetonitrile/water gradient on a C18 reverse-phase column (Luna 5u c18(2) 100 Å; 150 × 30 mm, Phenomenex, Le Pecq, France) the identity of the products was confirmed by MALDI TOF mass spectrometry.
Preparation of samples for CD spectroscopy
The CRAC-TM_model polypeptide was dissolved in chloroform/methanol 2/1 v/v and mixed with a solution of lipid in the same solvent to obtain a peptide-to-lipid ratio of 1/100. CRAC-TM_gp41 was dissolved in trifluoro ethanol (TFE)/water 85/15 v/v and homogenously mixed with lipid in TFE/chloroform 1/1 v/v. The solvent was evaporated under a stream of nitrogen and in high vacuum overnight. The lipid film was then hydrated in 10 mM phosphate buffer, pH 7 to obtain a peptide concentration of 25 µM. Small unilamellar vesicles were prepared by tip sonication (Bandelin Sonoplus HD 200, Bandelin, Berlin, Germany) in an ice bath. Aggregates and titanium debris from the sonicator tip were removed by centrifugation at 10,000g for 5 min at room temperature.
CD spectra
CD spectra were recorded on a JASCO-810 spectrometer (Jasco, Tokyo, Japan) at 20 °C between 190 and 250 nm with a band width of 1 nm. 4 scans of adaptive integration time between 1 and 8 s were averaged. A spectrum of buffer was subtracted. The spectra were analyzed for secondary structure composition of the peptides using CDPRO and the CONTIN/LL algorithm with reference set SMP5659,60.
ATR FTIR measurements
For ATR FTIR measurements CRAC-TM_model samples were prepared by dissolving 1 mg of peptide in chloroform/methanol 2/1 v/v. Lipids (POPC, Cholesterol) in the same solvent were added to reach a peptide-to-lipid (P/L) molar ratio of 2%. A few µL of sample solution in 10 mM sodium acetate buffer prepared in D2O were deposited on the germanium crystal of a Nicolet 6700 ATR-FTIR spectrometer (Thermo Scientific, Waltham, MA). Typically, five ATR spectra, each of 128 scans, were recorded between 1000 and 1800 cm−1 with a resolution of 4 cm−1. The spectra were normalized to the amide I absorption band near 1655 cm−1 to facilitate their comparison and slightly smoothed by means of a binomial filter numerical procedure. In order to obtain the relative contribution of each secondary structure element, the deconvolution of the amide I signal is necessary (Origin 8.5 software).
Preparation of supported lipid bilayers
The peptides were dissolved and mixed with lipids in the same solvents as used above (CD samples) to obtain a peptide-to-lipid ratio of 2 mol%. The solvents were evaporated under a stream of nitrogen until a viscous solution was obtained (~ 200 μL). The latter was applied to 22 ultrathin glass plates (8 × 22 mm or 6 × 11 mm, thickness 00, Marienfeld, Lauda-Königshofen, Germany) and dried first in air, then in high vacuum overnight. The glass plates were equilibrated in 93% relative humidity at ambient temperature, stacked on top of each other and sealed in plastic wrapping (illustrated protocol in61).
Solid-state NMR spectroscopy
The samples were inserted into a flat-coil NMR probe62 with the normal parallel to the magnetic field direction of the NMR spectrometer.
Proton-decoupled 15N solid-state NMR spectra were recorded at 40.54 MHz on a Bruker Avance 400 solid-state NMR spectrometer (Bruker Biospin, Rheinstetten, Germany) at room temperature. A cross polarization (CP) pulse sequence63 was used with the following parameters: spectral width 37.5 kHz, acquisition time 3.5 ms, CP contact time 0.8 ms, recycle delay 3 s, B1 fields during CP 42 kHz. Typically, 40 k scans were accumulated. An exponential multiplication with line broadening of 100 Hz was applied before Fourier transformation. The spectra were calibrated relative to external ammonium chloride (40 ppm)64. The precision of the chemical shift determination depends on the line width and the signal-to-noise of the corresponding spectra, therefore, several samples have been recorded repeatedly. The data were analyzed by taking into consideration the angular dependence of the chemical shift σzz which is σzz = σ11 sin2Θ cos2Φ + σ22 sin2Θ sin2Φ + σ33 cos2Θ. In this formula σ11, σ22 and σ33 are the main tensor elements64, and the Euler angles Θ and Φ position the chemical shift tensor relative to the magnetic field direction of the NMR spectrometer65.
2H solid-state NMR spectra were recorded at 61.4 MHz on a Bruker Avance 400 solid-state NMR spectrometer (Bruker Biospin, Rheinstetten, Germany) at room temperature. A quadrupolar echo pulse sequence was used66 with a spectral window, an acquisition time, echo delay, and a recycle delay of 125 kHz, 6.8 ms and 1.5 s, respectively. The echo delay was twice 40–50 μs. Typically, 40 k scans were accumulated. An exponential multiplication with line broadening of 300 Hz was applied before Fourier transformation. D2O was used as an external reference.
Results
Peptide design
Domains considered important during the viral life cycle were prepared by solid-phase peptide synthesis and HPLC purification in order to investigate their membrane interactions. During the synthesis 15N and 2H isotopic labels were included into the sequences for solid-state NMR structural investigations. The sequences thus prepared and investigated are listed in Table 1. The CRAC sequence of gp41 was anchored at the membrane interface by its covalent linkage to a transmembrane anchor. The membrane anchor is either the wild type sequence of gp41 which has been shown to adopt helical conformations in previous investigations43,44,45 (noted: TM_gp41), or a helical hydrophobic model peptide made of alanines and leucines similar to those investigated in previous studies67,68,69 (TM_model). As for the CRAC sequence, the native LWYIK was incorporated and its interactions compared to those of a scrambled sequence (scr) which does not follow the published amino acid consensus motif33. Thereby it is possible to distinguish if chemical shift changes upon addition of cholesterol are due to peptide-lipid interactions or rather reflect alterations in the physico-chemical properties of the membrane. Furthermore, an I/L variant was investigated which also fits the CRAC motif (model_2). Finally, the position of the 15N label was varied to test if the topological findings at Leu 8 are reproduced at Leu 11 thereby reflecting the CRAC domain as a whole (15N-L8 or 15N-I11).
