Abstract
Wild germplasm resources are crucial for gene mining and molecular breeding because of their special trait performance. Haplotype-resolved genome is an ideal solution for fully understanding the biology of subgenomes in highly heterozygous species. Here, we surveyed the genome of a wild walnut tree from Gongliu County, Xinjiang, China, and generated a haplotype-resolved reference genome of 562.99 Mb (contig N50 = 34.10 Mb) for one haplotype (hap1) and 561.07 Mb (contig N50 = 33.91 Mb) for another haplotype (hap2) using PacBio high-fidelity (HiFi) reads and Hi-C technology. Approximately 527.20 Mb (93.64%) of hap1 and 526.40 Mb (93.82%) of hap2 were assigned to 16 pseudochromosomes. A total of 41039 and 39744 protein-coding gene models were predicted for hap1 and hap2, respectively. Moreover, 123 structural variations (SVs) were identified between the two haplotype genomes. Allele-specific expression genes (ASEGs) that respond to cold stress were ultimately identified. These datasets can be used to study subgenome evolution, for functional elite gene mining and to discover the transcriptional basis of specific traits related to environmental adaptation in wild walnut.
Similar content being viewed by others
Background & Summary
Common walnut (Juglans regia L.) (2n = 32), which belongs to the Juglans genus of Juglandaceae, is an important economic tree species and the source of the fourth most consumed nut1. The fruits of common walnut are mainly used for the production of oil and health care products due to their lipid profile and abundance of antioxidants, such as phenolic compounds, vitamins and micronutrients2,3,4, which have potential beneficial roles in health maintenance and disease prevention5,6. These properties have promoted walnut cultivation and production in many countries, such as the USA, Iran, Turkey and China7.
Walnut likely originated and was domesticated in Central Asia8,9 and has been found in China for more than 7000 years10. Currently, China is regarded as a secondary centre of origin, harbouring wide genetic diversity11,12. Wild walnut is widely distributed in a nature reserve in Gongliu County, Xinjiang, China13,14,15. Wild relatives often harbour elite alleles conferring tolerance to extreme biotic or abiotic stress in specific environments16,17,18, which can provide resistance resources for genetic improvement. Xinjiang wild walnut shows relatively high stress resistance19,20 and good nutritional components21. Specifically, the elite cold resistance of wild walnut can guarantee its survival and propagation in long-term extreme cold, whether in winter or late spring, in Gongliu County. Therefore, wild walnut in this area is an ideal material for cold-improvement molecular breeding and the identification of cold stress regulatory mechanisms. However, walnut is a nonmodel tree species, and most studies have focused on the quality of cold-stressed walnut kernels22,23 or phenotyping the cold resistance of walnut varieties24, with few focusing on the molecular mechanism of cold tolerance25.
The draft genome of walnut was released in 201626 and improved in 201827. Since then, reference genome assemblies have been published successively with higher genome continuity or at the chromosome scale by means of long-read sequencing and Hi-C anchoring28,29. In addition, the genome sequences of six species of Juglandaceae (J. sigillata, J. cathayensis, J. mandshurica, J. hindsii, J. microcarpa and J. nigra30) are also available (http://xhhuanglab.cn/data/juglans.html). However, the walnut genome has high heterozygosity between diploid chromosomes27,28,29, and the current genomes have mosaic assemblies of haplotypes. Recently, haplotype-defined genome assembly was realized in highly heterozygous organisms such as Manihot esculenta31, Camellia sinensis32, Artemisia annua33 and Malus spp.34, which provided novel insight into allelic variation calling and functional differentiation of divergent alleles in heterozygous species.
Diploid organisms have two allelic copies in their genome, but in a given individual, the alleles are not necessarily both active or active at the same level. This unequal expression of gene copies caused by cis-acting genetic variants or CHG methylation is called allele-specific expression (ASE)35,36. ASE was first reported in yeast in 200237 and is involved in many plant biological processes, such as the response to mild water deficit in tomato38 and heterosis via partial to full dominance or overdominance effects on the traits being regulated and chromatin accessibility alterations in rice39,40. In recent years, genome-wide ASE features and ASE-related genes have been well characterized, benefitting from haplotype-resolved genome assembly in this third-generation sequencing era41,42. In the woody perennial plant apple, approximately 19% of the expressed genes were allele specific, and many ASE genes, such as genes encoding ACC oxidase and RIN-like MADS-box transcription factors, were associated with fruit quality43. Furthermore, Tian et al.34 reported that transposable elements (TEs) can regulate ASE to determine flower colour in apple.
This study reports a high-quality haplotype-resolved reference genome of wild walnut. First, Illumina sequencing reads were used to estimate the genome size, heterozygosity and proportion of repetitive sequences. PacBio high-fidelity (HiFi) long reads were adopted to primarily assemble highly continuous contigs, followed by HiFi read phasing to reassemble the resulting haplotypes. Then, two haplotypes (hap1 and hap2) at the chromosome scale were identified by Hi-C technology-assisted assignment. The final assembly lengths were 562.99 Mb and 561.07 Mb for hap1 and hap2, respectively, with contig N50 values of 33.91 Mb and 22.40 Mb, respectively. Approximately 93.64% of the contigs (527.2 Mb) were anchored onto 16 pseudochromosomes for hap1, and 93.82% (526.4 Mb) were anchored onto 16 pseudochromosomes for hap2. The LTR assembly index (LAI v2.9.4)44, BUSCO v5 completeness45, genome collinearity and phasing switch errors32 were used to evaluate the completeness, continuity and phasing accuracy of the haplotype genomes. We predicted 41039 gene models for hap1 and 39744 gene models for hap2, and 34617 and 34562 genes were functionally annotated by four conventional databases, respectively. A total of 358869 and 358104 simple sequence repeats (SSRs) were also identified for hap1 and hap2, respectively. The genome variation between hap1 and hap2, including different types of structural variations (SVs), is highlighted. Finally, 358, 457, 465, 494 and 386 ASEGs were identified at 0 h, 3 h, 6 h, 12 h and 24 h after cold treatment, respectively. This high-quality haplotype-resolved genome assembly will shed light on subgenome evolution, molecular genetics and ASE regulatory mechanisms underlying adaptation to the extreme cold environments of wild walnut.
