Abstract
The calcium-sensing receptor (CaSR) is a family C G-protein-coupled receptor1 (GPCR) that has a central role in regulating systemic calcium homeostasis2,3. Here we use cryo-electron microscopy and functional assays to investigate the activation of human CaSR embedded in lipid nanodiscs and its coupling to functional Gi versus Gq proteins in the presence and absence of the calcimimetic drug cinacalcet. High-resolution structures show that both Gi and Gq drive additional conformational changes in the activated CaSR dimer to stabilize a more extensive asymmetric interface of the seven-transmembrane domain (7TM) that involves key protein–lipid interactions. Selective Gi and Gq coupling by the receptor is achieved through substantial rearrangements of intracellular loop 2 and the C terminus, which contribute differentially towards the binding of the two G-protein subtypes, resulting in distinct CaSR–G-protein interfaces. The structures also reveal that natural polyamines target multiple sites on CaSR to enhance receptor activation by zipping negatively charged regions between two protomers. Furthermore, we find that the amino acid l-tryptophan, a well-known ligand of CaSR extracellular domains, occupies the 7TM bundle of the G-protein-coupled protomer at the same location as cinacalcet and other allosteric modulators. Together, these results provide a framework for G-protein activation and selectivity by CaSR, as well as its allosteric modulation by endogenous and exogenous ligands.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
Data availability
The cryo-EM density maps and corresponding coordinates have been deposited in the Electron Microscopy Data Bank (EMDB) and the PDB, respectively, under the following accession codes: EMD-40914 and 8SZF (cinacalcet-bound active-state CaSR); EMD-40915 and 8SZG (cinacalcet-bound CaSR–Gq complex); EMD-40916 and 8SZH (cinacalcet-bound CaSR–Gi complex); and EMD-40917 and 8SZI (PAM-free CaSR–Gi complex). Source data are provided with this paper.
References
Kniazeff, J., Prézeau, L., Rondard, P., Pin, J.-P. & Goudet, C. Dimers and beyond: the functional puzzles of class C GPCRs. Pharmacol. Ther. 130, 9–25 (2011).
Hofer, A. M. & Brown, E. M. Extracellular calcium sensing and signalling. Nat. Rev. Mol. Cell Biol. 4, 530–538 (2003).
Hannan, F. M., Kallay, E., Chang, W., Brandi, M. L. & Thakker, R. V. The calcium-sensing receptor in physiology and in calcitropic and noncalcitropic diseases. Nat. Rev. Endocrinol. 15, 33–51 (2019).
Vahe, C. et al. Diseases associated with calcium-sensing receptor. Orphanet J. Rare Dis. 12, 19 (2017).
Massy, Z. A., Henaut, L., Larsson, T. E. & Vervloet, M. G. Calcium-sensing receptor activation in chronic kidney disease: effects beyond parathyroid hormone control. Semin. Nephrol. 34, 648–659 (2014).
Eckardt, K.-U. et al. Evolving importance of kidney disease: from subspecialty to global health burden. Lancet 382, 158–169 (2013).
Niswender, C. M. & Conn, P. J. Metabotropic glutamate receptors: physiology, pharmacology, and disease. Annu. Rev. Pharmacol. Toxicol. 50, 295–322 (2010).
Gregory, K. J. & Goudet, C. International union of basic and clinical pharmacology. CXI. Pharmacology, signaling, and physiology of metabotropic glutamate receptors. Pharmacol. Rev. 73, 521–569 (2021).
Leach, K. et al. International Union of Basic and Clinical Pharmacology. CVIII. Calcium-sensing receptor nomenclature, pharmacology, and function. Pharmacol. Rev. 72, 558–604 (2020).
Seven, A. B. et al. G-protein activation by a metabotropic glutamate receptor. Nature 595, 450–454 (2021).
Lin, S. et al. Structures of Gi-bound metabotropic glutamate receptors mGlu2 and mGlu4. Nature 594, 583–588 (2021).
Shen, C. et al. Structural basis of GABAB receptor–Gi protein coupling. Nature 594, 594–598 (2021).
Magno, A. L., Ward, B. K. & Ratajczak, T. The calcium-sensing receptor: a molecular perspective. Endocr. Rev. 32, 3–30 (2011).
Centeno, P. P. et al. Phosphate acts directly on the calcium-sensing receptor to stimulate parathyroid hormone secretion. Nat. Commun. 10, 4693 (2019).
Leach, K. et al. Towards a structural understanding of allosteric drugs at the human calcium-sensing receptor. Cell Res. 26, 574–592 (2016).
Ling, S. et al. Structural mechanism of cooperative activation of the human calcium-sensing receptor by Ca2+ ions and l-tryptophan. Cell Res. 31, 383–394 (2021).
