Abstract
Cerebral arachnoid cysts (ACs) are one of the most common and poorly understood types of developmental brain lesion. To begin to elucidate AC pathogenesis, we performed an integrated analysis of 617 patient–parent (trio) exomes, 152,898 human brain and mouse meningeal single-cell RNA sequencing transcriptomes and natural language processing data of patient medical records. We found that damaging de novo variants (DNVs) were highly enriched in patients with ACs compared with healthy individuals (P = 1.57 × 10−33). Seven genes harbored an exome-wide significant DNV burden. AC-associated genes were enriched for chromatin modifiers and converged in midgestational transcription networks essential for neural and meningeal development. Unsupervised clustering of patient phenotypes identified four AC subtypes and clinical severity correlated with the presence of a damaging DNV. These data provide insights into the coordinated regulation of brain and meningeal development and implicate epigenomic dysregulation due to DNVs in AC pathogenesis. Our results provide a preliminary indication that, in the appropriate clinical context, ACs may be considered radiographic harbingers of neurodevelopmental pathology warranting genetic testing and neurobehavioral follow-up. These data highlight the utility of a systems-level, multiomics approach to elucidate sporadic structural brain disease.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout



Similar content being viewed by others
Data availability
The sequencing data for all AC parent–offspring trios and singletons from the healthcare-acquired cohort have been deposited in the NCBI database of Genotypes and Phenotypes and AnVIL (https://anvilproject.org/data/studies/phs000744/) under the accession number phs000744.v4.p2. Patients referred to GeneDx are consented to aggregate, de-identified research and subject to US Health Insurance Portability and Accountability Act (HIPAA) privacy protections. The patient-level alignment, phenotypic and variant call data for the GeneDx cohort cannot be shared without a HIPAA Business Associate Agreement. Access to the de-identified, aggregate data used in this analysis is available upon request to GeneDx, provided that a HIPAA Business Associate Agreement is established. Under those conditions, researchers can request the de-identified, aggregate data from GeneDx by contacting smcgee@genedx.com and can expect to receive the requested data within approximately 26 weeks.
Code availability
The software utilized in this study is available at the following web addresses: SAMtools version 1.3.1 (https://github.com/samtools/samtools); GATK HaplotypeCaller version 3.7.0 (https://github.com/broadinstitute/gatk/releases); GATK GenotypeGVCFs version 3.7.0 (https://github.com/broadinstitute/gatk/releases); GATK VariantRecalibrator version 3.7.0 (https://github.com/broadinstitute/gatk/releases); TrioDeNovo version 0.6.0 (http://genome.sph.umich.edu/wiki/Triodenovo); denovolyzeR version 0.2.0 (http://denovolyzer.org); DeNovoWEST 42 version 1.0.0 (https://github.com/queenjobo/DeNovoWEST); PLINK version 1.9 (http://pngu.mgh.harvard.edu/~purcell/plink); MetaSVM/cadd13/ANNOVAR version 4.2 (http://annovar.openbioinformatics.org); R version 3.5.0 (https://www.r-project.org/); Python version 2.7 (https://www.python.org/downloads/); EIGENSTRAT version 7.2.1 (https://github.com/DReichLab/EIG/tree/master/EIGENSTRAT); DMLE+ version 2.3 (http://dmle.org/); enrichR R package version 3.0 (https://cran.r-project.org/web/packages/enrichR/index.html); GOrilla (http://cbl-gorilla.cs.technion.ac.il/); QIAGEN December 2021 release (http://www.ingenuity.com); txt2hpo version 0.2.3 (https://github.com/GeneDx/txt2hpo); phenopy version 0.3.0 (https://github.com/GeneDx/phenopy); Monocle R package version 3 (https://cole-trapnell-lab.github.io/monocle3/); and disgenet2r R package version 0.0.9 (https://www.disgenet.org/static/disgenet2r/disgenet2r.html). Our in-house pipelines and codes are available at https://github.com/Kahle-Lab/Arachnoid-Cyst.
References
White, T., Su, S., Schmidt, M., Kao, C. Y. & Sapiro, G. The development of gyrification in childhood and adolescence. Brain Cogn. 72, 36–45 (2010).
Juric-Sekhar, G. & Hevner, R. F. Malformations of cerebral cortex development: molecules and mechanisms. Annu. Rev. Pathol. 14, 293–318 (2019).