Secondary structure in lipid bilayers by CD- and FTIR spectroscopies
In a next step the secondary structure of the polypeptides in membrane environments was investigated by CD spectroscopy. The hydrophobic anchoring sequence and the lipid reconstitution protocol starting from a homogenous solution of peptides and lipids in organic solvent assure that virtually all of the polypeptide is associated with the liposomes. The polypeptides were designed in such a manner to adopt a high degree of helical conformation in membranes. Thereby, the spectroscopic investigations also provide a first test if the peptide design and the membrane reconstitution protocol worked as expected.
The CD spectra of CRAC-TM_model or of CRAC-TM_gp41 exhibit the typical features of helical polypeptides when investigated in POPC/POPS 3/1 mol/mole membranes, i.e. a positive maximum at 195 nm and two negative intensities at 208 nm and 222 nm (Fig. 2). Indeed, line fitting analysis using the Dichroweb software indicates helical contributions > 50%59,60. At higher cholesterol concentrations the spectra flatten and a more quantitative analysis has to be considered with caution because of light scattering artifacts in the presence of vesicles of a size approaching the wave length of the incoming light70. Therefore, the fit results shown in Fig. 2C should be considered semiquantitative.
CD spectra of (A) CRAC-TM_model and (B) CRAC-TM_gp41 after reconstitution into POPC/POPS 3/1 mol/mole SUVs with increasing concentrations of cholesterol at a peptide-to-lipid ratio of 1 mol%. The peptide concentration was 25 µM in 10 mM phosphate buffer, pH 7, at 20 °C. (C) Fit results of the CD spectra of CRAC-TM_model (closed symbols) and of CRAC-TM_gp41 (open symbols) using the CONTIN/LL algorithm and the base set SMP56 of CDPRO. Comparison of the experimental spectra and the fits are shown in Figure S1.
Because peptide oligomerization/aggregation and vesicle agglutination can have a pronounced effect on the CD spectra, when at the same time they depend on all of lipid composition, preparation protocol, exact P/L ratio and lipid concentration, the secondary structure preferences were further investigated by ATR-FTIR spectroscopy where a lipid suspension deposited on a solid support is investigated. In contrast to CD spectroscopy light diffraction artefacts due to the agglutination and/or fusion of vesicles are avoided. The ATR-FTIR spectra of CRAC-TM_model reconstituted in POPC or POPC/cholesterol 70/30 mol/mole lipid bilayers exhibit a single resonance with a peak position characteristic of helical conformations (Fig. 3A)71. In contrast CRAC-TM_gp41 shows additional intensities > 1665 cm−1 characteristic for β-turn conformations (Fig. 3B).
Solid-state NMR investigations of the CRAC domain in oriented lipid bilayers
Previous structural studies have come to different conclusions with regard to the membrane interactions of the MPER which includes the CRAC domain57. The data are suggestive of an orientation along the surface29, a helix-kink-helix structure where the helices are at about 45° relative to the membrane normal44 or a continuous extension of the transmembrane helix43. Whereas such previous structures were calculated from a large number of short range NMR parameters static oriented solid-state NMR spectroscopy is a complementary approach based on bond orientations relative to the membrane derived from the anisotropy of chemical shift, dipolar and quadrupolar interactions58,65.
Therefore, in order in investigate in more detail the interactions of the CRAC-TMD polypeptides with lipid bilayers, and in particular to get direct insight into the alignment of the CRAC sequence relative to the membrane surface, the peptides were prepared carrying isotope labels at specific sites, reconstituted into supported liquid crystalline phospholipid bilayers and solid-state NMR spectra were recorded. When inserted into the solid-state NMR probe with the sample normal parallel to the magnetic field direction the resulting 15N chemical shift of the peptide amides correlates in a direct manner with the alignment of the amide 15N-1H vector and thereby the helical tilt angle65. Whereas 15N–1H bonds that are oriented parallel to the membrane normal (i.e. transmembrane helices) exhibit 15N chemical shifts around 200 ppm those that are oriented along the surface resonate at < 100 ppm. Finally, unstructured domains either aggregate and show broad powder pattern line shapes (50–230 ppm) or isotropic peak positions (ca 120 ppm)72.
In a first set of experiments the CRAC-TM_model sequence labelled at the Leu8 position, i.e. within the CRAC motif, was investigated (Fig. 4, Table 2). The spectra show well-oriented 15N solid-state NMR spectra in the 200 ppm region which are typically observed for transmembrane helical sequences. In POPC the chemical shift is 197 ppm which changes to 208 ppm upon addition of 30 mol% cholesterol (Fig. 4A,D). Closely related spectra are obtained for CRAC-TM_model_2 which only differs in a single I-L substitution (Tables 1 and 2, Figure S2). When the CRAC sequence is scrambled the chemical shift is about 210 ppm in the absence or presence of cholesterol (Fig. 4B,E, Table 2). For all three peptides, in the presence of cholesterol the peaks become broader by about two-fold suggesting that in the cholesterol containing membrane motional averaging is less dynamic1.
Proton-decoupled 15N solid-state NMR spectra of [15N-Leu8]-CRAC-TM_model (A,D) [15N-Leu8]-CRAC-TM_model-scr with a scrambled amino acid arrangement of the CRAC region (B,E) and of [15N-Leu8]-CRAC-TM_gp41 (C,F). The polypeptides were reconstituted into uniaxially aligned POPC lipid bilayers without (A–C) and with 30 mol% cholesterol (D–F) at a peptide-to-lipid ratio of 2 mol%. The membrane normal was parallel to the magnetic field of the NMR spectrometer and the experiments were performed at room temperature.
In a second series of experiments the CRAC-TM_gp41 sequence was labelled at the Leu8 or the Ile11 position, reconstituted into oriented membranes and investigated in the presence of 0, 2, 10 or 30 mol% cholesterol (Fig. 4C,F, Table 3). Irrespective of the cholesterol content, 15N chemical shifts around 198 ppm were observed for Leu8, and at about 215 ppm for Ile11. Although the chemical shift position did not change significantly with the addition of cholesterol the line width at half-height increased from 15 to 40 ppm for Leu8 (Fig. 4C,F, Table 3) and from 7 to 14 ppm for Ile11 (Table 3).
Solid-state NMR investigations of the TM domain in oriented lipid bilayers
Whereas 15N solid-state NMR investigations provide data on the membrane alignment and interactions of the CRAC domain (Fig. 4, Tables 2 and 3) the membrane interactions of the TMD are analyzed by recording 2H solid-state NMR spectra from the same samples. For these experiments 2H3-alanine was incorporated at position 24 of the CRAC-TM_model or position 29 of the CRAC-TM_gp41 sequences. The deuterium labels thus represent the situation within the transmembrane anchor. Previous NMR investigations indicate a stable helical conformation of the gp41 membrane anchor when studied in bicellar environments43,44,45. Furthermore, hydrophobic model sequences that are closely related to the one investigated here and carrying isotopic labels at comparable positions exhibit solid-state NMR spectra typical of stable helical structures in liquid crystalline bilayers67,68,69.