Gao, Y. et al. Asymmetric activation of the calcium-sensing receptor homodimer. Nature 595, 455–459 (2021).
Park, J. et al. Symmetric activation and modulation of the human calcium-sensing receptor. Proc. Natl Acad. Sci. USA 118, e2115849118 (2021).
Olsen, R. H. et al. TRUPATH, an open-source biosensor platform for interrogating the GPCR transducerome. Nat. Chem. Biol. 16, 841–849 (2020).
Masuho, I. et al. Distinct profiles of functional discrimination among G proteins determine the actions of G protein–coupled receptors. Sci. Signal. 8, ra123 (2015).
Wall, M. A. et al. The structure of the G-protein heterotrimer Giα1β1γ2. Cell 83, 1047–1058 (1995).
Duan, J. et al. Cryo-EM structure of an activated VIP1 receptor-G protein complex revealed by a NanoBiT tethering strategy. Nat. Commun. 11, 4121 (2020).
Timmers, H., Karperien, M., Hamdy, N., De Boer, H. & Hermus, A. Normalization of serum calcium by cinacalcet in a patient with hypercalcaemia due to a de novo inactivating mutation of the calcium-sensing receptor. J. Intern. Med. 260, 177–182 (2006).
Hannan, F. M. et al. Identification of 70 calcium-sensing receptor mutations in hyper-and hypo-calcaemic patients: evidence for clustering of extracellular domain mutations at calcium-binding sites. Hum. Mol. Genet. 21, 2768–2778 (2012).
Nagase, T. et al. A family of autosomal dominant hypocalcemia with a positive correlation between serum calcium and magnesium: identification of a novel gain of function mutation (Ser820Phe) in the calcium-sensing receptor. J. Clin. Endocrinol. Metab. 87, 2681–2687 (2002).
Shiohara, M. et al. A novel gain-of-function mutation (F821L) in the transmembrane domain of calcium-sensing receptor is a cause of severe sporadic hypoparathyroidism. Eur. J. Pediatr. 163, 94–98 (2004).
Hu, J. et al. A region in the seven-transmembrane domain of the human Ca2+ receptor critical for response to Ca2+. J. Biol. Chem. 280, 5113–5120 (2005).
Quinn, S. J. et al. The Ca2+-sensing receptor: a target for polyamines. Am. J. Physiol. 273, C1315–C1323 (1997).
Michael, A. J. Polyamines in eukaryotes, bacteria, and archaea. J. Biol. Chem. 291, 14896–14903 (2016).
Schamber, M. R. & Vafabakhsh, R. Mechanism of sensitivity modulation in the calcium-sensing receptor via electrostatic tuning. Nat. Commun. 13, 2194 (2022).
Cole, D. E. et al. Calcium-sensing receptor mutations and denaturing high performance liquid chromatography. J. Mol. Endocrinol. 42, 331–339 (2009).
Tan, Y. et al. Autosomal dominant hypocalcemia: a novel activating mutation (E604K) in the cysteine-rich domain of the calcium-sensing receptor. J. Clin. Endocrinol. Metab. 88, 605–610 (2003).
Kwan, B. et al. A novel CASR mutation (p. Glu757Lys) causing autosomal dominant hypocalcaemia type 1. Endocrinol. Diabetes Metab. Case Rep. 2018, 18-0107 (2018).
Conigrave, A. D., Quinn, S. J. & Brown, E. M. l-amino acid sensing by the extracellular Ca2+-sensing receptor. Proc. Natl Acad. Sci. USA 97, 4814–4819 (2000).
Mangmool, S. & Kurose, H. Gi/o protein-dependent and-independent actions of pertussis toxin (PTX). Toxins 3, 884–899 (2011).
Vargas-Poussou, R. et al. Familial hypocalciuric hypercalcemia types 1 and 3 and primary hyperparathyroidism: similarities and differences. J. Clin. Endocrinol. Metab. 101, 2185–2195 (2016).
Koltin, D. et al. Mild infantile hypercalcemia: diagnostic tests and outcomes. J. Pediatr. 159, 215–221 (2011).
Ray, K., Fan, G.-F., Goldsmith, P. K. & Spiegel, A. M. The carboxyl terminus of the human calcium receptor: requirements for cell-surface expression and signal transduction. J. Biol. Chem. 272, 31355–31361 (1997).
Nesbit, M. A. et al. Mutations affecting G-protein subunit α11 in hypercalcemia and hypocalcemia. N. Engl. J. Med. 368, 2476–2486 (2013).
Bai, M. et al. Protein kinase C phosphorylation of threonine at position 888 in Ca2+o-sensing receptor (CaR) inhibits coupling to Ca2+ store release. J. Biol. Chem. 273, 21267–21275 (1998).
