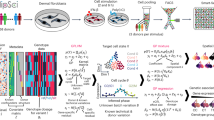
Abstract
Humans display remarkable interindividual variation in their immune response to identical challenges. Yet, our understanding of the genetic and epigenetic factors contributing to such variation remains limited. Here we performed in-depth genetic, epigenetic and transcriptional profiling on primary macrophages derived from individuals of European and African ancestry before and after infection with influenza A virus. We show that baseline epigenetic profiles are strongly predictive of the transcriptional response to influenza A virus across individuals. Quantitative trait locus (QTL) mapping revealed highly coordinated genetic effects on gene regulation, with many cis-acting genetic variants impacting concomitantly gene expression and multiple epigenetic marks. These data reveal that ancestry-associated differences in the epigenetic landscape can be genetically controlled, even more than gene expression. Lastly, among QTL variants that colocalized with immune-disease loci, only 7% were gene expression QTL, while the remaining genetic variants impact epigenetic marks, stressing the importance of considering molecular phenotypes beyond gene expression in disease-focused studies.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout







Similar content being viewed by others
Data availability
Sequence data have been deposited at the European Genome-Phenome Archive (EGA), under accession numbers EGAD00001008422 (RNA-seq, ATAC–seq and ChIPmentation) and EGAD00001008359 (WGS and WGBS). In addition, all data generated in this study are freely accessible via a custom web-based browser that enables easy querying and visualization of all the data generated (https://computationalgenomics.ca/tools/epivar). Full DNA methylation and QTL mapping results as well as inputs for analyses are available at https://zenodo.org/records/10108241 (ref. 94). Reagent and resource requests should be addressed and will be fulfilled by the lead contacts, L.B.B. (lbarreiro@uchicago.edu) and G.B. (guil.bourque@mcgill.ca).
Code availability
All original code is available at https://doi.org/10.5281/zenodo.10515250 (ref. 95). Code for ASE analysis can be found at https://doi.org/10.5281/zenodo.10511587 (ref. 96).
References
Brinkworth, J. F. & Barreiro, L. B. The contribution of natural selection to present-day susceptibility to chronic inflammatory and autoimmune disease. Curr. Opin. Immunol. 31, 66–78 (2014).
Duffy, D. et al. Functional analysis via standardized whole-blood stimulation systems defines the boundaries of a healthy immune response to complex stimuli. Immunity 40, 436–450 (2014).
Pennington, R., Gatenbee, C., Kennedy, B., Harpending, H. & Cochran, G. Group differences in proneness to inflammation. Infect. Genet. Evol. 9, 1371–1380 (2009).
Bakker, O. B. et al. Integration of multi-omics data and deep phenotyping enables prediction of cytokine responses. Nat. Immunol. 19, 776–786 (2018).
Nédélec, Y. et al. Genetic ancestry and natural selection drive population differences in immune responses to pathogens. Cell 167, 657–669.e21 (2016).
Piasecka, B. et al. Distinctive roles of age, sex, and genetics in shaping transcriptional variation of human immune responses to microbial challenges. Proc. Natl Acad. Sci. USA 115, E488–E497 (2018).
Quach, H. et al. Genetic adaptation and Neandertal admixture shaped the immune system of human populations. Cell 167, 643–656.e17 (2016).
Randolph, H. E. et al. Genetic ancestry effects on the response to viral infection are pervasive but cell type specific. Science 374, 1127–1133 (2021).
Barreiro, L. B. et al. Deciphering the genetic architecture of variation in the immune response to Mycobacterium tuberculosis infection. Proc. Natl Acad. Sci. USA 109, 1204–1209 (2012).
Schirmer, M. et al. Linking the human gut microbiome to inflammatory cytokine production capacity. Cell 167, 1125–1136.e8 (2016).
Snyder-Mackler, N. et al. Social status alters immune regulation and response to infection in macaques. Science 354, 1041–1045 (2016).
Snyder-Mackler, N. et al. Social determinants of health and survival in humans and other animals. Science 368, eaax9553 (2020).
Cole, S. W. Human social genomics. PLoS Genet. 10, e1004601 (2014).
Bekkering, S., Dominguez-Andres, J., Joosten, L. A. B., Riksen, N. P. & Netea, M. G. Trained immunity: reprogramming innate immunity in health and disease. Annu. Rev. Immunol. 39, 667–693 (2021).