Siegenthaler, J. A. et al. Retinoic acid from the meninges regulates cortical neuron generation. Cell 139, 597–609 (2009).
Borrell, V. & Marin, O. Meninges control tangential migration of hem-derived Cajal–Retzius cells via CXCL12/CXCR4 signaling. Nat. Neurosci. 9, 1284–1293 (2006).
Al-Holou, W. N. et al. Prevalence and natural history of arachnoid cysts in adults. J. Neurosurg. 118, 222–231 (2013).
Mustansir, F., Bashir, S. & Darbar, A. Management of arachnoid cysts: a comprehensive review. Cureus 10, e2458 (2018).
De Keersmaecker, B. et al. Outcome of 12 antenatally diagnosed fetal arachnoid cysts: case series and review of the literature. Eur. J. Paediatr. Neurol. 19, 114–121 (2015).
Katzman, G. L., Dagher, A. P. & Patronas, N. J. Incidental findings on brain magnetic resonance imaging from 1000 asymptomatic volunteers. J. Am. Med. Assoc. 282, 36–39 (1999).
Hayes, M. J., TerMaath, S. C., Crook, T. R. & Killeffer, J. A. A review on the effectiveness of surgical intervention for symptomatic intracranial arachnoid cysts in adults. World Neurosurg. 123, e259–e272 (2019).
Jafrani, R., Raskin, J. S., Kaufman, A. & Lam, S. Intracranial arachnoid cysts: pediatric neurosurgery update. Surg. Neurol. Int. 10, 15 (2019).
Choi, J. U. & Kim, D. S. Pathogenesis of arachnoid cyst: congenital or traumatic? Pediatr. Neurosurg. 29, 260–266 (1998).
Starkman, S. P., Brown, T. C. & Linell, E. A. Cerebral arachnoid cysts. J. Neuropathol. Exp. Neurol. 17, 484–500 (1958).
Zeegers, M. et al. Radiological findings in autistic and developmentally delayed children. Brain Dev. 28, 495–499 (2006).
Nikolic, I. et al. The association of arachnoid cysts and focal epilepsy: hospital based case control study. Clin. Neurol. Neurosurg. 159, 39–41 (2017).
Al-Holou, W. N. et al. Prevalence and natural history of arachnoid cysts in children. J. Neurosurg. Pediatr. 5, 578–585 (2010).
Wiener, S. N., Pearlstein, A. E. & Eiber, A. MR imaging of intracranial arachnoid cysts. J. Comput. Assist. Tomogr. 11, 236–241 (1987).
Gosalakkal, J. A. Intracranial arachnoid cysts in children: a review of pathogenesis, clinical features, and management. Pediatr. Neurol. 26, 93–98 (2002).
Hall, S. et al. Clinical and radiological outcomes following surgical treatment for intra-cranial arachnoid cysts. Clin. Neurol. Neurosurg. 177, 42–46 (2019).
Cilluffo, J. M., Gomez, M. R., Reese, D. F., Onofrio, B. M. & Miller, R. H. Idiopathic (“congenital”) spinal arachnoid diverticula. Clinical diagnosis and surgical results. Mayo Clin. Proc. 56, 93–101 (1981).
Zafeiriou, D. I. & Batzios, S. P. Brain and spinal MR imaging findings in mucopolysaccharidoses: a review. AJNR Am. J. Neuroradiol. 34, 5–13 (2013).
Qureshi, H. M. et al. Familial and syndromic forms of arachnoid cyst implicate genetic factors in disease pathogenesis. Cereb. Cortex 18, bhac257 (2022).
Furey, C. G. et al. De novo mutation in genes regulating neural stem cell fate in human congenital hydrocephalus. Neuron 99, 302–314.e4 (2018).
Jin, S. C. et al. Exome sequencing implicates genetic disruption of prenatal neuro-gliogenesis in sporadic congenital hydrocephalus. Nat. Med. 26, 1754–1765 (2020).
Bilgüvar, K. et al. Whole-exome sequencing identifies recessive WDR62 mutations in severe brain malformations. Nature 467, 207–210 (2010).
Barak, T. et al. Recessive LAMC3 mutations cause malformations of occipital cortical development. Nat. Genet. 43, 590–594 (2011).