Methods
Plant material preparation, Illumina short-read library construction and sequencing
Approximately 20 g of young leaves from a wild walnut tree was collected from the nature reserve of Gongliu County, Xinjiang, China (E82°16′28″, N43°20′52″). The altitude of the wild walnut habitat ranges from 1241 m to 1670 m, and the annual average temperature is 7.6 °C, with an average of 19.7 °C in July and −9.9 °C in January46,47. Under cold stress, wild walnut plants exhibit better performance than seedling walnut48. The location and appearance are shown in Fig. 1. Then, the leaves were cut into pieces and rapidly placed in liquid nitrogen. The genomic DNA of the fresh leaves was isolated using a Super Plant Genomic DNA DP360 Kit (Tiangen Biotech, Beijing, China), followed by DNA concentration measurement using a NanoDrop spectrophotometer (ND2000, Thermo Fisher Scientific, USA) and quality monitoring by electrophoresis on 0.80% agarose gels. High-quality DNA was subjected to ultrasonic fragmentation (350 bp) using a Covaris S2 instrument (Covaris, Woburn, MA, USA). A 350 bp paired-end resequencing library was prepared using a TruSeq DNA Sample Prep Kit (Illumina, San Diego, CA, USA) according to the manufacturer’s standard protocols. An Agilent 2100 bioanalyzer (Agilent Technologies, Palo Alto, CA) and q-PCR were used to measure the fragment size and quality of the library. To aid in gene model prediction, total RNA was extracted from mixed samples containing leaf, root, shoot and flower tissues (~1 g) from the same tree using the RNeasy Plant Mini Kit (Qiagen, Germany) according to the manufacturer’s instructions and treated with an RNase-free DNase I digestion kit (Aidlab, China). RNA degradation was monitored using a 1% agarose gel, and the purity of the isolated RNA was measured using a NanoDrop 2000 spectrophotometer (Thermo Scientific, Wilmington, DE, USA). The integrity of the RNA was assessed on an Agilent 2100 Bioanalyzer (Agilent, Palo Alto, CA, USA), and the RNA library was constructed with the TruSeq RNA Library Prep Kit v.2 (Illumina, San Diego, CA, USA). Both the DNA and RNA libraries were sequenced on an Illumina NovaSeq. 6000 sequencer (Illumina, San Diego, CA, USA) according to the manufacturer’s recommendations. The raw reads were filtered to generate high-quality clean reads by (i) removing adaptor sequences, (ii) filtering reads with >10% unidentified nucleotides, and (iii) removing reads with >50% bases with a low Phred quality score (≤10). As a result, 31.47 Gb (~56×) and 10 Gb (RNA-seq data) of high-quality clean reads were generated for the genome survey and gene model prediction, respectively.
PacBio HiFi long-read library generation and sequencing
High-integrity genomic DNA extraction was performed according to Mayjonade’s methods49. The HiFi library was used according to PacBio’s standard protocols. Briefly, 10 μg of high-quality gDNA was randomly disrupted by a Covaris g-TUBE device (Covaris, Woburn, MA) and transferred for DNA damage repair and end repair. Then, dumbbell-shaped adaptors were ligated. Thereafter, the products were subjected to exonuclease digestion followed by size selection to screen segments between 10 kb and 30 kb in size using BluePippin (Sege Science, USA). The final library was sequenced on a PacBio Sequel II platform (circular consensus sequencing (CCS) model) using two flow cells for 35 h (Adsen Biotechnology Company, Urumqi, China). Finally, we obtained approximately 57.86 Gb of HiFi reads (~ 103 × depth) with a CCS contig N50 and greatest length of 15.91 kb and 50.33 kb, respectively.
Hi-C library preparation and sequencing
As living samples can provide integrated and real interaction states in vivo, seeds from wild walnuts (for DNA extraction) were collected and germinated indoors. Seedlings at the four-leaf stage were used for Hi-C library preparation according to Lieberman–Aiden’s method50. In brief, fresh leaves were immediately crossed with formaldehyde, and the cross-linked DNA was then digested with Hind III (NEB). Subsequently, the sticky ends of these enzyme-digested fragments were end-repaired by a biotin-modified base. Circular DNA molecules were continuously generated using blunt-end proximity-ligated fragments and fragmented into 300–700 bp fragments. Finally, the fragments were enriched by biotin beads, and after library quality control by a Qbit 2.0 (Life Technologies, Carlsbad, CA, US) and an Agilent 2100 bioanalyzer (Agilent Technologies, Palo Alto, CA, USA), the library was subjected to paired-end 150 bp (PE150) sequencing in one lane (Illumina, San Diego, CA, USA). This Hi-C library generated approximately 64.61 Gb (~115 × depth) of clean data.