Zhang, Q. & Cao, X. Epigenetic remodeling in innate immunity and inflammation. Annu. Rev. Immunol. 39, 279–311 (2021).
Chen, L. et al. Genetic drivers of epigenetic and transcriptional variation in human immune cells. Cell 167, 1398–1414.e24 (2016).
Degner, J. F. et al. DNase I sensitivity QTLs are a major determinant of human expression variation. Nature 482, 390–394 (2012).
Carja, O. et al. Worldwide patterns of human epigenetic variation. Nat. Ecol. Evol. 1, 1577–1583 (2017).
Husquin, L. T. et al. Exploring the genetic basis of human population differences in DNA methylation and their causal impact on immune gene regulation. Genome Biol. 19, 222 (2018).
Kasowski, M. et al. Extensive variation in chromatin states across humans. Science 342, 750–752 (2013).
McVicker, G. et al. Identification of genetic variants that affect histone modifications in human cells. Science 342, 747–749 (2013).
Waszak, S. M. et al. Population variation and genetic control of modular chromatin architecture in humans. Cell 162, 1039–1050 (2015).
The ENCODE Project Consortium. Identification and analysis of functional elements in 1% of the human genome by the ENCODE pilot project. Nature 447, 799–816 (2007).
The ENCODE Project Consortium. An integrated encyclopedia of DNA elements in the human genome. Nature 489, 57–74 (2012).
Roadmap Epigenomics Consortium. Integrative analysis of 111 reference human epigenomes. Nature 518, 317–329 (2015).
Fernández, J. M. et al. The BLUEPRINT data analysis portal. Cell Syst. 3, 491–495.e5 (2016).
Killip, M. J., Fodor, E. & Randall, R. E. Influenza virus activation of the interferon system. Virus Res. 209, 11–22 (2015).
Ciancanelli, M. J., Abel, L., Zhang, S.-Y. & Casanova, J.-L. Host genetics of severe influenza: from mouse Mx1 to human IRF7. Curr. Opin. Immunol. 38, 109–120 (2016).
Froggatt, H. M., Harding, A. T., Chaparian, R. R. & Heaton, N. S. ETV7 limits antiviral gene expression and control of influenza viruses. Sci. Signal. 14, eabe1194 (2021).
Pezzè, L. et al. ETV7 regulates breast cancer stem-like cell features by repressing IFN-response genes. Cell Death Dis. 12, 742 (2021).
Alexander, D. H., Novembre, J. & Lange, K. Fast model-based estimation of ancestry in unrelated individuals. Genome Res. 19, 1655–1664 (2009).
The 1000 Genomes Project Consortium. A global reference for human genetic variation. Nature 526, 68–74 (2015).
Storey, J. D. & Tibshirani, R. Statistical significance for genomewide studies. Proc. Natl Acad. Sci. 100, 9440–9445 (2003).
Urbut, S. M., Wang, G., Carbonetto, P. & Stephens, M. Flexible statistical methods for estimating and testing effects in genomic studies with multiple conditions. Nat. Genet. 51, 187–195 (2019).
Maples, B. K., Gravel, S., Kenny, E. E. & Bustamante, C. D. RFMix: a discriminative modeling approach for rapid and robust local-ancestry inference. Am. J. Hum. Genet. 93, 278–288 (2013).
Liberzon, A. et al. The Molecular Signatures Database Hallmark Gene Set Collection. Cell Syst. 1, 417–425 (2015).
Banovich, N. E. et al. Methylation QTLs are associated with coordinated changes in transcription factor binding, histone modifications, and gene expression levels. PLoS Genet. 10, e1004663 (2014).
Huan, T. et al. Genome-wide identification of DNA methylation QTLs in whole blood highlights pathways for cardiovascular disease. Nat. Commun. 10, 4267 (2019).
Ellegren, H. Microsatellites: simple sequences with complex evolution. Nat. Rev. Genet. 5, 435–445 (2004).
Gemayel, R., Vinces, M. D., Legendre, M. & Verstrepen, K. J. Variable tandem repeats accelerate evolution of coding and regulatory sequences. Annu. Rev. Genet. 44, 445–477 (2010).