Mishra-Gorur, K. et al. Mutations in KATNB1 cause complex cerebral malformations by disrupting asymmetrically dividing neural progenitors. Neuron 84, 1226–1239 (2014).
Kundishora, A. J. et al. DIAPH1 variants in non-East Asian patients with sporadic moyamoya disease. JAMA Neurol. 78, 993–1003 (2021).
De Ligt, J. et al. Diagnostic exome sequencing in persons with severe intellectual disability. N. Engl. J. Med. 367, 1921–1929 (2012).
Neale, B. M. et al. Patterns and rates of exonic de novo mutations in autism spectrum disorders. Nature 485, 242–245 (2012).
Timberlake, A. T. et al. Two locus inheritance of non-syndromic midline craniosynostosis via rare SMAD6 and common BMP2 alleles. eLife 5, e20125 (2016).
Krumm, N. et al. Excess of rare, inherited truncating mutations in autism. Nat. Genet. 47, 582–588 (2015).
Shi, C. et al. Down-regulation of the forkhead transcription factor Foxp1 is required for monocyte differentiation and macrophage function. Blood 112, 4699–4711 (2008).
Li, X. et al. MEK is a key regulator of gliogenesis in the developing brain. Neuron 75, 1035–1050 (2012).
Chakraborty, R. et al. Mutually exclusive recurrent somatic mutations in MAP2K1 and BRAF support a central role for ERK activation in LCH pathogenesis. Blood 124, 3007–3015 (2014).
Aoidi, R. et al. Mek1Y130C mice recapitulate aspects of human cardio-facio-cutaneous syndrome. Dis. Model Mech. 11, dmm031278 (2018).
Nie, Z. et al. A specificity and targeting subunit of a human SWI/SNF family-related chromatin-remodeling complex. Mol. Cell. Biol. 20, 8879–8888 (2000).
Tumber, A. et al. Potent and selective KDM5 inhibitor stops cellular demethylation of H3K4me3 at transcription start sites and proliferation of MM1S myeloma cells. Cell Chem. Biol. 24, 371–380 (2017).
Stuart, J. M., Segal, E., Koller, D. & Kim, S. K. A gene-coexpression network for global discovery of conserved genetic modules. Science 302, 249–255 (2003).
Castro Dias, M., Mapunda, J. A., Vladymyrov, M. & Engelhardt, B. Structure and junctional complexes of endothelial, epithelial and glial brain barriers. Int. J. Mol. Sci. 20, 5372 (2019).
Pinero, J. et al. DisGeNET: a comprehensive platform integrating information on human disease-associated genes and variants. Nucleic Acids Res. 45, D833–D839 (2017).
Arriola, G., de Castro, P. & Verdu, A. Familial arachnoid cysts. Pediatr. Neurol. 33, 146–148 (2005).
Martinez, J. O. et al. Intracranial arachnoid cysts and epilepsy in children: should this be treated surgically? Our 29-year experience and review of the literature. Neurocirugía 33, 157–164 (2021).
Valencia, A. M. & Pasca, S. P. Chromatin dynamics in human brain development and disease. Trends Cell Biol. 32, 98–101 (2022).
Sokpor, G., Xie, Y., Rosenbusch, J. & Tuoc, T. Chromatin remodeling BAF (SWI/SNF) complexes in neural development and disorders. Front. Mol. Neurosci. 10, 243 (2017).
Eissenberg, J. C. & Shilatifard, A. Histone H3 lysine 4 (H3K4) methylation in development and differentiation. Dev. Biol. 339, 240–249 (2010).
Bragin, E. et al. DECIPHER: database for the interpretation of phenotype-linked plausibly pathogenic sequence and copy-number variation. Nucleic Acids Res. 42, D993–D1000 (2014).
De Rubeis, S. et al. Synaptic, transcriptional and chromatin genes disrupted in autism. Nature 515, 209–215 (2014).
Zaidi, S. et al. De novo mutations in histone-modifying genes in congenital heart disease. Nature 498, 220–223 (2013).
Kadoch, C. & Crabtree, G. R. Mammalian SWI/SNF chromatin remodeling complexes and cancer: mechanistic insights gained from human genomics. Sci. Adv. 1, e1500447 (2015).