Notably, in POPC membranes well-oriented 2H solid-state NMR spectra are observed for all four sequences i.e. the CRAC-TM_model carrying the native CRAC, the scrambled CRAC (CRAC-TM_model_scr) or the I11L substitution CRAC as well as the CRAC-TM_gp41 sequence (Fig. 5A–D, Table 4). It should be noted that relatively narrow angular distributions of only 5° or 10° can result in considerable line broadening of the 2H solid-state NMR spectra recorded from deuterated alanines68,73. In contrast the 15N spectra of backbone labelled transmembrane helices are much less sensitive to angular variations65. Because the 2H quadrupolar splitting is so sensitive to even small orientational changes58,68 this indicates that in POPC the TMDs exhibit unique membrane alignments with very little mosaicity.
2H solid-state NMR spectra of 2 mol% [3,3,3-2H3-Ala24]-CRAC-TM_model (A,E), [3,3,3-2H3-Ala24]-CRAC-TM_model2 with a single I to L substitution within the CRAC domain (B,F), [3,3,3-2H3-Ala24]-CRAC-TM_model with a scrambled CRAC amino acid arrangement of the CRAC domain (C,G) and of [3,3,3-2H3-Ala29]-CRAC-TM_gp41 reconstituted into uniaxially aligned POPC lipid bilayers (D,H) without (A–D) and with cholesterol (E–H). The membrane normal was parallel to the magnetic field of the NMR spectrometer and the experiments were performed at room temperature. The central contributions exhibiting small quadrupolar splittings and isotropic resonances are from residual HDO74.
When 30% cholesterol is added to the POPC bilayers the spectra of the [3,3,3-2H3-Ala24]-model sequence broaden considerably (Fig. 5E–G) whereas the gp41 transmembrane domain remains largely unaffected (Fig. 5H, Table 4). The broad distribution of quadrupolar splittings covering ≤ 35 kHz is indicative of an increased range of conformations and/or topologies that result in Cα–Cβ orientations in slow (Fig. 5E,G) or intermediate exchange (Fig. 5F) on a time scale of 10–5 s.
In contrast, a single alignment persists for the gp41 TMD with little topological heterogeneity (Fig. 5H). In the presence of cholesterol, the quadrupolar splitting of [3,3,3-2H3-Ala29]-CRAC-TM_gp41 changes by 5 kHz (Table 4) which is indicative of a change in Cα–Cβ orientation by a few degrees and/or a decrease in motional averaging.
Solid-state NMR spectra of deuterated cholesterol
Whereas in the experiments just presented the effects of cholesterol on the structure and alignment of the CRAC-TM sequences were tested (Figs. 4 and 5, Tables 2, 3 and 4) a complementary view on these interactions is obtained from NMR spectra of the surrounding lipids. Therefore, in the next series of experiments, the 2H solid-state NMR spectra of deuterated cholesterol were recorded in order to monitor its interactions with the CRAC-TMD sequences in liquid crystalline bilayers. Cholesterol was deuterated at either six sites at the 2, 3, 4 and 6 positions, which are expected to locate in the interfacial region in mixed phospholipid/cholesterol membranes, or at seven sites of carbons 25, 26 and 27 which reside in the hydrophobic membrane interior (Fig. 6E). The 2H solid-state NMR spectrum of a pure lipid membrane made of POPC/cholesterol-d6 90/10 mol/mole is shown in Fig. 6A and compared to the spectrum obtained in the presence of 2 mol% CRAC-TM_ gp41 (Fig. 6B). Furthermore, Fig. 6C,D show the 2H solid-state NMR spectra of POPC/cholesterol-d7 90/10 mol/mole in the absence or presence of this polypeptide, respectively.
Deuterium solid-state NMR spectra of [2,2,3,4,4,6]-2H6-cholesterol (A,B) or [25, 263, 273]-2H7-cholesterol (C,D) in the absence (A,C) or presence (B,D) of 2 mol% CRAC-TM_gp41 in POPC/cholesterol 90/10 mol/mole membranes. (E) Structure of cholesterol where the deuterated sites are numbered. (F) The deuterium quadrupolar splittings extracted from the corresponding 2H solid-state NMR spectra as a function of temperature in the absence (solid lines, closed symbols) or presence of 1 mol% CRAC-TM_model (hatched line, open symbols). Positions 2 and 4 carry two deuterons each with different alignments (axial and equatorial relative to the ring system) and therefore different quadrupolar splittings.
In the presence of 2% CRAC-TM_gp41 a small increase in the quadrupolar splittings is observed for membranes enriched with 10% cholesterol-d6 (positions 2,3,4,5,6) when investigated at ambient temperatures (Fig. 6A,B). The same changes are observed in the presence of CRAC-TM_gp41_scr carrying a scrambled CRAC motif (not shown) suggesting that these alterations are unrelated to specific interactions with the cholesterol recognition motif.
Furthermore, a small decrease of the splittings assigned to the CD label at position 25 as well as of the CD3 groups at positions 26 and 27 occurs in the presence of 2% CRAC-TM_gp41 (Fig. 6C,D). Interestingly, the splittings of the methyl groups not only decrease but also split up into two distinguishable contributions upon addition of CRAC-TM_gp41 (Fig. 6C,D) suggesting a decrease in molecular order parameter and/or conformational changes of the cholesterol alkyl chain.
When the interactions of CRAC-TM_gp41with cholesterol were investigated neither the 15N chemical shifts of the peptide (Figs. 4C,F, Table3) nor the 2H spectra of deuterated cholesterol (Fig. 6A–D) showed any changes that would indicate a strong and specific interaction between the CRAC motif and the lipid. Because a small 15N chemical shift change due to the presence of cholesterol was observed for the CRAC motif when associated with the transmembrane model sequence (CRAC-TM_model; Fig. 4A,D), we performed additional measurements by reconstituting this peptide into membranes carrying deuterated cholesterol (Figs. 6F and S3).
The temperature dependent decrease of the quadrupolar splittings of the various sites in the presence of CRAC-TM_model is shown in Fig. 6F where the assignment has been taken from previous publications75,76. The comparison shows that the quadrupolar splittings of the 2H-labels at the 2, 3 and 4 positions decreases by about 4% due to the presence of CRAC-TM_model, while the deuteron at the 25 position does not show any difference due to the incorporation of this polypeptide (Figs. 6F and S3). Because the C2H3-signals of the 26 and 27 positions exhibit a small splitting they potentially overlap with residual HDO resonances and have not been analyzed further.