Alasoo, K. et al. Shared genetic effects on chromatin and gene expression indicate a role for enhancer priming in immune response. Nat. Genet. 50, 424–431 (2018).
Lee, M. N. et al. Common genetic variants modulate pathogen-sensing responses in human dendritic cells. Science 343, 1246980 (2014).
Fairfax, B. P. et al. Innate immune activity conditions the effect of regulatory variants upon monocyte gene expression. Science 343, 1246949 (2014).
Zhang, Q. & Cao, X. Epigenetic regulation of the innate immune response to infection. Nat. Rev. Immunol. 19, 417–432 (2019).
Giambartolomei, C. et al. Bayesian test for colocalisation between pairs of genetic association studies using summary statistics. PLoS Genet. 10, e1004383 (2014).
Wallace, C. Eliciting priors and relaxing the single causal variant assumption in colocalisation analyses. PLoS Genet. 16, e1008720 (2020).
Finucane, H. K. et al. Partitioning heritability by functional annotation using genome-wide association summary statistics. Nat. Genet. 47, 1228–1235 (2015).
Gazal, S. et al. Linkage disequilibrium-dependent architecture of human complex traits shows action of negative selection. Nat. Genet. 49, 1421–1427 (2017).
Bulik-Sullivan, B. et al. LD score regression distinguishes confounding from polygenicity in genome-wide association studies. Nat. Genet. 47, 291–295 (2015).
Gamazon, E. R. et al. A gene-based association method for mapping traits using reference transcriptome data. Nat. Genet. 47, 1091–1098 (2015).
Ding, Z. et al. Quantitative genetics of CTCF binding reveal local sequence effects and different modes of X-chromosome association. PLoS Genet. 10, e1004798 (2014).
Tehranchi, A. K. et al. Pooled ChIP–seq links variation in transcription factor binding to complex disease risk. Cell 165, 730–741 (2016).
Fanucchi, S., Domínguez-Andrés, J., Joosten, L. A. B., Netea, M. G. & Mhlanga, M. M. The intersection of epigenetics and metabolism in trained immunity. Immunity 54, 32–43 (2021).
GTEx Consortium. Genetic effects on gene expression across human tissues. Nature 550, 204–213 (2017).
Võsa, U. et al. Large-scale cis- and trans-eQTL analyses identify thousands of genetic loci and polygenic scores that regulate blood gene expression. Nat. Genet. 53, 1300–1310 (2021).
GTEx Consortium. The GTEx Consortium atlas of genetic regulatory effects across human tissues. Science 369, 1318–1330 (2020).
Lovering, R. C., Camon, E. B., Blake, J. A. & Diehl, A. D. Access to immunology through the Gene Ontology. Immunology 125, 154–160 (2008).
Oliva, M. et al. The impact of sex on gene expression across human tissues. Science 369, eaba3066 (2020).
Pacis, A. et al. Gene activation precedes DNA demethylation in response to infection in human dendritic cells. Proc. Natl Acad. Sci. USA 116, 6938–6943 (2019).
Corces, M. R. et al. An improved ATAC–seq protocol reduces background and enables interrogation of frozen tissues. Nat. Methods 14, 959–962 (2017).
Schmidl, C., Rendeiro, A. F., Sheffield, N. C. & Bock, C. ChIPmentation: fast, robust, low-input ChIP–seq for histones and transcription factors. Nat. Methods 12, 963–965 (2015).
Bourgey, M. et al. GenPipes: an open-source framework for distributed and scalable genomic analyses. GigaScience 8, giz037 (2019).
Jiang, H., Lei, R., Ding, S. W. & Zhu, S. Skewer: a fast and accurate adapter trimmer for next-generation sequencing paired-end reads. BMC Bioinform. 15, 182 (2014).
Li, H. & Durbin, R. Fast and accurate short read alignment with Burrows-Wheeler transform. Bioinformatics 25, 1754–1760 (2009).
McKenna, A. et al. The genome analysis toolkit: a MapReduce framework for analyzing next-generation DNA sequencing data. Genome Res. 20, 1297–1303 (2010).
Picard. GitHub http://broadinstitute.github.io/picard/ (2014).
Chang, C. C. et al. Second-generation PLINK: rising to the challenge of larger and richer datasets. GigaScience 4, 7 (2015).