Genome survey analysis
A total of approximately 31.47 Gb of clean reads were generated for the genome survey using GenomeScope v151 with the parameters “k = 15, 17, 19, 21, 23; length = 100; and max coverage = 10000”. We obtained an estimated genome size of 470.69 Mb-513.17 Mb with a heterozygosity of 0.29%–0.41% for this wild walnut (Table 1).
De novo assembly of the haplotype-resolved chromosome-scale genome
After adaptor filtering, the HiFi reads were employed to assemble a primary contig dataset using hifiasm v0.16.1 with default parameters52. Second, all CCS reads were remapped to the above assembled contigs using minimap2 v2.2453, and the parameters were set as -secondary = n -cx map-pb. Third, Longshot (default settings) was used to call variations54. Fourth, variation phasing was realized by using WhatsHap with default parameters55. Fifth, CCS reads were separated into two haplotypes (hap1 and hap2) and an unassigned group according to the phased variations. Sixth, CCS reads of the hap1 combined unassigned group and hap2 combined unassigned group were assembled independently using hifiasm v0.16.1 with default parameters52. For genome scaffolding, clean Hi-C data were mapped to the contigs of hap1 and hap2 by BWA v0.7.1056. HiCExplorer57 was used to calculate mapping reads and interaction pairs. For hap1, a total of 120.89 million reads were uniquely mapped, of which 51.16 million were valid interactions, accounting for 42.32%. For hap2, 119.03 million reads were uniquely mapped, and 50.58 million reads were observed to interact with other pairs, for a percentage of valid data of 42.49%. Second, valid interaction read pairs were used to cluster and orient the assembled haplotype-resolved contigs onto pseudochromosomes by using HiCAssembler58 (options: -min_scaffold_length 100000; -bin_size 20000; -misassembly_zscore_threshold 1.0; -num_iterations 4; -num_processors 30). The Juicebox tool (https://github.com/aidenlab/Juicebox) was used to fine-tune the pseudochromosomes manually. This final assembly resulted in a total of 562.99 Mb (contig N50 = 34.10 Mb) assigned to 16 pseudochromosomes for hap1, of which 527.20 Mb could be oriented, accounting for 93.64% of the haplotype genome size, while 526.40 Mb of hap2 was oriented, accounting for 93.82% of the 561.07 Mb (contig N50 = 33.91 Mb) of the assembled sequences (Table 2; Fig. 2c,d). Heatmaps were produced to display the interactions of genome read pairs at a resolution of 20 kb (Fig. 2a,b).
Hi-C heatmap and Circos plot for hap1 and hap2. (a,b) show the interaction links of the Hi-C data at a bin size of 20 kb. The interaction intensity is indicated using colours ranging from light yellow to dark red, which denote the frequency of Hi-C interactions from low to high. (c,d) Circos plot of the genomes of hap1 and hap2 with a window size of 500 kb. This Circos plot includes six components: (a) LAI (LTR assembly index) values; (b) density of GC content; (c) gene density; (d) density of Copia retrotransposons; (e) density of Gypsy retrotransposons; and (f) density of TEs (transposable elements).
Genome-wide repetitive sequence annotation
A comprehensive transposable element (TE) prediction tool, Extensive Denovo TE Annotator59 (options: -t 20 -step all -sensitive 1 -anno 1), was adopted to construct a nonredundant TE library with Helitron and long terminal repeat (LTR) identification. TIR-Learner v2.560 was used to search for terminal inverted repeats (TIRs) under default recommendations. RepeatMasker v4.0561 was used to predict the final repeat sequences of the genome under the parameters -nolow -no_is -norna -engine wublast. Finally, we identified 46.26% and 46.23% repetitive elements in the hap1 and hap2 assemblies, respectively, including similar components of the LTR type (4%–8%), TIR type (0–5%), non-LRT type (~0.50%) and non-TIR type (2%–9%) between hap1 and hap2 (Table 3; Fig. 2c,d).
Gene model prediction and functional annotation
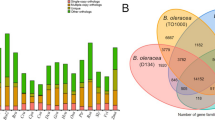
Three steps were combined for protein-coding gene model prediction. First, AUGUSTUS v3.4.062 and GeneMark-ES v4.6863 were used to search for gene models via ab initio prediction. Second, GeMoMa v1.3.164 was used for homology prediction. Then, PASA (parameters: -align_tools gmap, -maxIntronLen 20000) was used for gene prediction via RNA-seq65. Finally, 41039 gene models were annotated for hap1, and 39744 gene models were annotated for hap2 (Table 4; Fig. 2c,d). For functional annotation, the above gene model sequences were translated into protein sequences and subsequently compared against the COG v2.1.466, GO67, Pfam68 and KEGG69 databases. In total, 34617 hap1 and 34562 hap2 nonredundant gene models were annotated by these biological databases, accounting for 84.35% and 86.96% of all gene models, respectively (Table 4). The gene model numbers of hap1 and hap2 were greater than those reported in Chandler26 (32498), Serr (Payne × PI 159568)70 (31425) and Ding et al.71 (33430) and lower than those reported in Chandler28 (37554) and Zhongmucha-129 (39432).