Discussion
Because in previous structural investigations ambiguous data have been obtained about the membrane alignment and interactions of the CRAC sequence (reviewed in reference57), here we have used oriented solid-state NMR to obtain angular information on several labelled sites in the presence and absence of cholesterol. In contrast to the distance information obtained by solution and MAS solid-state NMR from which the membrane topology of protein domains is deduced in an indirect manner29,43,44,57 solid-state NMR spectroscopy on supported lipid bilayers works in a complementary way by directly measuring angular information of peptide chemical bonds relative to the membrane normal58,65.
The CRAC motif of gp41 was positioned at the membrane interface via transmembrane helical anchor sequences, and its structural response to cholesterol was tested using CD- and FTIR- as well as oriented solid-state NMR spectroscopies. The mutual influence of the transmembrane helical domain and the MPER was investigated by replacing the native gp41 sequence by a simple hydrophobic model anchor. Furthermore, control experiments consisted in a scrambled CRAC sequence, a variant of the CRAC motif and sequences that carried the isotopic labels at different positions (Table 1).
Secondary structure and membrane topology of CRAC-TM
In a first series of experiments the constructs were studied for secondary structure preferences. As expected from the experimental design CD- and FTIR spectra were dominated by helical contributions in the presence or absence of cholesterol (Figs. 2 and 3). Interestingly, the FTIR technique reveals some β-turn contribution for the polypeptides anchored with the native gp41 TMD (Fig. 3B). Helical preferences in the presence of DPC micelles were also found when peptides presenting the pre-transmembrane region of gp41 were investigated by solution NMR-, CD- and infra-red spectroscopies29 while a 13C solid-state NMR chemical shift analysis revealed a tendency for the carboxy-terminal part of the gp41 TMD to adopt some β-sheet character in complex cholesterol-containing membranes mimicking the HIV envelope57.
When the CRAC motif LWYIK was labelled with 15N at the Leu or the Ile position and the resulting CRAC-TMD constructs incorporated into uniaxially oriented POPC lipid bilayers the 15N spectra exhibit well oriented line shapes with chemical shifts around 200 ppm typically observed for helical domains that are aligned with the helix axis parallel to the membrane normal (Figs. 4 and 7A)65,77,78. Whereas such spectra have often been observed for hydrophobic transmembrane regions the CRAC sequence is part of the MPER. Therefore, the spectra are in line with a continuous helical arrangement as has been observed for a related gp41 sequence in bicellar environments by solution state NMR43. A continuous CRAC-TMD helical conformation has also be observed in the MAS solid-state NMR structure of TSPO79. Homology modelling with the crystal structures of bacterial counterparts indicated an ≈ 7° change in tilt angle of two helices upon interaction with cholesterol79. While the CRAC domain itself tends to stick into the water phase (Fig. 7A) comparison with other structures are suggestive that this configuration is not exclusive and can be affected by interaction contributions from upstream residues that pull the membrane external residues closer to the membrane29,57 (Fig. 7B) or stabilize a helix-kink-helix structure involving the CRAC motif44.
Sketches the conformational flexibility of gp41 where the TMD shown in green and the MPER/CRAC regions shown in yellow exhibit predominantly helical structures. The protein appears to exhibit considerable conformational flexibility. Whereas the data shown in this paper indicate that both helical domains are aligned in the same direction, possibly forming a continuous structure (A), helix-kink-helix conformations have also been observed (B). Possibly, the two conformers shown represent different states of the membrane fusion process (see text for details).
Differential effects of cholesterol on CRAC-TM structure and topology
When the effect of adding 30% cholesterol to CRAC-TM_model reconstituted into supported POPC lipid bilayers is investigated the 15N chemical shift of the labelled Leu increases from 197 to 208 ppm (Table 2), which corresponds to a topological change in tilt/pitch angles by a few degrees65. Notably, the change was absent in presence of the scrambled CRAC sequence (Table 2). Thereby the spectral alterations point to an interaction between the CRAC motif and cholesterol, albeit when CRAC is attached to TM_gp41 such changes are absent (Table 3). Interestingly, when sub-nanometer proximities between cholesterol and the somewhat more extended MPER-TM_gp41 were tested in a recent MAS solid-state NMR investigation it turned out that cholesterol is close to two hot spots made up residues 673–677 and 684–688 but not in proximity of residues 679–683 which make up the CRAC motif80. In combination with the results of this paper this data suggests a weak interaction of cholesterol with the CRAC motif which can be overruled in a competitive manner by interactions with other domains. In this context it is of interest that residues 683–689 of TM_gp41 cover a CARC sequence80.
To gain further insight into the polypeptide topology the transmembrane domains were labelled with 2H3 at one of the alanine methyl groups and investigated by 2H solid-state NMR. This approach has previously been demonstrated to be highly sensitive to even small changes of the relative alignment of the alanine Cα–Cβ bond with respect to the membrane normal58. The addition of cholesterol has only a small effect on the wt TM_gp41 whereas a range of Cα–Cβ alignments is observed for the TM_model sequence (Fig. 5, Table 4). The broadening of the 2H NMR signal in case of TM_model can reflect conformational heterogeneity and/or many different topologies in slow exchange. It is possible that TM_model scans different alignments also in pure POPC bilayers but that in this lipid environment exchange is fast thus only an average orientation becomes apparent on the time scale of the 2H solid-state NMR spectra (10–5 s). In contrast, such motions are slowed sufficiently to make them distinguishable in the presence of cholesterol.
When the inverse experiment is performed where unlabeled peptide is added to deuterated cholesterol only in the presence of CRAC-TM_ model a significant reduction in the deuterium quadrupolar splittings of the interfacial cholesterol deuterons was observed whereas CRAC-TM_gp41 or the scrambled CRAC motif did not have an effect (Fig. 6). Thus, interactions between CRAC-TM_gp41 and cholesterol do not result in significant structural changes of either interaction partner (Tables 3 and 4, Fig. 6).