Purcell, S. et al. PLINK: a tool set for whole-genome association and population-based linkage analyses. Am. J. Hum. Genet. 81, 559–575 (2007).
Sherry, S. T. et al. dbSNP: the NCBI database of genetic variation. Nucleic Acids Res. 29, 308–311 (2001).
Delaneau, O., Coulonges, C. & Zagury, J. F. Shape-IT: new rapid and accurate algorithm for haplotype inference. BMC Bioinform. 9, 540 (2008).
Danecek, P. et al. Twelve years of SAMtools and BCFtools. GigaScience 10, giab008 (2021).
Bolger, A. M., Lohse, M. & Usadel, B. Trimmomatic: a flexible trimmer for Illumina sequence data. Bioinformatics 30, 2114–2120 (2014).
Dobin, A. et al. STAR: ultrafast universal RNA-seq aligner. Bioinformatics 29, 15–21 (2013).
Anders, S., Pyl, P. T. & Huber, W. HTSeq—a Python framework to work with high-throughput sequencing data. Bioinformatics 31, 166–169 (2015).
Zhang, Y. et al. Model-based analysis of ChIP–seq (MACS). Genome Biol. 9, R137 (2008).
Ernst, J. & Kellis, M. ChromHMM: automating chromatin-state discovery and characterization. Nat. Methods 9, 215–216 (2012).
Krueger, F. & Andrews, S. R. Bismark: a flexible aligner and methylation caller for Bisulfite-Seq applications. Bioinformatics 27, 1571–1572 (2011).
Pacis, A. et al. Bacterial infection remodels the DNA methylation landscape of human dendritic cells. Genome Res. 25, 1801–1811 (2015).
Robinson, M. D., McCarthy, D. J. & Smyth, G. K. edgeR: a Bioconductor package for differential expression analysis of digital gene expression data. Bioinformatics 26, 139–140 (2010).
Ritchie, M. E. et al. Limma powers differential expression analyses for RNA-sequencing and microarray studies. Nucleic Acids Res. 43, e47 (2015).
Park, Y. & Wu, H. Differential methylation analysis for BS-seq data under general experimental design. Bioinformatics 32, 1446–1453 (2016).
Grömping, U. Relative importance for linear regression in R: the package relaimpo. J. Stat. Softw. 17, 1–27 (2006).
Korotkevich, G. et al. Fast gene set enrichment analysis. Preprint at bioRxiv https://doi.org/10.1101/060012 (2021).
Yu, G., Wang, L. G. & He, Q. Y. ChIP seeker: an R/Bioconductor package for ChIP peak annotation, comparison and visualization. Bioinformatics 31, 2382–2383 (2015).
Li, Z. et al. Identification of transcription factor binding sites using ATAC-seq. Genome Biol. 20, 45 (2019).
Harrison, G. F. et al. Natural selection contributed to immunological differences between hunter-gatherers and agriculturalists. Nat. Ecol. Evol. 3, 1253–1264 (2019).
Hänzelmann, S., Castelo, R. & Guinney, J. GSVA: gene set variation analysis for microarray and RNA-seq data. BMC Bioinform. 14, 7 (2013).
Friedman, J., Hastie, T. & Tibshirani, R. Regularization paths for generalized linear models via coordinate descent. J. Stat. Softw. 33, 1–22 (2010).
Shabalin, A. A. Gene expression matrix eQTL: ultra fast eQTL analysis via large matrix operations. Bioinformatics 28, 1353–1358 (2012).
Kalita, C. A. et al. QuASAR-MPRA: accurate allele-specific analysis for massively parallel reporter assays. Bioinformatics 34, 787–794 (2018).
Harvey, C. T. et al. QuASAR: quantitative allele-specific analysis of reads. Bioinformatics 31, 1235–1242 (2015).
Willems, T. et al. Genome-wide profiling of heritable and de novo STR variations. Nat. Methods 14, 590–592 (2017).
Wang, G., Sarkar, A., Carbonetto, P. & Stephens, M. A simple new approach to variable selection in regression, with application to genetic fine mapping. J. R. Stat. Soc. Ser. B 82, 1273–1300 (2020).
Aracena, K. A. et al. Epigenetic variation impacts individual differences in the transcriptional response to influenza infection [data set]. Zenodo https://doi.org/10.5281/zenodo.10108240 (2023).