Noncoding RNA annotation
tRNAscan-SE v2.0.972 (default settings), barrnap v0.973 (options: -kingdom euk -threads 8) and Infernal v1.074 (option: -Z $Z -cut_ga -rfam –nohmmonly -fmt 2) combined with Rfam75 were used for the tRNA, rRNA and miRNA/snoRNA searches, respectively. Overall, 1817 (500 tRNA, 72 rRNA, 207 miRNA and 1038 snoRNA) and 1919 (500 tRNA, 104 rRNA, 210 miRNA and 1105 snoRNA) noncoding RNAs were discovered in the genomes of hap1 and hap2, respectively (Table 4).
Simple sequence repeat (SSR) recognition and variation calling
Hap1 and hap2 were analysed by using TBtools v 1.09875176 for SSR recognition under default settings. We identified 358869 SSRs for hap1 and 358104 SSRs for hap2. In addition, we used hap2 as a query to map it to hap1 using minimap2 v2.2453 under the settings -eqx -ax asm5, followed by variation detection using SyRI77 under default settings. Afterwards, we detected 21 inversions, 40 translocations, 575 presence/absence variants (PAVs), 2 duplications in hap1 and 60 duplications in hap2 (Table 5; Fig. 3). Notably, the lengths of the PAVs were 274,629 bp in hap1 and 274.840 bp in hap2, accounting for 303 and 272 genes in hap1 and hap2, respectively (see repository files78). Similarly, the structural variations between hap1 and three released genomes29,70,71 of Juglans were identified (see repository files78).
SV (structural variation) distribution between the haplotypes. The horizontal blue lines indicate the chromosomes of hap1, and the red lines indicate the chromosomes of hap1. The grey, orange, light blue and green lines indicate the syntenic, inverted, duplicated and trans-located regions, respectively.
Allele-specific expression genes (ASEGs) under cold stress
To explore the response of ASEGs to cold stress, three-leaf-stage seedlings derived from the kernels of wild walnut trees were treated at 4 °C in a climate chamber (Shanghai Boxun Industrial Co., Ltd., Shanghai, China). Leaves (approximately 0.5 g) were sampled after treatment for 0 h, 3 h, 6 h, 12 h and 24 h, with three biological repetitions, and immediately placed in liquid nitrogen for quick freezing. RNA-seq libraries were constructed, and sequencing was performed (see Methods: Plant material preparation, Illumina short-read library construction and sequencing). MCScanX79 was used to identify collinear block gene pairs between hap1 and hap2 with the default settings. The scaffold-involved collinear gene pairs were removed. Subsequently, the collinear gene pairs were checked by BLAST80 with a similarity between 90% and 100%. Ultimately, we obtained 18770 gene pairs. The clean RNA-seq data were mapped to the walnut genomes using HISAT2 v2.2081, and TPM values were used for expression quantification by StringTie v 2.1.282. DESeq 283 was used to determine ASEGs with the criteria |log2Fold Change| > 1 and adjusted p < 0.05. We obtained 358, 457, 465, 494 and 386 ASEGs at 0 h, 3 h, 6 h, 12 h and 24 h, respectively, accounting for approximately 2% of all the identified collinear genes (see repository files78; Fig. 4). Of all the ASEGs, 234 were shared, and 12, 37, 22, 32 and 20 were specifically detected in the 0 h, 3 h, 6 h, 12 h and 24 h samples, respectively (Fig. 5b). We also noticed that more frequent ASEGs were dominantly expressed on hap2 (Fig. 5a), indicating uncoordinated allelic expression patterns for the ASEGs under cold stress in wild walnut.
Allele-specific gene expression in hap1 and hap2 under cold stress. (a) Allele-specific expression genes (ASEGs) on haplotype chromosomes at five time points under cold stress (0 h, 3 h, 6 h, 12 h, 24 h). The numbers of ASEGs are displayed as the mean ± s.d. (b) Venn diagram of the ASEGs at five time points under cold stress. The numbers indicate unique and common ASEGs for different time points.
Data Records
Illumina reads for genome survey, RNA-seq reads and Hi-C reads were deposited in the Sequence Read Archive (SRA) (accession numbers: PRJNA858167, PRJNA859241, and PRJNA858917, respectively)84,85,86. HiFi long reads were co-deposited in the National Genomics Data Center (https://ngdc.cncb.ac.cn/) under the accession number CRA007543 and SRA under accession number PRJNA94732987. The genome assembly of wild walnut (Juglans regia L.) has been deposited on the Figshare platform (https://doi.org/10.6084/m9.figshare.22266730)88 and GenBank with accession numbers GCA_034508915.1 and GCA_034509015.1, respectively89,90. The RNA-seq reads of cold-stressed samples were deposited in the SRA under accession number PRJNA94242691.
Technical Validation
Sequencing read quality control
For Illumina-based sequencing data (sequencing for genome survey, Hi-C reads and RNAseq reads), raw reads were removed to obtain clean data according to the following steps: (1) filtering adaptors; (2) removing reads with >10% unidentified nucleotides; and (3) removing reads with >50% bases with a Q value ≤ 10. The clean data were then evaluated in terms of the insert size, GC content, Q30 value, quality distribution and base composition. Specifically, we used HiCUP92 (v0.6.1) for preprocessing and to evaluate the Hi-C library quality. After trimming the restriction enzyme site of HindIII by calling the HiCUP_truncater script, the Hi-C clean data were aligned to the haplotype genomes by BWA v0.7.1056, and only the unique mapped sequences with a mapping score of at least 20 were used for the following process. For the HiFi long reads, subreads from the same template strand were consistently corrected, and adapters and low-quality reads were filtered. Before processing, 2000 randomly selected long reads were subjected to contamination evaluation by BLAST80 against the Nt database (ftp://ftp.ncbi.nih.gov/blast/db) under the threshold of P < 1e-05.