Rylaarsdam, L. & Guemez-Gamboa, A. Genetic causes and modifiers of autism spectrum disorder. Front. Cell Neurosci. 13, 385 (2019).
Jahed, Z., Shams, H., Mehrbod, M. & Mofrad, M. R. Mechanotransduction pathways linking the extracellular matrix to the nucleus. Int. Rev. Cell Mol. Biol. 310, 171–220 (2014).
Rengachary, S. S. & Watanabe, I. Ultrastructure and pathogenesis of intracranial arachnoid cysts. J. Neuropathol. Exp. Neurol. 40, 61–83 (1981).
Kanton, S. et al. Organoid single-cell genomic atlas uncovers human-specific features of brain development. Nature 574, 418–422 (2019).
Rabiei, K., Hogfeldt, M. J., Doria-Medina, R. & Tisell, M. Surgery for intracranial arachnoid cysts in children—a prospective long-term study. Childs Nerv. Syst. 32, 1257–1263 (2016).
Tamburrini, G., Dal Fabbro, M., & Di Rocco, C. Sylvian fissure arachnoid cysts: a survey on their diagnostic workout and practical management. Childs Nerv. Syst. 24, 593–604 (2008).
Schulz, M. et al. Surgical management of intracranial arachnoid cysts in pediatric patients: radiological and clinical outcome. J. Neurosurg. Pediatr. 28, 102–112 (2021).
Sadler, B. et al. Rare and de novo coding variants in chromodomain genes in Chiari I malformation. Am. J. Hum. Genet. 108, 100–114 (2021).
Duran, D. et al. Mutations in chromatin modifier and ephrin signaling genes in vein of galen malformation. Neuron 101, 429–443.e4 (2019).
Timberlake, A. T. et al. Genetic influence on neurodevelopment in nonsyndromic craniosynostosis. Plast. Reconstr. Surg. 149, 1157–1165 (2022).
Retterer, K. et al. Clinical application of whole-exome sequencing across clinical indications. Genet. Med. 18, 696–704 (2016).
McKenna, A. et al. The Genome Analysis Toolkit: a MapReduce framework for analyzing next-generation DNA sequencing data. Genome Res. 20, 1297–1303 (2010).
Karczewski, K. J. et al. The mutational constraint spectrum quantified from variation in 141,456 humans. Nature 581, 434–443 (2020).
Taliun, D. et al. Sequencing of 53,831 diverse genomes from the NHLBI TOPMed program. Nature 590, 290–299 (2021).
Mills, R. E. et al. Natural genetic variation caused by small insertions and deletions in the human genome. Genome Res. 21, 830–839 (2011).
Kaplanis, J. et al. Evidence for 28 genetic disorders discovered by combining healthcare and research data. Nature 586, 757–762 (2020).
Ware, J. S., Samocha, K. E., Homsy, J. & Daly, M. J. Interpreting de novo variation in human disease using denovolyzeR. Curr. Protoc. Hum. Genet. 87, 7.25.1–7.25.15 (2015).
Purcell, S. et al. PLINK: a tool set for whole-genome association and population-based linkage analyses. Am. J. Hum. Genet. 81, 559–575 (2007).
Lango Allen, H. et al. Hundreds of variants clustered in genomic loci and biological pathways affect human height. Nature 467, 832–838 (2010).
Grove, J. et al. Identification of common genetic risk variants for autism spectrum disorder. Nat. Genet. 51, 431–444 (2019).
Jin, S. C. et al. Contribution of rare inherited and de novo variants in 2,871 congenital heart disease probands. Nat. Genet. 49, 1593–1601 (2017).
Song, L. et al. STAB: a spatio-temporal cell atlas of the human brain. Nucleic Acids Res. 49, D1029–D1037 (2021).
Zhu, Y. et al. Spatiotemporal transcriptomic divergence across human and macaque brain development. Science 362, eaat8077 (2018).
Langfelder, P. & Horvath, S. WGCNA: an R package for weighted correlation network analysis. BMC Bioinformatics 9, 559 (2008).
Walker, R. L. et al. Genetic control of expression and splicing in developing human brain informs disease mechanisms. Cell 179, 750–771.e22 (2019).
Kuleshov, M. V. et al. Enrichr: a comprehensive gene set enrichment analysis web server 2016 update. Nucleic Acids Res. 44, W90–W97 (2016).