Davies, S. L., Ozawa, A., McCormick, W. D., Dvorak, M. M. & Ward, D. T. Protein kinase C-mediated phosphorylation of the calcium-sensing receptor is stimulated by receptor activation and attenuated by calyculin-sensitive phosphatase activity. J. Biol. Chem. 282, 15048–15056 (2007).
Jiang, Y. F. et al. Protein kinase C (PKC) phosphorylation of the Ca2+o-sensing receptor (CaR) modulates functional interaction of G proteins with the CaR cytoplasmic tail. J. Biol. Chem. 277, 50543–50549 (2002).
Lazarus, S. et al. A novel mutation of the primary protein kinase C phosphorylation site in the calcium-sensing receptor causes autosomal dominant hypocalcemia. Eur. J. Endocrinol. 164, 429–435 (2011).
Brown, E. M. et al. Cloning and characterization of an extracellular Ca2+-sensing receptor from bovine parathyroid. Nature 366, 575–580 (1993).
Robertson, M. J., Meyerowitz, J. G., Panova, O., Borrelli, K. & Skiniotis, G. Plasticity in ligand recognition at somatostatin receptors. Nat. Struct. Mol. Biol. 29, 210–217 (2022).
Maeda, S. et al. Development of an antibody fragment that stabilizes GPCR/G-protein complexes. Nat. Commun. 9, 3712 (2018).
Bayburt, T. H., Grinkova, Y. V. & Sligar, S. G. Self-assembly of discoidal phospholipid bilayer nanoparticles with membrane scaffold proteins. Nano Lett. 2, 853–856 (2002).
Peisley, A. & Skiniotis, G. 2D projection analysis of GPCR complexes by negative stain electron microscopy. Methods Mol. Biol. 1335, 29–38 (2015).
Mastronarde, D. N. Automated electron microscope tomography using robust prediction of specimen movements. J. Struct. Biol. 152, 36–51 (2005).
Punjani, A., Rubinstein, J. L., Fleet, D. J. & Brubaker, M. A. cryoSPARC: algorithms for rapid unsupervised cryo-EM structure determination. Nat. Methods 14, 290–296 (2017).
Zivanov, J. et al. New tools for automated high-resolution cryo-EM structure determination in RELION-3. eLife 7, e42166 (2018).
Pettersen, E. F. et al. UCSF Chimera—a visualization system for exploratory research and analysis. J. Comput. Chem. 25, 1605–1612 (2004).
Zhang, X. et al. Structures of the human cholecystokinin receptors bound to agonists and antagonists. Nat. Chem. Biol. 17, 1230–1237 (2021).
Emsley, P., Lohkamp, B., Scott, W. G. & Cowtan, K. Features and development of Coot. Acta Crystallogr. D 66, 486–501 (2010).
Liebschner, D. et al. Macromolecular structure determination using X-rays, neutrons and electrons: recent developments in Phenix. Acta Crystallogr. D 75, 861–877 (2019).
Chen, V. B. et al. MolProbity: all-atom structure validation for macromolecular crystallography. Acta Crystallogr. D 66, 12–21 (2010).
Pettersen, E. F. et al. UCSF ChimeraX: structure visualization for researchers, educators, and developers. Protein Sci. 30, 70–82 (2021).
Falk‐Petersen, C. B. et al. Development of a robust mammalian cell‐based assay for studying recombinant α4β1/3δ GABAA receptor subtypes. Basic Clin. Pharmacol. Toxicol. 121, 119–129 (2017).
Acknowledgements
We thank E. Montabana at the Stanford Cryo-EM Center (cEMc) for support with data collection. This work was supported by R01 DK132902 (G.S.), R01 NS122394 (G.S. and J.M.M.), R01 DK071801 (L.L.), S10 RR029531 (L.L.), the Lundbeck Foundation Experiment R346-2020-1829 (S.N.R. and J.M.M.) and the Independent Research Fund Denmark (Medical Sciences, 1030-00050A) (S.N.R. and J.M.M.).
Author information
Authors and Affiliations
Contributions
F.H. and Y.G. generated expression constructs for structural studies. F.H., C.-G.W. and Y.G. expressed and purified CaSR and G proteins. F.H. designed and generated the CaSR mutants. F.H. reconstituted and purified the inactive-state CaSR and the CINA-bound active-state CaSR and CaSR–Gq complexes in nanodiscs. C.-G.W. and F.H. reconstituted and purified the PAM-free CaSR–Gi complex in nanodiscs. Y.G. reconstituted and purified CaSR–Gi in detergents and the CINA-bound CaSR–Gi complex in nanodiscs. C.-G.W., F.H. and Y.G. prepared cryo-EM grids. M.Z., F.H., C.-G.W. and Y.G. collected cryo-EM data. F.H., Y.G. and C.-G.W. processed cryo-EM data. F.H. and Y.G. built and refined the structural models. C.-G.W. and Y.G. expressed and purified MSP1E3D1. S.N.R. optimized and performed in vitro cellular G-protein activation and signalling assays and assisted in figure preparation under the supervision of J.M.M. C.-G.W. and F.H. performed TRUPATH assays. J.M.M. conceived, optimized and performed in vitro cellular G-protein activation and signalling mutagenesis assays and interpreted results. M.M.P.-S. performed the GTPγS activity assay and analysed the data. T.-J.G. collected and analysed mass spectrometry data under the supervision of L.L. M.J.R. provided constructs for Gi expression and assisted in modelling. A.B.S. assisted in data collection and processing. F.H. and G.S. analysed data and wrote the manuscript with input from all authors. G.S. supervised the project.