Haplotype genome assembly evaluation
BUSCO v545 evaluation revealed that more than 94% of Embryophyta genes were successfully assembled in both hap1 and hap2 (Fig. 6), indicating that the completeness of these haplotype-resolved genomes was in line with that of the published genome29. The LAI44 values of hap1 and hap2 were 13.36 and 13.41, respectively. The CC ratios93 of hap1 and hap2 were 43.25 and 42.94, respectively. Furthermore, we used hap1 and hap2 as queries for mapping against two published genomes29,71 in minimap2 v2.2453 under the settings -eqx -ax asm5. Good collinearities were observed between our haplotype genomes and the two published genomes29,71 (Fig. 7). Finally, the phasing accuracy of the haplotype-resolved assembly was assessed by the percentage of switch errors using the modified calc_switchErr pipeline32,94. As 10x Genomics linked reads were not available, the criterion consistency phased SNP step was excluded from the phasing evaluation pipeline. A total of 184957 SNPs were detected, including 33061 switched SNPs, indicating a switch error rate of 17.87%, which is similar to that in the genome of Solanum tuberosum L. (17.1%) and lower than that in the genomes of Malus domestica cv. Gala (22.2%) and Vanilla planifolia (44.0%)94. We further used published Nanopore sequencing data (SRR10001245) from the Juglans regia L. cultivar Chandler28 to evaluate phasing accuracy. We first polished the ONT data using the genome resequencing data of wild walnut with Ratatosk95 v0.7.0 under default settings; then, the same method was used to calculate the switch error. Finally, we obtained a switch error of 6.25%. Collectively, these results indicate that a reference-level high-quality haplotype-resolved genome of wild walnut was obtained.
Summary assessment bar chart for BUSCO evaluation in hap1, hap2 and published genomes of Juglans (http://xhhuanglab.cn/data/juglans.html). Light blue indicates the percentage of complete and single-copy genes, while darker blue indicates the percentage of complete and duplicated genes; yellow indicates the percentage of fragmented genes; and red indicates the percentage of missing genes in the assemblies.
Collinearity evaluation between haplotypes and the published genome of Juglans regia L29,71. (a) The collinearity between hap1 and the published Juglans regia L. genome29. (b) The collinearity between hap2 and the published Juglans regia L. genome29. (c) The collinearity between hap1 and the published Juglans regia L. genome71. (d) The collinearity between hap2 and the published Juglans regia L. genome71. To distinguish the two published genomes, chromosome71 collinearity was marked using different colours.
Published QTL confirmation
To further validate the accuracy of the genome, we downloaded published reduced-representation sequencing data from Juglans regia L., which were used for high-density genetic map construction and QTL mapping96. A total of 2540 out of 2577 markers (98.56%), developed by specific length amplified fragment sequencing (SLAF-seq), on the genetic map can be mapped to the genome of hap1 with an identity >90% and an E threshold value < E-10 (see repository files78). Specifically, a QTL identified by the authors for anthracnose resistance harbouring 10 markers was found to be best mapped to chr1:14148922–14220999 of hap1, a region of approximately 721 kb (Table 6). A similar discovery was made for hap2. These results indicated the high accuracy of the subgenomes of this wild walnut.
Code availability
No specific code was developed in this work. All the data were analyzed following the manuals suggested by the developers of the bioinformatic tools, which have been described in the Methods section.
References
Ji, F. et al. A genome variation map provides insights into the genetics of walnut adaptation and agronomic traits. Genome Biol. 22, 300 (2021).
Miraliakbari, H. & Shahidi, F. Antioxidant activity of minor components of tree nut oils. Food Chem. 111, 421–427 (2008).
Şen, S. M. & Karadeniz, T. The nutritional value of walnut. J. Hyg. Eng. Design 11, 68–71 (2015).
Ros, E., Izquierdo-Pulido, M. & Sala-Vila, A. Beneficial effects of walnut consumption on human health: Role of micronutrients. Curr. Opin. Clin. Nutr. 21, 498–504 (2018).
Sánchez-González, C., Ciudad, C. J., Noe, V. & Izquierdo-Pulido, M. Health benefits of walnut polyphenols: An exploration beyond their lipid profile. Crit. Rev. Food Sci. 57, 3373–3383 (2017).
Ni, Z. J. et al. Exploration of walnut components and their association with health effects. Crit. Rev. Food Sci. 62, 1881439 (2021).
Bernard, A., Lheureux, F. & Dirlewanger, E. Walnut: past and future of genetic improvement. Tree Genet. Genom. 14, 1 (2018).
Vahdati, K. et al. Advances in Persian walnut (Juglans regia L.) breeding strategies Vol. 4 (eds. Al-Khayri, J., Jain, S. & Johnson, D.) Ch. 11 (Springer Cham, 2019).
Vahdati, K., Arab, M. M. & Sarikhani, S. Advances in Biotechnology and Propagation of Nut Trees in Iran. In: BIO Web of Conferences 25, p.01003 (EDP Sciences, 2020).
Xi, R. T. Discussion on the origin of walnut in China. Acta Hortic. 284, 353–62 (1990).