Interestingly, SDS gel electrophoresis is indicative of higher molecular weight oligomers formed by CRAC-TM_gp41 when compared to CRAC-TM_model (Figure S4). Therefore, the observed differences could be related to the association of the native TMD into oligomers (probably trimers)57 which is less likely for the model sequence. Thereby, within the oligomeric assembly of the wild-type sequence structural and topological changes of the CRAC or TM domains are energetically too costly to be triggered by the relatively weak interactions with cholesterol57. Indeed, the gp41 TMD carries a GxxxG motif (amino acids 19–23 of CRAC-TM_gp41) as well as an arginine at position 25, both being known to drive helix-helix association within the hydrophobic environment of the membrane interior81,82,83,84.
Possible cholesterol recognition sites within gp41
Although the sequence and interfacial localization of the CRAC fit to the requirements for interaction with cholesterol it has been shown that there is a high statistical probability to find an amino acid arrangement fulfilling the criteria of a CRAC motif23,25. Thus, the motif has been found at high abundance in the proteins of a cholesterol-free bacterium85. Furthermore, of 19 high-resolution crystallographic structures where cholesterol has been detected a single carrier protein but none of the membrane proteins interacted with cholesterol via the CRAC or CARC motifs25. It is interesting to note that recent solid-state NMR studies of MPER- gp41_TMD revealed that cholesterol is localized too far from the membrane external CRAC motif to efficiently interact with this region80. Furthermore, the Influenza virus hemagglutinin M2 TMD tetramer revealed interactions of cholesterol with a hydrophobic cleft but not the CRAC sequence of this protein76,86.
When membrane protein structures which include cholesterol25 are investigated the cholesterol molecules are aligned parallel to the helical long axes with the hydroxyl being situated within the interface (e.g. pdb 4HYT at the level of the lipid phosphate). In this context it is interesting to note that the CRAC-TM_model sequence has an influence on the deuterium labels at positions 2, 3 and 4 of cholesterol but did not affect the deuterium NMR spectra of the 25-position of cholesterol confirming that the interactions between protein and lipid are more pronounced within the interface (Fig. 6). Because here we found that the membrane external CRAC motif (residues 679–683) is an extension of the TM helix it is probably positioned too far out for efficient direct interactions with cholesterol (Fig. 7A) even though the bilayer hydrophobic thickness increases by 4 Å in the presence of 30% cholesterol87. Thus, the weak interactions between cholesterol and the gp41 constructs investigated here probably involve other amino acid side chains than the CRAC motif.
For example, the model TMD starts with KLFIMIALA which positions Leu and Ile residues close to the interface (Table 1). These residues have been shown to be involved in knobs-into-hole interactions with the C19 methyl of cholesterol25. Furthermore, Phe residues have been shown to interact with the C5=C6 double bond by π–π interactions, and finally Lys and Met could establish a hydrogen bond with the cholesterol OH25. As for the gp41 TMD sequence KLFIMIVGG similar considerations apply.
Thus, the membrane proximal and transmembrane regions of gp41 carry several potential interaction sites for cholesterol. The cumulated data suggest that these interactions are relatively weak and interactions are reversible. Thus, different sites compete with each other and with oligomerization interactions within the gp41 transmembrane helix. However, even in the absence of strong, direct molecular interactions cholesterol can have regulatory, structural and dynamic effects on the membrane-embedded protein by changing the physico-chemical properties of the bilayer1. Recent work by the Tamm group has established that the gp41 fusion peptide requires the coexistence of Lo/Ld phases where the line tension at the lipid domain boundaries contributes significantly to overcome the energetic barriers of fusion20. The need for cholesterol to assure efficient fusion of viral and cellular membranes12,13,14 could thereby be associated with the coexistence of Lo and Ld domains, rather than strong direct interactions.
Structural transitions during the fusion process
An extended helix made of residues of the MPER and the gp41 transmembrane domain was observed in this paper as well as in a previous investigation43 (Fig. 7A). In this arrangement cholesterol induces negative curvature strain to the lipid bilayer, a feature that favors the formation of a stalk during the initial stages of the fusion process1. It remains possible that during the membrane fusion events a kink between the MPER and the TMD forms44 which allows the MPER to insert into the membrane interface29. In this configuration the MPER, similar to other amphipathic helices, would exert considerable positive curvature strain29,70 a feature that stabilizes fusion pores1. Thereby, the dynamic nature of the MPER-TMD where different conformations are transiently adopted has the potential to optimize the fusion process by modulating the physical properties of the membrane (Fig. 7B). The lipid interactions, the fusion peptide and the heptadrepeat domains (Fig. 1) have also been postulated to have an active role during the fusion events31,32,88,89,90. Thus, during the various intermediate stages of the fusion process not only different domains of the gp41 protein may be involved but the processes are also associated with different conformations of the MPER that have been observed in structural investigations29,43,44.
Abbreviations
- AIDS:
-
Acquired immune deficiency syndrome
- CARC:
-
Reversed CRAC
- CRAC:
-
Cholesterol recognition amino acid consensus
- CCM:
-
Cholesterol consensus motif
- CD:
-
Circular dichroism
- DHPC:
-
1,2-Di-hexanoyl-sn-glycero-3-phosphocholine
- DMPC:
-
1,2-Di-myristoyl-sn-glycero-3-phosphocholine
- EPR:
-
Electron paramagnetic resonance
- ATR FTIR:
-
Attenuated total reflection Fourier transform infrared
- HIV:
-
Human immunodeficiency virus
- HPLC:
-
High performance liquid chromatography
- MALDI TOF:
-
Matrix assisted laser desorption ionization time of flight
- MPER:
-
Membrane proximal external region
- NMR:
-
Nuclear magnetic resonance
- POPC:
-
1-Palmitoyl-2-oleoyl-sn-glycero-3-phosphocholine
- PRE:
-
Paramagnetic relaxation enhancement
- RDC:
-
Residual dipolar coupling
- SIV:
-
Simian immunodeficiency virus
- SSD:
-
Sterol sensing domain
- TFA:
-
Trifluoroacetic acid
- TFE:
-
Trifluoroethanol
- TMD:
-
Transmembrane domain
References
Yang, S. T., Kreutzberger, A. J. B., Lee, J., Kiessling, V. & Tamm, L. K. The role of cholesterol in membrane fusion. Chem. Phys. Lipids 199, 136–143 (2016).
Harrison, S. C. Viral membrane fusion. Nat. Struct. Mol. Biol. 15, 690–698 (2008).
Zhu, P. et al. Distribution and three-dimensional structure of AIDS virus envelope spikes. Nature 441, 847–852 (2006).
Bartesaghi, A., Merk, A., Borgnia, M. J., Milne, J. L. & Subramaniam, S. Prefusion structure of trimeric HIV-1 envelope glycoprotein determined by cryo-electron microscopy. Nat. Struct. Mol. Biol. 20, 1352–1357 (2013).