Eden, E., Navon, R., Steinfeld, I., Lipson, D. & Yakhini, Z. GOrilla: a tool for discovery and visualization of enriched GO terms in ranked gene lists. BMC Bioinformatics 10, 48 (2009).
Kramer, A., Green, J., Pollard, J. Jr. & Tugendreich, S. Causal analysis approaches in Ingenuity Pathway Analysis. Bioinformatics 30, 523–530 (2014).
DeSisto, J. et al. Single-cell transcriptomic analyses of the developing meninges reveal meningeal fibroblast diversity and function. Dev. Cell 54, 43–59.e4 (2020).
Cao, J. et al. The single-cell transcriptional landscape of mammalian organogenesis. Nature 566, 496–502 (2019).
Campello, R. J. G. B., Moulavi, D. & Sander, J. in Advances in Knowledge Discovery and Data Mining (eds. Pei, J. et al.) 160–172 (Springer Berlin Heidelberg, 2013).
Acknowledgements
We are grateful to the patients and families who participated in this research for their invaluable role in this study. This work is supported by the Yale–National Institutes of Health (NIH) Center for Mendelian Genomics (5U54HG006504); R01 NS111029-01A1, R01 NS109358, K12 228168 and the Rudi Schulte Research Institute (to K.T.K.); the NIH Medical Scientist Training Program (NIH/National Institute of General Medical Sciences grant T32GM007205); an NIH Clinical and Translational Science Award from the National Center for Advancing Translational Sciences (TL1 TR001864); the K99/R00 Pathway to Independence Award R00HL143036 (to S.C.J.); the Children’s Discovery Institute Faculty Scholar award CDI-FR-2021-926 (to S.C.J.); the Vernon W. Lippard Research Fellowship; and the Howard Hughes Medical Institute.
Author information
Authors and Affiliations
Contributions
A.J.K. and K.T.K. designed and conceptualized the study. A.J.K., G.A., S. McGee, K.Y.M., V.G., E.K., P.Q.D., H.S., J.O., J.S., A.A., M.L.D., C.G.F., A.T.T., H.M.Q., A.A.E., B.S.C., M.G., R.P.L., F.M., R.I.T., S.C.J. and K.T.K. performed cohort ascertainment, recruitment and phenotypic characterization. I.R.T., C.C., F.L.-G. and S. Mane produced and validated the exome sequencing data. G.A., S. McGee, V.G., A.J.K., S.Z., Y.-C.W., A.T.T., J.R.K., P.-Y.F., W.D., F.M., R.I.T., S.C.J. and K.T.K. performed the exome sequencing analysis. G.A., E.K. and K.T.K. performed the integrative genomics analysis. A.J.K., S. McGee, K.Y.M., V.G., A.M.-D.-L. and K.T.K. performed the phenomics analysis. G.A., A.J.K., S.C.J. and W.D. performed the statistical analysis. C.N.-W. performed Sanger sequencing validation. A.J.K., A.M.-D.-L. and K.T.K. performed neuroimaging characterization. S.H. performed the biophysical simulation. C.N.-W., S. Mane, M.G., R.P.L, R.I.T., S.C.J. and K.T.K. provided resources. A.J.K., G.A., S. McGee, K.Y.M., E.K., S.L.A., M.G., R.P.L., F.M., R.I.T., S.C.J. and K.T.K. wrote and reviewed the manuscript. A.J.K., G.A., S. McGee, K.Y.M., R.I.T., S.C.J. and K.T.K. performed project administration. R.P.L., S.C.J. and K.T.K. acquired funding and supervised the project.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Medicine thanks Alan Shuldiner, Abhaya Kulkarni and the other, anonymous, reviewer(s) for their contribution to the peer review of this work. Primary Handling Editor: Anna Maria Ranzoni, in collaboration with the Nature Medicine team.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 Graphical summary of the methodological framework of the study.
Graphical summary of the methodological framework of the study.
Extended Data Fig. 2 De novo variation (DNV) rate closely approximated Poisson distribution in AC cases and controls.
The observed number of DNVs per subject (bars) compared to the numbers expected (lines) from the Poisson distribution in the case (red) and control (blue) cohorts. ‘p’ denotes chi-squared p-value. P-values determined by Chi-squared goodness of fit test, two sided. Not adjusted.
