Abstract
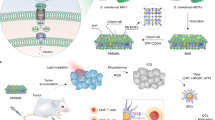
Microbial pathogens, including bacteria, fungi and viruses, can develop resistance to clinically used drugs; therefore, finding new therapeutic agents is an ongoing challenge. Recently, we reported the photoimmuno-antimicrobial strategy (PIAS), a type of photoimmunotechnology, that enables molecularly targeted elimination of a wide range of microbes, including the viral pathogen severe acute respiratory syndrome coronavirus 2 and the multidrug-resistant bacterial pathogen methicillin-resistant Staphylococcus aureus (MRSA). PIAS works in the same way as photoimmunotherapy (PIT), which has been used to treat recurrent head and neck cancer in Japan since 2020. Both PIAS and PIT use a monoclonal antibody conjugated to a phthalocyanine derivative dye that undergoes a shape change when photoactivated. This shape change induces a structural change in the antibody–dye conjugate, resulting in physical stress within the binding sites of the conjugate and disrupting them. Therefore, targeting accuracy and flexibility can be determined based on the specificity of the antibody used. In this protocol, we describe how to design a treatment strategy, label monoclonal antibodies with the dye and characterize the products. We provide detailed examples of how to set up and perform PIAS and PIT applications in vitro and in vivo. These examples are PIAS against microbes using MRSA as a representative subject, PIAS against viruses using severe acute respiratory syndrome coronavirus 2 in VeroE6/TMPRSS2 cells, PIAS against MRSA-infected animals, and in vitro and in vivo PIT against cancer cells. The in vitro and in vivo protocols can be completed in ~3 h and 2 weeks, respectively.
Key points
-
Photoimmunotherapy, an effective treatment for head and neck cancer, is a technique that uses a monoclonal antibody labeled with a phthalocyanine derivative to disrupt the binding site after photoactivation.
-
This technique allows for both ‘target specificity’ and ‘flexibility in target selection’, leading to the development of a novel antimicrobial strategy that enables targeted elimination of microbes, regardless of the target species or drug resistance status of the target.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout







Similar content being viewed by others
Data availability
Data of this study are available from the corresponding authors upon reasonable request. Source data are provided with this paper.
References
Levy, S. B. & Marshall, B. Antibacterial resistance worldwide: causes, challenges and responses. Nat. Med. 10, S122–S129 (2004).
Antimicrobial Resistance Collaborators. Global burden of bacterial antimicrobial resistance in 2019: a systematic analysis. Lancet 99, 629–655 (2022).
Chambers, H. F. & Deleo, F. R. Waves of resistance: Staphylococcus aureus in the antibiotic era. Nat. Rev. Microbiol. 7, 629–641 (2009).
Kluytmans, J., van Belkum, A. & Verbrugh, H. Nasal carriage of Staphylococcus aureus: epidemiology, underlying mechanisms, and associated risks. Clin. Microbiol. Rev. 10, 505–520 (1997).
Iwase, T. et al. Staphylococcus epidermidis Esp inhibits Staphylococcus aureus biofilm formation and nasal colonization. Nature 465, 346–349 (2010).
Guan, W. J. et al. China medical treatment expert group for COVID-19. Clinical characteristics of coronavirus disease 2019 in China. N. Engl. J. Med. 382, 1708–1720 (2020).
Laxminarayan, R. et al. Access to effective antimicrobials: a worldwide challenge. Lancet 387, 168–175 (2016).
Coates, A., Hu, Y., Bax, R. & Page, C. The future challenges facing the development of new antimicrobial drugs. Nat. Rev. Drug Discov. 1, 895–910 (2002).
Mitsunaga, M. et al. Cancer cell-selective in vivo near infrared photoimmunotherapy targeting specific membrane molecules. Nat. Med. 17, 1685–1691 (2011).
Mitsunaga, M. et al. Antimicrobial strategy for targeted elimination of different microbes, including bacterial, fungal and viral pathogens. Commun. Biol. 5, 647 (2022).
Mitsunaga, M. et al. A flexible target-specific anti-infection therapeutic platform that can be applied to different microbial species. Preprint at bioRxiv https://doi.org/10.1101/2020.04.20.049791 (2020).
Sato, K. et al. Photoinduced ligand release from a silicon phthalocyanine dye conjugated with monoclonal antibodies: a mechanism of cancer cell cytotoxicity after near-infrared photoimmuno therapy. ACS Cent. Sci. 4, 1559–1569 (2018).
Kato, T. et al. Electron donors rather than reactive oxygen species needed for therapeutic photochemical reaction of near-infrared photoimmunotherapy. ACS Pharmacol. Transl. Sci. 4, 1689–1701 (2021).
Hatayama, Y. et al. Development of a monoclonal antibody targeting HTLV-1 envelope gp46 glycoprotein and its application to near-infrared photoimmuno-antimicrobial strategy. Viruses 14, 2153 (2022).
Yasui, H. et al. Near infrared photo-antimicrobial targeting therapy for Candida albicans. Adv. Ther. 4, 2000221 (2021).