Lyumkis, D. et al. Cryo-EM structure of a fully glycosylated soluble cleaved HIV-1 envelope trimer. Science 342, 1484–1490 (2013).
Sattentau, Q. J. & Weiss, R. A. The CD4 antigen: physiological ligand and HIV receptor. Cell 52, 631–633 (1988).
Alkhatib, G. et al. CC CKR5: a RANTES, MIP-1alpha, MIP-1beta receptor as a fusion cofactor for macrophage-tropic HIV-1. Science 272, 1955–1958 (1996).
Wu, L. et al. CD4-induced interaction of primary HIV-1 gp120 glycoproteins with the chemokine receptor CCR-5. Nature 384, 179–183 (1996).
Feng, Y., Broder, C. C., Kennedy, P. E. & Berger, E. A. HIV-1 entry cofactor: functional cDNA cloning of a seven-transmembrane G protein-coupled receptor. Science 272, 872–877 (1996).
Furuta, R. A., Wild, C. T., Weng, Y. & Weiss, C. D. Capture of an early fusion-active conformation of HIV-1 gp41. Nat. Struct. Biol 5, 276–279 (1998).
Blumenthal, R., Durell, S. & Viard, M. HIV entry and envelope glycoprotein-mediated fusion. J. Biol. Chem. 287, 40841–40849 (2012).
Viard, M. et al. Role of cholesterol in human immunodeficiency virus type 1 envelope protein-mediated fusion with host cells. J. Virol 76, 11584–11595 (2002).
Liao, Z., Graham, D. R. & Hildreth, J. E. Lipid rafts and HIV pathogenesis: virion-associated cholesterol is required for fusion and infection of susceptible cells. AIDS Res. Hum Retroviruses 19, 675–687 (2003).
Liao, Z., Cimakasky, L. M., Hampton, R., Nguyen, D. H. & Hildreth, J. E. Lipid rafts and HIV pathogenesis: host membrane cholesterol is required for infection by HIV type 1. AIDS Res. Hum. Retroviruses 17, 1009–1019 (2001).
Sato, R. Recent advances in regulating cholesterol and bile acid metabolism. Biosci. Biotechnol. Biochem. 1–8 (2020).
Aguilar-Ballester, M., Herrero-Cervera, A., Vinue, A., Martinez-Hervas, S. & Gonzalez-Navarro, H. Impact of cholesterol metabolism in immune cell function and atherosclerosis. Nutrients 12 (2020).
Sankaram, M. B. & Thompson, T. E. Modulation of phospholipid acyl chain order by cholesterol. A solid-state 2H nuclear magnetic resonance study. Biochemistry 29, 10676–10684 (1990).
Ipsen, J. H., Mouritsen, O. G. & Zuckermann, M. J. Theory of thermal anomalies in the specific heat of lipid bilayers containing cholesterol. Biophys. J. 56, 661–667 (1989).
Sengupta, P. & Lippincott-Schwartz, J. Revisiting membrane microdomains and phase separation: a viral perspective. Viruses 12 (2020).
Yang, S. T., Kiessling, V. & Tamm, L. K. Line tension at lipid phase boundaries as driving force for HIV fusion peptide-mediated fusion. Nat. Commun. 7, 11401 (2016).
Yang, S. T. et al. HIV virions sense plasma membrane heterogeneity for cell entry. Sci. Adv. 3, e1700338 (2017).
Yang, S. T., Kiessling, V., Simmons, J. A., White, J. M. & Tamm, L. K. HIV gp41-mediated membrane fusion occurs at edges of cholesterol-rich lipid domains. Nat. Chem. Biol. 11, 424–431 (2015).
Epand, R. M. Cholesterol and the interaction of proteins with membrane domains. Prog. Lipid Res. 45, 279–294 (2006).
Fantini, J. & Barrantes, F. J. How cholesterol interacts with membrane proteins: an exploration of cholesterol-binding sites including CRAC, CARC, and tilted domains. Front. Physiol. 4, 31 (2013).
Song, Y. L., Kenworthy, A. K. & Sanders, C. R. Cholesterol as a co-solvent and a ligand for membrane proteins. Protein Sci. 23, 1–22 (2014).
Hanson, M. A. et al. A specific cholesterol binding site is established by the 2.8 A structure of the human beta2-adrenergic receptor. Structure 16, 897–905 (2008).
Motamed, M. et al. Identification of luminal Loop 1 of Scap protein as the sterol sensor that maintains cholesterol homeostasis. J. Biol. Chem. 286, 18002–18012 (2011).
Vincent, N., Genin, C. & Malvoisin, E. Identification of a conserved domain of the HIV-1 transmembrane protein gp41 which interacts with cholesteryl groups. Biochim. Biophys. Acta 1567, 157–164 (2002).
Lorizate, M., Huarte, N., Saez-Cirion, A. & Nieva, J. L. Interfacial pre-transmembrane domains in viral proteins promoting membrane fusion and fission. Biochim. Biophys. Acta 1778, 1624–1639 (2008).
Salzwedel, K., West, J. T. & Hunter, E. A conserved tryptophan-rich motif in the membrane-proximal region of the human immunodeficiency virus type 1 gp41 ectodomain is important for Env-mediated fusion and virus infectivity. J. Virol. 73, 2469–2480 (1999).
Roche, J., Louis, J. M., Grishaev, A., Ying, J. & Bax, A. Dissociation of the trimeric gp41 ectodomain at the lipid-water interface suggests an active role in HIV-1 Env-mediated membrane fusion. Proc. Natl. Acad. Sci. USA 111, 3425–3430 (2014).
Aisenbrey, C. & Bechinger, B. Structure, interactions and membrane topology of HIV gp41 ectodomain sequences Biochim. Biophys. Acta 1862, 183274 (2020).
Li, H. & Papadopoulos, V. Peripheral-type benzodiazepine receptor function in cholesterol transport. Identification of a putative cholesterol recognition/interaction amino acid sequence and consensus pattern. Endocrinology 139, 4991–4997 (1998).
Chen, S. S. L. et al. Identification of the LWYIK motif located in the human immunodeficiency virus type 1 transmembrane gp41 protein as a distinct determinant for viral infection. J. Virol. 83, 870–883 (2009).
Muster, T. et al. A conserved neutralizing epitope on gp41 of human immunodeficiency virus type 1. J. Virol. 67, 6642–6647 (1993).
Huang, J. H. et al. Broad and potent neutralization of HIV-1 by a gp41-specific human antibody. Nature 491, 406 (2012).