Extended Data Fig. 3 Quantile-quantile (Q-Q) plot comparing observed versus expected p-values.
(a) DeNovoWEST derived plots for de novo variants (DNVs) in each gene in 617 AC cases. ADNP, ARIDB1, KDM5C, PURA, FOXP1, and MAP2K1 exhibit exome-wide significant enrichment for all DNVs in AC cases. ARID1B, ADNP, and FOXP1 exhibit significant enrichment of loss-of-function (LoF) DNVs comprising premature termination, frameshift, or splice-site variants. KDM5C and MAP2K1 exhibit significant enrichment of missense variants. ARID1B, FOXP1, ADNP, and KDM5C exhibit significant enrichment of protein-altering variants, including missense and predictive LoF DNVs. ARID1B, ADNP, FOXP1, MAP2K1, PURA, and KDM5C exhibit significant enrichment of protein-damaging variants, including D-mis and LoF DNVs. There is no significant enrichment of synonymous DNVs among the 617 cases. Grey areas within graphs represents 95% confidence interval for expected values. (b) DenovolyzeR derived plots for DNVs in each gene in 617 AC cases. ARID1B, PURA, ADNP, and FOXP1 exhibit exome-wide significant enrichment for all DNVs in AC cases. ARID1B and ADNP exhibit significant enrichment of LoF DNVs. MAP2K1 exhibits significant enrichment of damaging-missense (D-mis) variants (MetaSVM = ‘D’ or MPC > 2 damaging missense). ARID1B, ADNP, FOXP1, MAP2K1, and KDM5C exhibit significant enrichment of protein-altering variants. ARID1B, ADNP, FOXP1, MAP2K1, and DDX3X exhibit significant enrichment of protein-damaging variants. There is no significant enrichment of tolerated-missense (T-mis) DNVs or synonymous DNVs among the 617 cases. The grey areas within graphs represents 95% confidence interval centered around the observed = expected line.
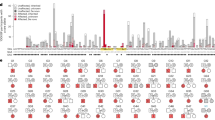
Extended Data Fig. 4 Phenomic heat map of traits identified in AC patients harboring de novo variants (DNVs) in exome-wide significant AC risk genes.
Subject phenotypes were determined by text2HPO natural language processing of medical record data (https://github.com/GeneDx/txt2hpo).
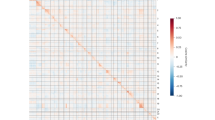
Extended Data Fig. 5 Integrative genomic findings within meningeal cell dataset.
(a) Enrichment of AC genes in meningeal gene modules. Numbers displayed exceed the Bonferroni-corrected statistical significance threshold tested by one sided Fisher’s exact test and are -log10(p-value). pAC: possible AC gene set; hcAC; high-confidence AC gene set; EWS exome-wide significant; Mod: module. (b) GOrilla and WikiPathways analyses of enriched arachnoid cell module 3. P-values determined by one sided Fisher’s exact test. Bonferroni-corrected significance threshold denoted by the vertical yellow line. Top terms displayed. (c) Enrichment of gene modules in specific meningeal cell types. P-values by one sided Fisher’s exact test. Modules in red have similar meningeal cell-type enrichment compared to AC risk gene meningeal cell-type enrichment. The red asterisk highlights significant enrichment (Bonferroni corrected) for cell types in the pAC gene set.
Supplementary information
Supplementary Information
Supplementary Figs. 1 and 2 and Tables 1–5.
Supplementary Table 6
Top 20 gene markers per cell cluster in the Spatio-Temporal Cell Atlas of the Human Brain dataset ranked by log2[fold change].
Supplementary Table 7
Top 20 gene markers per cell cluster in the the embryonic forebrain meningeal dataset ranked by log2[fold change].
Supplementary Table 8
Phenotype groupings of HPO terms.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Kundishora, A.J., Allington, G., McGee, S. et al. Multiomic analyses implicate a neurodevelopmental program in the pathogenesis of cerebral arachnoid cysts. Nat Med 29, 667–678 (2023). https://doi.org/10.1038/s41591-023-02238-2
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41591-023-02238-2
This article is cited by
-
An arachnoid cyst rupture complicated with subdural hygroma in a middle-aged woman: a case report and review of the literature
Egyptian Journal of Neurosurgery (2023)