Huang, M., Shen, A., Ding, J. & Geng, M. Molecularly targeted cancer therapy: some lessons from the past decade. Trends Pharmacol. Sci. 35, 41–50 (2014).
Scott, A., Wolchok, J. & Old, L. Antibody therapy of cancer. Nat. Rev. Cancer 12, 278–287 (2012).
Loibl, S. & Gianni, L. HER2-positive breast cancer. Lancet 389, 2415–2429 (2017).
Hughes, J. P., Rees, S., Kalindjian, S. B. & Philpott, K. L. Principles of early drug discovery. Br. J. Pharmacol. 162, 1239–1249 (2011).
Arora, A. & Scholar, E. M. Role of tyrosine kinase inhibitors in cancer therapy. J. Pharmacol. Exp. Ther. 315, 971–979 (2005).
Matsuyama, S. et al. Enhanced isolation of SARS-CoV-2 by TMPRSS2-expressing cells. Proc. Natl Acad. Sci. USA 117, 7001–7003 (2020).
Kokai-Kun, J. F. The cotton rat as a model for Staphylococcus aureus nasal colonization in humans: cotton rat S. aureus nasal colonization model. Methods Mol. Biol. 431, 241–254 (2008).
Mitsunaga, M. et al. Immediate in vivo target-specific cancer cell death after near infrared photoimmunotherapy. BMC Cancer 12, 345 (2012).
Nakajima, T., Sano, K., Mitsunaga, M., Choyke, P. L. & Kobayashi, H. Real-time monitoring of in vivo acute necrotic cancer cell death induced by near infrared photoimmunotherapy using fluorescence lifetime imaging. Cancer Res. 72, 4622–4628 (2012).
Sano, K., Mitsunaga, M., Nakajima, T., Choyke, P. L. & Kobayashi, H. Acute cytotoxic effects of photoimmunotherapy assessed by 18F-fluorodeoxyglucose (FDG) PET. J. Nucl. Med. 54, 770–775 (2013).
Kosaka, N. et al. Semiquantitative assessment of the microdistribution of fluorescence- labeled monoclonal antibody in small peritoneal disseminations of ovarian cancer. Cancer Sci. 101, 820–825 (2010).
Kosaka, N. et al. Microdistribution of fluorescently-labeled monoclonal antibody in a peritoneal dissemination model of ovarian cancer. Proc. Spie. 7576, 7576041–7576049 (2010).
Mitsunaga, M., Nakajima, T., Sano, K., Choyke, P. L. & Kobayashi, H. Near-infrared theranostic photoimmunotherapy (PIT): repeated exposure of light enhances the effect of immunoconjugate. Bioconjugate Chem. 23, 604–609 (2012).
Acknowledgements
We acknowledge the staff at the National Institute of Infectious Diseases for providing SC isolate (EPI_ISL_408667) and thank JCRB Cell Bank in Japan for VeroE6/TMPRSS2 cells. We also thank Y. Hatayama and T. Morita for viral experiments, Y. Dan and H. Yamada (Hitachi High-Technologies), H. Saito and T. Tachibana (Jikei University) for preparing and acquiring SEM images, H. Ishizaka, and colleagues in our laboratory for technical support, and H. Kanuka, Y. Kinjo, Y. Manome, H. Tajiri, M. Saruta and A. Chiba (Jikei University) for their support and comments. We also thank the University of Alabama at Birmingham Bacterial Pathogenesis & Physiology Journal Club for comments on PIAS study. This study was partly supported by AMED (grant number: JP22fk0108611h0802 to T.I.), JSPS KAKENHI (grant numbers: JP22K07076 to T.I.; 21H03811, 18H03523, 26710010 and JP26670487 to M.M.; and JP17K16233 to K.I.), Takeda Science Foundation to M.M., and Intramural Research Program of the National Institutes of Health, National Cancer Institute, Center for Cancer Research grant number: ZIA BC 011513 to H.K. and The Jikei University Research Fund to M.M. and T.I.
Author information
Authors and Affiliations
Contributions
T.I., H.K. and M.M. designed the study, performed experiments and analyses, and wrote the manuscript. K.I. and T.N. performed the experiments and analyses. A.R. and K.M. contributed to the viral study. All authors commented critically on the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Protocols thanks Tayyaba Hasan, Mingdong Huang and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Related links
Key references using this protocol
Mitsunaga, M. et al. Commun. Biol. 5, 647 (2022): https://www.nature.com/articles/s42003-022-03586-4
Mitsunaga, M. et al. Nat. Med. 17, 1685–1691 (2011): https://www.nature.com/articles/nm.2554
Supplementary information
Supplementary Information
Supplementary Figs. 1 and 2.
Source data
Source Data Fig. 7
Source data for Fig. 7b.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Iwase, T., Ito, K., Nishimura, T. et al. Photoimmunotechnology as a powerful biological tool for molecular-based elimination of target cells and microbes, including bacteria, fungi and viruses. Nat Protoc 18, 3390–3412 (2023). https://doi.org/10.1038/s41596-023-00874-z
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41596-023-00874-z
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.