Williams, L. D. et al. Potent and broad HIV-neutralizing antibodies in memory B cells and plasma. Sci. Immunol. 2 (2017).
Stiegler, G. et al. A potent cross-clade neutralizing human monoclonal antibody against a novel epitope on gp41 of human immunodeficiency virus type 1. AIDS Res. Hum. Retroviruses 17, 1757–1765 (2001).
Rantalainen, K. et al. HIV-1 envelope and MPER antibody structures in lipid assemblies. Cell Rep. 31, 107583 (2020).
Pinto, D. et al. Structural basis for broad HIV-1 neutralization by the MPER-specific human broadly neutralizing antibody LN01. Cell Host Microbe 26, 623–637 (2019).
Coutant, J. et al. Both lipid environment and pH are critical for determining physiological solution structure of 3-D-conserved epitopes of the HIV-1 gp41-MPER peptide P1. FASEB J. 22, 4338–4351 (2008).
Schibli, D. J., Montelaro, R. C. & Vogel, H. J. The membrane-proximal tryptophan-rich region of the HIV glycoprotein, gp41, forms a well-defined helix in dodecylphosphocholine micelles. Biochemistry 40, 9570–9578 (2001).
Chiliveri, S. C., Louis, J. M., Ghirlando, R., Baber, J. L. & Bax, A. Tilted, uninterrupted, monomeric HIV-1 gp41 transmembrane helix from residual dipolar couplings. J. Am. Chem. Soc. 140, 34–37 (2018).
Fu, Q. et al. Structure of the membrane proximal external region of HIV-1 envelope glycoprotein. Proc. Natl. Acad. Sci. USA 115, E8892–E8899 (2018).
Dev, J. et al. Structural basis for membrane anchoring of HIV-1 envelope spike. Science 353, 172–175 (2016).
Piai, A., Dev, J., Fu, Q. & Chou, J. J. Stability and water accessibility of the trimeric membrane anchors of the HIV-1 envelope spikes. J. Am. Chem. Soc. 139, 18432–18435 (2017).
Buzon, V. et al. Crystal structure of HIV-1 gp41 including both fusion peptide and membrane proximal external regions. PLoS Pathog. 6, e1000880 (2010).
Ofek, G. et al. Structure and mechanistic analysis of the anti-human immunodeficiency virus type 1 antibody 2F5 in complex with its gp41 epitope. J. Virol. 78, 10724–10737 (2004).
Bryson, S. et al. Crystal structure of the complex between the F(ab)’ fragment of the cross-neutralizing anti-HIV-1 antibody 2F5 and the F(ab) fragment of its anti-idiotypic antibody 3H6. J. Mol. Biol. 382, 910–919 (2008).
Dai, Z. et al. Conditional trimerization and lytic activity of HIV-1 gp41 variants containing the membrane-associated segments. Biochemistry 54, 1589–1599 (2015).
Merk, A. & Subramaniam, S. HIV-1 envelope glycoprotein structure. Curr. Opin. Struct. Biol. 23, 268–276 (2013).
Munoz-Barroso, I., Salzwedel, K., Hunter, E. & Blumenthal, R. Role of the membrane-proximal domain in the initial stages of human immunodeficiency virus type 1 envelope glycoprotein-mediated membrane fusion. J. Virol 73, 6089–6092 (1999).
Dimitrov, D. S. & Blumenthal, R. Photoinactivation and kinetics of membrane fusion mediated by the human immunodeficiency virus type 1 envelope glycoprotein. J. Virol 68, 1956–1961 (1994).
Wild, C. T., Shugars, D. C., Greenwell, T. K., McDanal, C. B. & Matthews, T. J. Peptides corresponding to a predictive alpha-helical domain of human immunodeficiency virus type 1 gp41 are potent inhibitors of virus infection. Proc. Natl. Acad. Sci. USA 91, 9770–9774 (1994).
Douek, D. C., Kwong, P. D. & Nabel, G. J. The rational design of an AIDS vaccine. Cell 124, 677–681 (2006).
Caffrey, M. HIV envelope: challenges and opportunities for development of entry inhibitors. Trends Microbiol. 19, 191–197 (2011).
Kwon, B., Lee, M., Waring, A. J. & Hong, M. Oligomeric structure and three-dimensional fold of the HIV gp41 membrane-proximal external region and transmembrane domain in phospholipid bilayers. J. Am. Chem. Soc. 140, 8246–8259 (2018).
Bechinger, B., Resende, J. M. & Aisenbrey, C. The structural and topological analysis of membrane-associated polypeptides by oriented solid-state NMR spectroscopy: Established concepts and novel developments. Biophys. Chem. 153, 115–125 (2011).
Whitmore, L. & Wallace, B. A. Protein secondary structure analyses from circular dichroism spectroscopy: methods and reference databases. Biopolymers 89, 392–400 (2008).
Whitmore, L. & Wallace, B. A. DICHROWEB, an online server for protein secondary structure analyses from circular dichroism spectroscopic data. Nucleic Acids Res. 32, W668–W673 (2004).
Aisenbrey, C., Bertani, P. & Bechinger, B. in Antimicrobial Peptides Methods in Molecular Biology eds A Guiliani & A. C. Rinaldi) Ch. 14, 209–233 (Humana Press, Springer, 2010).
Bechinger, B. & Opella, S. J. Flat-coil probe for NMR spectroscopy of oriented membrane samples. J. Magn. Reson. 95, 585–588 (1991).
Pines, A., Gibby, M. G. & Waugh, J. S. Proton-enhanced NMR of dilute spins in solids. J. Chem. Phys. 59, 569–590 (1973).
Bertani, P., Raya, J. & Bechinger, B. 15N chemical shift referencing in solid state NMR. Solid-State NMR Spec. 61–62, 15–18 (2014).
Bechinger, B. & Sizun, C. Alignment and structural analysis of membrane polypeptides by 15N and 31P solid-state NMR spectroscopy. Concepts Magn. Reson. 18A, 130–145 (2003).
Davis, J. H., Jeffrey, K. R., Bloom, M., Valic, M. I. & Higgs, T. P. Quadrupolar echo deuteron magnetic resonance spectroscopy in ordered hydrocarbon chains. Chem. Phys. Lett. 42, 390–394 (1976).
Salnikov, E. S. et al. Membrane topologies of the PGLa antimicrobial peptide and a transmembrane anchor sequence by dynamic nuclear polarization/solid-state NMR spectroscopy. Sci. Rep. 6, 20895 (2016).