Chen, L., Ma, Q., Chen, Y., Wang, B. & Pei, D. Identification of major walnut cultivars grown in China based on nut phenotypes and SSR markers. Sci. Hortic. 168, 240–248 (2014).
Feng, X. et al. The phytogeographic history of common walnut in China. Front. Plant Sci. 9, 1399 (2018).
Dong, Y. Z. et al. Investigation and analysis on the wild walnut in Gongliu, Xinjiang. J. Plant Genet. Res. 13, 386–392 (2012). (In Chinese).
Deng, F. B. et al. Analysis of phenotypic diversity of nuts in wild walnut (Juglans cathayensis Dode) in Xinjiang. J. Fruit Sci. 35, 275–284 (2018).
Zhang, W., Li, H. Y., Cui, D. & Yang, Y. F. Age structure and growth characteristics of Juglans cathayensis seedling populations at different slope aspects in Wild Walnut Natural Conservation Area of Xinjiang, China. J. Appl. Ecol. 28, 382–390 (2017).
Reuscher, S. et al. Assembling the genome of the African wild rice Oryza longistaminata by exploiting synteny in closely related Oryza species. Commun. Biol. 1, 162 (2018).
Szymański, J. et al. Analysis of wild tomato introgression lines elucidates the genetic basis of transcriptome and metabolome variation underlying fruit traits and pathogen response. Nat. Genet. 52, 1111–1121 (2020).
Yu, D. et al. Multi-omics assisted identification of the key and species-specific regulatory components of drought‐tolerant mechanisms in Gossypium stocksii. Plant Biotechnol. J. 19, 1690 (2021).
Zhang, M. H. et al. Physiological response of Xinjiang wild walnut germplasm to low temperature stress. J. Appl. Ecol. 31, 2558–2566 (2020).
Yu, T., Zhang, P. & Chen, W. D. Effects of drought stress on protective enzyme and malondialdehyde content of three kinds of Xinjiang wild walnut. Jiangsu Agr. Sci. 46, 119–121 (2018).
Tian, J., Wu, Y., Wang, Y. & Han, F. Development and prospects of the walnut industry in China. Int. Soc. Hortic. Sci. 861, 31–38 (2010).
Christopoulos, M. V. & Tsantili, E. Participation of phenylalanine ammonia-lyase (PAL) in increased phenolic compounds in fresh cold stressed walnut (Juglans regia L.) kernels. Postharvest Biol. Tec. 104, 17–25 (2015).
Liu, D., Zhang, B., Sun, H., Peng, S. & Zhu, H. Comprehensive evaluation on cold resistance of early fruiting walnut cultivars. Acta Hortic. Sin. 42, 545–553 (2015).
Zhang, R. et al. Evaluation of cold tolerance of seven walnut varieties. Cryoletters 43, 74–82 (2022).
Han, L. et al. The cold-stress responsive gene DREB1A involved in low-temperature tolerance in Xinjiang wild walnut. PeerJ 10, e14021 (2022).
Martínez-García, P. J. et al. The walnut (Juglans regia) genome sequence reveals diversity in genes coding for the biosynthesis of non-structural polyphenols. The Plant J. 87, 507–532 (2016).
Stevens, K. A. et al. Genomic variation among and within six Juglans species. G3-Genes Genom. Genet. 8, 2153–2165 (2018).
Marrano, A. et al. High-quality chromosome-scale assembly of the walnut (Juglans regia L.) reference genome. Gigascience 9, giaa050 (2020).
Zhang, J. et al. A high-quality walnut genome assembly reveals extensive gene expression divergences after whole-genome duplication. Plant Biotechnol. J. 18, 1848–1850 (2020).
Zhou, H. et al. Pan-genome and transcriptome analyses provide insights into genomic variation and differential gene expression profiles related to disease resistance and fatty acid biosynthesis in eastern black walnut (Juglans nigra). Hortic. Res. 10, uhad015 (2023).
Hu, W. et al. Allele-defined genome reveals biallelic differentiation during cassava evolution. Mol. Plant 14, 851–854 (2021).
Zhang, X. et al. Haplotype-resolved genome assembly provides insights into evolutionary history of the tea plant Camellia sinensis. Nat. Genet. 53, 1250–1259 (2021).
Liao, B. et al. Allele-aware chromosome-level genome assembly of Artemisia annua reveals the correlation between ADS expansion and artemisinin yield. Mol. Plant 15, 1310–1328 (2022).
Tian, Y. et al. Transposon insertions regulate genome-wide allele‐specific expression and underpin flower colour variations in apple (Malus spp.). Plant Biotechnol. J. 20, 1285–1297 (2022).
Cleary, S. & Seoighe, C. Perspectives on allele-specific expression. Annu. Rev. Biomed. Data Sci. 4, 101–122 (2021).
Ma, X. et al. Parental variation in CHG methylation is associated with allelic-specific expression in elite hybrid rice. Plant Physiol. 186, 1025–1041 (2021).
Brem, R. B., Yvert, G., Clinton, R. & Kruglyak, L. Genetic dissection of transcriptional regulation in budding yeast. Science 296, 752–755 (2002).
Albert, E. et al. Allele-specific expression and genetic determinants of transcriptomic variations in response to mild water deficit in tomato. The Plant J. 96, 635–650 (2018).
Shao, L., Xing, F., Xu, C. & Zhang, Q. Patterns of genome-wide allele-specific expression in hybrid rice and the implications on the genetic basis of heterosis. P. Natl. Acad. Sci. USA 116, 5653–5658 (2019).