Aracena, K. A. katiearacena/EU_AF_ancestry_flu_code: Aracena et al code. Zenodo https://doi.org/10.5281/zenodo.10515250 (2024).
Pique-Regi, R. katiearacena/asepipeline: ASE code for Aracena et al. Zenodo https://doi.org/10.5281/zenodo.10511587 (2024).
Acknowledgements
We thank S. Vidal from McGill University for a gift of the influenza strain. We thank all members from the Barreiro and Bourque laboratories for their comments on the paper. This work was supported by National Institute of Health Research grants R01-GM134376 and P30-DK042086 to L.B.B. It is also supported by a Canada Institute of Health Research (CIHR) program grant (CEE-151618) for the McGill Epigenomics Mapping Center, which is part of the Canadian Epigenetics, Environment and Health Research Consortium (CEEHRC) Network, to G.B., L.B.B. and T.P. K.A.A. is supported by a grant to University of Chicago from the Howard Hughes Medical Institute through the James H. Gilliam Fellowships for Advanced Study program. G.B. is supported by a Canada Research Chair Tier 1 award, an FRQ-S, Distinguished Research Scholar award and by the World Premier International Research Center Initiative (WPI), NEXT, Japan. The Canadian Center for Computational Genomics (C3G) is supported by a Genome Canada Genome Technology Platform grant. Computational resources were provided by the University of Chicago Research Computing Center (Barreiro team) and Calcul Québec and Compute Canada (Bourque team). Figure 1a and Extended Data Fig. 8c were created with BioRender.com.
Author information
Authors and Affiliations
Contributions
L.B.B., G.B. and T.P. conceived the project. L.B.B. directed the study. V.Y., R.S., A. Pramatarova and M.-M.S. performed experimental work. K.A.A. led the computational analyses, with contributions from Y.-L.L, A. Pacis, S.G., Z.M., K.L., C.G., X.C., X.H., Y.L., C.B. and R.P.-R. A. Pacis and D.L. developed and implemented the EpiVar browser with help from R.G., D. Brownlee and D. Bujold. K.A.A. and L.B.B. wrote the manuscript, with input from all authors.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Genetics thanks Musa Mhlanga and the other, anonymous, reviewer(s) for their contribution to the peer review of this work. Peer reviewer reports are available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 Genome-wide impact of flu infection across regulatory marks.
(A) PVE by mock versus NI samples. (B) Comparison of infection effects for the union of genes tested in our study and Randolph et al.8. Pearson’s correlation coefficient and a 95% confidence interval are plotted). (C) Distribution depicting the relationship between gene expression changes and epigenetic changes in response to flu infection as seen in Fig. 1e but here focusing on epigenetic changes nearby genes that are downregulated in response to infection. Downregulated genes are defined as genes with beta < -0.5 and FDR < .01. Epigenetic changes are those with FDR < .01, except for methylation changes (FDR < .20).
Extended Data Fig. 2 Classification of ancestry-associated differences.
(A) Correlation of population differentially expressed (popDE) effects calculated with global or local ancestry effects (Pearson’s correlation coefficient reported). (B) Distribution depicting the relationship between popDE genes and popDE epigenetic changes across both conditions. Genes more highly expressed in individuals with high proportions of European ancestry (fold change < -0.5, FDR < 0.10) are nearby popDE epigenetic regions (FDR < .10) that show increased levels of chromatin accessibility, H3K27ac, H3K4me1 and H3K4me3 in individuals with increased European ancestry levels. Black lines represent means. (C) Distributions of individual mean scores of inflammatory pathways in the flu-infected condition comparable to Fig. 2c which shows non-infected condition distributions. A higher score indicates a strong expression of genes or epigenetic marks nearby genes within the Hallmark inflammatory response pathway. (D) Individual mean score differences between the population-groups for the Hallmark “inflammatory pathway” in the non-infected and (C) flu-infected conditions remain consistent when reducing popDE effects FDR from 10% to 5%. P values in panels C and D calculated using a two-sided Wilcoxon rank sum test. (E) Population-group differences utilizing popDR effects to calculate individual transcriptional response score across 6 immune pathways remain consistent with varying FDR thresholds (20%, 10% and 5%). (n = 35 individuals: 14 AF, 21 EU) P values calculated using a two-sided Wilcoxon rank sum test. The maxima and minima are the upper and lower points, respectively. The center line represents the median, and the top and bottom lines are the 75% and 25% percentile, respectively. (F) The distribution of Spearman’s correlation between the predicted and observed mean scores for the various pathways using different alphas.