Aisenbrey, C. & Bechinger, B. Tilt and rotational pitch angles of membrane-inserted polypeptides from combined 15N and 2H solid-state NMR spectroscopy Biochemistry 43, 10502–10512 (2004).
Harzer, U. & Bechinger, B. The alignment of lysine-anchored membrane peptides under conditions of hydrophobic mismatch: A CD, 15 N and 31 P solid-state NMR spectroscopy investigation. Biochemistry 39, 13106–13114 (2000).
Aisenbrey, C., Marquette, A. & Bechinger, B. The mechanisms of action of cationic antimicrobial peptides refined by novel concepts from biophysical investigations. Adv. Exp. Med. Biol. 1117, 33–64 (2019).
Goormaghtigh, E., Cabiaux, V. & Ruysschaert, J. M. Determination of soluble and membrane protein structure by Fourier transform infrared spectroscopy. III. Secondary structures. [Review]. Sub-Cell. Biochem. 23, 405–450 (1994).
Salnikov, E., Bertani, P., Raap, J. & Bechinger, B. Analysis of the amide (15)N chemical shift tensor of the C(alpha) tetrasubstituted constituent of membrane-active peptaibols, the alpha-aminoisobutyric acid residue, compared to those of di- and tri-substituted proteinogenic amino acid residues. J. Biomol. NMR 45, 373–387 (2009).
Salnikov, E., Aisenbrey, C., Vidovic, V. & Bechinger, B. Solid-state NMR approaches to measure topological equilibria and dynamics of membrane polypeptides. Biochim. Biophys. Acta 1798, 258–265 (2010).
Mendonca de Moraes, C. & Bechinger, B. Peptide-related alterations of membrane-associated water: Deuterium solid-state NMR investigations of phosphatidylcholine membranes at different hydration levels. Magn. Reson. Chem. 42, 155–161 (2004).
Marsan, M. P. et al. Cholesterol orientation and dynamics in dimyristoylphosphatidylcholine bilayers: A solid state deuterium NMR analysis. Biophys. J. 76, 351–359 (1999).
Elkins, M. R. et al. Cholesterol-binding site of the influenza M2 protein in lipid bilayers from solid-state NMR. Proc. Natl. Acad. Sci. USA 114, 12946–12951 (2017).
Aisenbrey, C., Kemayo-Koumkoua, P., Salnikov, E. S., Glattard, E. & Bechinger, B. Investigations of the structure, topology and interactions of the transmembrane domain of the lipid sorting protein p24 being highly selective for sphingomyelin-C18 Biochemistry 58, 2782–2795 (2019).
Aisenbrey, C., Salnikov, E. S. & Bechinger, B. Solid-state NMR investigations of the MHC II transmembrane domains: topological equilibria and lipid interactions. J. Membr. Biol. 252, 371–384 (2019).
Jaipuria, G. et al. Cholesterol-mediated allosteric regulation of the mitochondrial translocator protein structure. Nat. Commun. 8, 14893 (2017).
Kwon, B. et al. Cholesterol interaction with the trimeric HIV fusion protein gp41 in lipid bilayers investigated by solid-state NMR spectroscopy and molecular dynamics simulations. J. Mol. Biol. 432, 4705–4721 (2020).
Kim, J. H., Hartley, T. L., Curran, A. R. & Engelman, D. M. Molecular dynamics studies of the transmembrane domain of gp41 from HIV-1. Biochim. Biophys. Acta 1788, 1804–1812 (2009).
Long, Y., Meng, F., Kondo, N., Iwamoto, A. & Matsuda, Z. Conserved arginine residue in the membrane-spanning domain of HIV-1 gp41 is required for efficient membrane fusion. Protein Cell 2, 369–376 (2011).
Senes, A., Engel, D. E. & DeGrado, W. F. Folding of helical membrane proteins: the role of polar, GxxxG-like and proline motifs. Curr. Opin. Struct. Biol. 14, 465–479 (2004).
Curran, A. R. & Engelman, D. M. Sequence motifs, polar interactions and conformational changes in helical membrane proteins. Curr. Opin. Struct. Biol. 13, 412–417 (2003).
Palmer, M. Cholesterol and the activity of bacterial toxins. FEMS Microbiol. Lett. 238, 281–289 (2004).
Ekanayake, E. V., Fu, R. & Cross, T. A. Structural influences: cholesterol, drug, and proton binding to full-length influenza A M2 protein. Biophys. J. 110, 1391–1399 (2016).
Nezil, F. A. & Bloom, M. Combined influence of cholesterol and synthetic amphiphillic peptides upon bilayer thickness in model membranes. Biophys. J. 61, 1176–1183 (1992).
Schmick, S. D., Vogel, E. P., Young, K. M. & Weliky, D. P. High-resolution secondary and tertiary structure of the membrane-associated HIV fusion peptide by itself and in large gp41 ectodomain constructs: correlation between beta sheet registry, membrane insertion and perturbation, and fusion catalysis. Biophys. J. 100, 633–633 (2011).
Li, Y. & Tamm, L. K. Structure and plasticity of the human immunodeficiency virus gp41 fusion domain in lipid micelles and bilayers. Biophys. J. 93, 876–885 (2007).
Jaroniec, C. P. et al. Structure and dynamics of micelle-associated human immunodeficiency virus gp41 fusion domain. Biochemistry 44, 16167–16180 (2005).
Acknowledgements
We are thankful to Arnaud Marquette for his help with the FTIR spectra. The financial contributions of the Agence Nationale de la Recherche (projects membraneDNP 12-BSV5-0012, MemPepSyn 14-CE34-0001-01, InMembrane 15-CE11-0017-01, Biosupramol 17-CE18-0033-3 and the LabEx Chemistry of Complex Systems 10-LABX-0026_CSC), the University of Strasbourg, the CNRS, the Région Alsace and the RTRA International Center of Frontier Research in Chemistry are gratefully acknowledged. BB is grateful to the Institut Universitaire de France for providing additional time to be dedicated to research.
Author information
Authors and Affiliations
Contributions
C.A. and O.R. performed experiments helped in writing and prepared Figures. B.B. designed experiments, helped during the discussions and analysis, wrote the paper and assured funding.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Aisenbrey, C., Rifi, O. & Bechinger, B. Structure, membrane topology and influence of cholesterol of the membrane proximal region: transmembrane helical anchor sequence of gp41 from HIV. Sci Rep 10, 22278 (2020). https://doi.org/10.1038/s41598-020-79327-6
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-020-79327-6
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.