Wang, P. et al. Allele-specific expression and chromatin accessibility contribute to heterosis in tea plants (Camellia sinensis). The Plant J. (2022).
Cheng, S. P. et al. Haplotype-resolved genome assembly and allele-specific gene expression in cultivated ginger. Hortic. Res. 8, 188 (2021).
Qi, W. et al. The haplotype-resolved chromosome pairs of a heterozygous diploid African cassava cultivar reveal novel pan-genome and allele-specific transcriptome features. GigaScience. 11, giac028 (2022).
Sun, X. et al. Phased diploid genome assemblies and pan-genomes provide insights into the genetic history of apple domestication. Nat. Genet. 52, 1423–1432 (2020).
Ou, S., Chen, J. & Jiang, N. Assessing genome assembly quality using the LTR Assembly Index (LAI). Nucleic Acids Res. 46, e126–e126 (2018).
Simão, F. A., Waterhouse, R. M., Ioannidis, P., Kriventseva, E. V. & Zdobnov, E. M. BUSCO: assessing genome assembly and annotation completeness with single-copy orthologs. Bioinformatics 31, 3210–3212 (2015).
Zhang, W., Jia, F. Q., Na, S. B. & Yang, Y. F. Population dynamics and growth of Juglans cathayensis seedlings of different slope aspects in the west Tianshan Valley in Xinjiang, China. Chin J. Ecol. 33, 2596–2602 (2014).
Zeng, B. Present status, problems, and further development strategies on nature Juglans fallax dode resource and their exploitation in Xinjiang. N. Fruits 4, 1–3 (2005).
Han, L. Q. et al. Physiological response and evaluation of cold resistance of Xinjiang wild walnut under low temperature stress. J. Northwest Forestry Univ. 34, 98–101 (2019).
Mayjonade, B. et al. Extraction of high-molecular-weight genomic DNA for long-read sequencing of single molecules. Biotechniques 61, 203–205 (2016).
Lieberman-Aiden, E. et al. Comprehensive mapping of long-range interactions reveals folding principles of the human genome. Science 326, 289–293 (2009).
Vurture, G. W. et al. GenomeScope: fast reference-free genome profiling from short reads. Bioinformatics 33, 2202–2204 (2017).
Cheng, H., Concepcion, G. T., Feng, X., Zhang, H. & Li, H. Haplotype-resolved de novo assembly using phased assembly graphs with hifiasm. Nat. Methods 18, 170–175 (2021).
Li, H. Minimap2: pairwise alignment for nucleotide sequences. Bioinformatics 34, 3094–3100 (2018).
Edge, P. & Bansal, V. Longshot enables accurate variant calling in diploid genomes from single-molecule long read sequencing. Nat. Commun. 10, 4660 (2019).
Patterson, M. et al. WhatsHap: weighted haplotype assembly for future-generation sequencing reads. J. Comput. Biol. 22, 498–509 (2015).
Li, H. & Durbin, R. Fast and accurate short read alignment with Burrows-Wheeler transform. Bioinformatics 25, 1754–1760 (2009).
Ramírez, F. et al. High-resolution TADs reveal DNA sequences underlying genome organization in flies. Nat. Commun. 9, 189 (2018).
Renschler, G. et al. Hi-C guided assemblies reveal conserved regulatory topologies on X and autosomes despite extensive genome shuffling. Gene Dev. 33, 1591–1612 (2019).
Ou, S. et al. Benchmarking transposable element annotation methods for creation of a streamlined, comprehensive pipeline. Genome Biol. 20, 275 (2019).
Su, W., Gu, X. & Peterson, T. TIR-Learner, a new ensemble method for TIR transposable element annotation, provides evidence for abundant new transposable elements in the maize genome. Mol. Plant 12, 447–460 (2019).
Chen, N. Using RepeatMasker to identify repetitive elements in genomic sequences. Curr. Protoc. Bioinform. 25, 4–10 (2004).
Stanke, M., Steinkamp, R., Waack, S. & Morgenstern, B. AUGUSTUS: a web server for gene finding in eukaryotes. Nucleic Acids Res. 32, W309–W312 (2004).
Besemer, J. & Borodovsky, M. GeneMark: web software for gene finding in prokaryotes, eukaryotes and viruses. Nucleic Acids Res. 33, W451–W454 (2005).
Keilwagen, J. et al. Using intron position conservation for homology-based gene prediction. Nucleic Acids Res. 44, e89–e89 (2016).
Campbell, M. A. et al. Comprehensive analysis of alternative splicing in rice and comparative analyses with Arabidopsis. BMC Genomics 7, 327 (2006).
Tatusov, R. L. et al. The COG database: an updated version includes eukaryotes. BMC Bioinformatics 4, 41 (2003).
Dimmer, E. C. et al. The UniProt-GO annotation database in 2011. Nucleic Acids Res. 40, D565–D570 (2012).
Bateman, A. et al. The Pfam protein families database. Nucleic Acids Res. 32, D138–D141 (2004).
Kanehisa, M. & Goto, S. KEGG: kyoto encyclopedia of genes and genomes. Nucleic Acids Res. 28, 27–30 (2000).
Zhu, T. et al. Sequencing a Juglans regia× J. microcarpa hybrid yields high-quality genome assemblies of parental species. Hortic. Res. 6, 55 (2019).