Extended Data Fig. 3 Power calculations and validation of the QTL identified using external data sets.
(A) Power calculations for QTL with effect sizes ranging from 0.1-0.3. Power to detect QTL increases as the effect size of the variant increases. (B) Validation of significant FDR < .10 QTL in our dataset. Pearson’s correlation coefficient and a 95% confidence interval are shown. First row: Left- Comparison of non-infected eQTL with Randolph et al. Middle- Comparison of flu-infected eQTL with Randolph et al. Right- Comparison of non-infected eQTL with Nedelec et al. Second row: Left- Comparison of non-infected caQTL with Alasoo et al. Right- Comparison of non-infected meQTL with Husquin et al. (C) ASE hits are enriched for QTL. Mean and 95% confidence interval measured by logistic regression.
Extended Data Fig. 4 QTL mapping of the different molecular traits.
(A) Proportion and number of SNP-QTL at a significance threshold of FDR < .10 in each condition (B) Proportion and number of STR-QTL at a significance threshold of FDR < .10 in each condition. (C) Proportion and number of genes/features associated with at least one SNP or STR QTL in non-infected macrophages. Shared QTL were defined as those genes/features associated with a QTL at an FDR < .10 when performing the QTL mapping against SNPs and STRs separately. SNP- or STR-specific are those only identified as significant (FDR < 0.1) against either SNPs or STRs. (D) The mean percent variance explained by the top SNP and STR across all features in the non-infected condition. Both is the sum of the PVE of the top SNP and top STR (E) The enrichment of TF binding sites across non-infected specific SNP-QTL using a logistic regression. TF clusters are shown. (See Supplementary Table 4 for full results).
Extended Data Fig. 5 Overlap of QTL across molecular traits.
(A) Left: The number of overlaps for each QTL type for the permuted analysis in the non-infected condition. More than one overlap indicates the QTL is shared with at least one other datatype. Center: The number of overlaps for each QTL type in the flu-infected condition. Right: The number of overlaps for each QTL type for the permuted analysis in the flu-infected condition. (B) The percentage of QTL in one data type that are also QTL for another data type in the flu-condition. The starting QTL (rows) are the QTL that are tested for sharing while the overlapping QTL (columns) are the percentage of each starting QTL that are shared with that datatype. The color of each circle corresponds to the percentage of sharing. (C) QTL sharing patterns for those QTL overlapping 2≥ data types) in the non-infected condition. Y axis the proportion of overlapping QTL (that is, the denominator is the number of QTL that are shared in at least 2 or more data types). (D) QTL sharing patterns for those QTL overlapping 3≥ data types) in the non-infected condition highlighting that caQTL, K4me1 QTL and meQTL are the most commonly shared. The Y axis is the same as described in (C) above.
Extended Data Fig. 6 Genetically driven variation in epigenetic levels has no impact on the magnitude of transcriptional responses upon IAV infection.
(A) Genotypes for epigenetic QTL at baseline have no impact on the transcriptional response of nearby genes. The light blue marks the mean for each genotype and gray the median across all genotypes. The center line of the boxplot represents the median, and the top and bottom lines are the 75% and 25% percentile, respectively. The maxima and minima are the upper and lower points, respectively. As detailed in Fig. 5, we restricted to QTL nearby upregulated genes that are not eQTL (P < .30). (B) Association between genetically encoded baseline differences in chromatin accessibility and baseline differences in other epigenetic marks. Left- Meta caQTL plot (at baseline condition) across caQTLs for accessibility regions associated with up-regulated genes (n = 681 caQTLs associated with 506 genes). Individuals with genotypes associated with increased chromatin accessibility also show significantly increased levels of H3K4me1 and H3K27ac (P < 2.2×10-16), and to a lesser extent, a reduction in the repressive mark.
Extended Data Fig. 7 Calculating the contribution of cis-acting regulatory variants to ancestry-associated differences.