Ding, Y. M. et al. Genome structure-based Juglandaceae phylogenies contradict alignment-based phylogenies and substitution rates vary with DNA repair genes. Nat. Commun. 14, 617 (2023).
Lowe, T. M. & Eddy, S. R. tRNAscan-SE: a program for improved detection of transfer RNA genes in genomic sequence. Nucleic Acids Res. 25, 955–964 (1997).
Seeman, T. Source code for: tseemann/barrnap: bacterial ribosomal RNA predictor (0.9-2). https://github.com/tseemann/barrnap (2018).
Nawrocki, E. P., Kolbe, D. L. & Eddy, S. R. Infernal 1.0: inference of RNA alignments. Bioinformatics 25, 1335–1337 (2009).
Nawrocki, E. P. et al. Rfam 12.0: updates to the RNA families database. Nucleic Acids Res. 43, D130–D137 (2015).
Chen, C. et al. TBtools: an integrative toolkit developed for interactive analyses of big biological data. Mol. Plant 13, 1194–1202 (2020).
Goel, M., Sun, H., Jiao, W. B. & Schneeberger, K. SyRI: finding genomic rearrangements and local sequence differences from whole-genome assemblies. Genome Biol. 20, 277 (2019).
Han, L. Q. et al. A haplotype-resolved reference genome provides insight into allele-specific expression in wild walnut (Juglans regia L.). figshare https://doi.org/10.6084/m9.figshare.23692980 (2024).
Wang, Y. et al. MCScanX: a toolkit for detection and evolutionary analysis of gene synteny and collinearity. Nucleic Acids Res. 40, e49–e49 (2012).
Altschul, S. F., Gish, W., Miller, W., Myers, E. W. & Lipman, D. J. Basic local alignment search tool. J. Mol. Biol. 215, 403–410 (1990).
Kim, D., Paggi, J. M., Park, C., Bennett, C. & Salzberg, S. L. Graph-based genome alignment and genotyping with HISAT2 and HISAT-genotype. Nat. Biotechnol. 37, 907–915 (2019).
Pertea, M., Kim, D., Pertea, G. M., Leek, J. T. & Salzberg, S. L. Transcript-level expression analysis of RNA-seq experiments with HISAT, StringTie and Ballgown. Nat. Protoc. 11, 1650–1667 (2016).
Love, M. I., Huber, W. & Anders, S. Moderated estimation of fold change and dispersion for RNA-seq data with DESeq. 2. Genome Biol. 15, 550 (2014).
NCBI Sequence Read Archive https://identifiers.org/ncbi/insdc.sra:SRP386543 (2022).
NCBI Sequence Read Archive https://identifiers.org/ncbi/insdc.sra:SRP386695 (2022).
NCBI Sequence Read Archive https://identifiers.org/ncbi/insdc.sra:SRP386523 (2022).
NCBI Sequence Read Archive https://identifiers.org/ncbi/insdc.sra:SRP429004 (2023).
Han, L. Q. et al. A haplotype-resolved reference genome provides insight into allele-specific expression in wild walnut (Juglans regia L.). figshare https://doi.org/10.6084/m9.figshare.22266730 (2023).
Han, L. Q. et al. GenBank https://identifiers.org/ncbi/insdc.gca:GCA_034508915.1 (2023).
Han, L. Q. et al. GenBank https://identifiers.org/ncbi/insdc.gca:GCA_034509015.1 (2023).
NCBI Sequence Read Archive https://identifiers.org/ncbi/insdc.sra:SRP426360 (2023).
Wingett, S. et al. HiCUP: pipeline for mapping and processing Hi-C data. F1000 Res. 4, 1310–1310 (2015).
Wang, P. & Wang, F. A proposed metric set for evaluation of genome assembly quality. Trends Genet. 39, 175–186 (2023).
Guk, J. Y., Jang, M. J., Choi, J. W., Lee, Y. M. & Kim, S. De novo phasing resolves haplotype sequences in complex plant genomes. Plant Biotechnol. J. 20, 1031–1041 (2022).
Holley, G. et al. Ratatosk: hybrid error correction of long reads enables accurate variant calling and assembly. Genome Biol. 22, 28 (2021).
Zhu, Y. et al. Construction of a high-density genetic map using specific length amplified fragment markers and identification of a quantitative trait locus for anthracnose resistance in walnut (Juglans regia L.). BMC Genomics 16, 614 (2015).
Acknowledgements
This work was supported by the the Opening Foundation of Xinjiang Key Laboratory (XJTSGS-2023001), the National Natural Science Foundation of China (31960584/32060665), and the Xinjiang Production and Construction Corps’ Key Industry R & D Program of the Southern Xinjiang (2022DB022).
Author information
Authors and Affiliations
Contributions
Y.X. and K.M. conceived the study and supervised all parts of the project. Y.Z., L.H., N.L. and K.M. collected the samples. Y.Z. prepared walnut seedlings and extracted the genomic DNA, RNA and conducted sequencing. Y.X. and L.H. performed genome assembly related analysis and ASE analysis. Y.X. and K.M. wrote the manuscript. Y.X., X.L. and L.H. revised this manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Han, L., Luo, X., Zhao, Y. et al. A haplotype-resolved genome provides insight into allele-specific expression in wild walnut (Juglans regia L.). Sci Data 11, 278 (2024). https://doi.org/10.1038/s41597-024-03096-4
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41597-024-03096-4