(A) Relationship between the observed and predicted betas for significant population differentially expressed (popDE) features (FDR < .10) for each of the data types in both conditions (adjusted R2 reported). (B) Boxplots of individual transcriptional response scores after regressing out the effects of the top SNP and STR in each condition for the 6 immune response pathways. (n = 35 individuals: 14 AF, 21 EU) P-values were calculated using a two-sided Wilcoxon rank sum test. The maxima and minima are the upper and lower points, respectively. The center line represents the median, and the top and bottom lines are the 75% and 25% percentile, respectively.
Extended Data Fig. 8 Epigenetic QTLs overlap with genetic variants associated with immune-related diseases.
(A) Summary of colocalization results for duplicated immune related diseases (11 diseases were investigated through 14 GWAS). Points represent the number of significant hits defined as PP3 + PP4 > 0.5 and PP4/(PP3 + PP4) > 0.8 in either condition. (B) Summary of PrediXcan results. Each point represents the total number of genes (Bonferroni corrected p = 0.05) associated with the disease trait in either condition. A gene is only counted once even if multiple peaks are associated with the gene. (C) Schematic depicting the proposed hypothesis that epigenetic QTL may act as a proxy for genetic variation that under particular environmental conditions has an impact on gene expression levels. Blue boxes represent gene exons and green peaks represent ATACseq peaks. A genetic variant at the QTL location impacts TF binding, such that differential binding of the TF is associated with variation in chromatin accessibility (that is, an caQTL). If the activity of this enhancer requires the recruitment of an additional TF (here labelled “environment-induced TF”) only induced in response to specific environmental/developmental conditions, the caQTL will not be associated with variation in gene expression levels. Yet, this caQTL will be a proxy for a genetic variant that on the “right environment” will ultimately be associated with an eQTL. Under this model, epigenetic QTLs that colocalize with GWAS variants (but not with eQTLs) can be thought of as a means to identify genetic variants that have an impact on gene expression in a yet unmeasured environment. Created with BioRender.com.
Extended Data Fig. 9 Heritability explained by molecular QTL.
(A) Heritability enrichment results for the 9 additional GWAS not shown in Fig. 6c. A 95% confidence interval is displayed. (B) Bar plots, mean values +/- SEM, representing the percent of heritability explained by each of the molecular QTL in all conditions. (C) Average heritability enrichment across independent GWAS traits (allergy and eczema, adult-onset asthma, MS, RA and IBD), comparing s-LDSC results using generic baseline, generic baseline and adjusting for s-LDSC’s histone marks, as well as generic baseline and adjusting for histone marks from the current study. s-LDSC analysis was conducted on finemapped molecular QTLs (using fine-mapping tool SuSiE), treating PIPs from fine-mapping results as continuous annotations. The average enrichments across all independent traits (error bars represent standard errors) are plotted with the p-value of enrichments from a random effects meta-analysis. (D) Height, BMI, and schizophrenia are shown as examples of negative controls. Mean values +/- SEM are reported. Little or no enrichment is seen, significantly less than reported immune traits.
Supplementary information
Supplementary Information
Supplementary methods and Figs. 1–4 with captions.
Supplementary Table 1
Description of the samples and libraries generated for this study.
Supplementary Table 2
List of differentially expressed, accessible and methylated features in response to flu infection.
Supplementary Table 3
GSEA results for infection effects and popDE effects with and without top QTL regressed.
Supplementary Table 4
TF activity scores and TF enrichment results in condition specific QTL.
Supplementary Table 5
List of popDE and responsive features.
Supplementary Table 6
List of cis-regulatory QTL identified in NI and flu-infected macrophages using both SNPs and STRs.
Supplementary Table 7
QTL integration results.
Supplementary Table 8
eRNA-QTL enrichments.
Supplementary Table 9
Colocalization results for immune-related GWAS.
Supplementary Table 10
LDSC-computed heritability results for immune-related GWAS.
Supplementary Table 11
PrediXcan results for immune-related GWAS.
Supplementary Table 12
ATAC–seq primers.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Aracena, K.A., Lin, YL., Luo, K. et al. Epigenetic variation impacts individual differences in the transcriptional response to influenza infection. Nat Genet 56, 408–419 (2024). https://doi.org/10.1038/s41588-024-01668-z
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41588-024-01668-z