Corresponding author
Ethics declarations
Competing interests
G.S. is a co-founder of and consultant for Deep Apple Therapeutics. The remaining authors declare no competing interests.
Peer review
Peer review information
Nature thanks Qiuyan Chen, Reza Vafabakhsh and the other, anonymous, reviewer(s) for their contribution to the peer review of this work. Peer reviewer reports are available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data figures and tables
Extended Data Fig. 1 G-protein signalling of CaSR and reconstitution of human CaSR–G-protein complexes.
a, Survey of Emax of Ca2+-dependent CaSRΔ894 activation of 11 different G-protein BRET-based TRUPATH biosensors using benchmark receptors NTSR1 and β2AR. The BRET signal monitors dissociation of the heterotrimeric Gαβγ complex induced by receptor-mediated guanine nucleotide exchange at the Gα subunit19. nd, no coupling detected. b, Concentration response curves of Gi3 and Gq activation pathways of CaSRΔ894 by Ca2+ using the BRET-based TRUPATH assay. c, Ca2+ concentration response curves of Gi3, Gq and Gs activation by CaSRΔ894 using the BRET-based G-protein-activation assay. d, G-protein-activation assays measuring the functional responses of 15 Gα subtypes by full-length CaSR (WT) and CaSRΔ894 after stimulation with 10 mM Ca2+, by MOR after stimulation with 10 mM morphine, by H1R after stimulation of 100 mM histamine and by β2AR after stimulation with 10 mM isoproterenol, respectively. The BRET signal monitors the binding of the released Gβγ from receptor-mediated Gαβγ dissociation to a membrane-associated reporter20. e, G-protein-activation assays monitoring functional responses of the CaSRΔ894 dimer fused to C1(WT-C1)/C2(WT-C2) tails to Ca2+ alone or in combination with cinacalcet. f–i, Size-exclusion chromatography profiles of purified CINA-bound CaSR/Gi in LMNG detergent (f), CINA-bound CaSR–Gi complex in nanodiscs (g), PAM-free CaSR–Gi complex in nanodiscs (h), CINA-bound CaSR–Gq complex in nanodiscs (i). Schematic diagram of the constructs used for CaSR and G protein is shown for each complex. The sample sizes and descriptions of error bars for all Extended Data Figures are described in the Methods under the section ‘Statistics and reproducibility’.
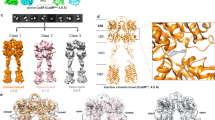
Extended Data Fig. 2 Cryo-EM processing and reconstruction of CaSR–Gi in LMNG detergent and the inactive-state CaSR in nanodiscs.
a, Cryo-EM data processing workflow for CaSR/Gi in detergents. b, Representative 2D averages. c, Local refinement maps of the class of Gi alone bound to detergent micelles. d, Low-resolution local refinement maps of the class of CaSR–Gi in detergents with a soft mask on receptor or G protein. e, Map of Gi alone with docked structure of inactive Gi (PDB ID: 1GP2). f, Low-resolution composite map of CaSR–Gi in LMNG detergent with docked structures of active-state CaSR (PDB ID: 7M3F) and inactive Gi (PDB ID: 1GP2). g, Model of the CINA-bound CaSR–Gi structure in nanodiscs docked into the density of receptor in the map of CaSR–Gi in LMNG detergent. h, Cryo-EM data processing workflow for inactive CaSR in nanodiscs. i, Representative 2D averages of inactive CaSR in nanodiscs. j, Local refinement maps of 7TM and VFT–CRD for inactive CaSR in nanodiscs. k, Low-resolution composite map of inactive CaSR in nanodiscs with open-closed VFTs and flexible 7TMs.
Extended Data Fig. 3 Cryo-EM processing and reconstruction of the CINA-bound and PAM-free CaSR–Gi complexes in lipid nanodiscs.
a, Cryo-EM data processing workflow for the CINA-bound CaSR–Gi complex in nanodiscs. b, Representative cryo-EM micrograph (out of 20528 similar micrographs) and 2D averages for the CINA-bound CaSR–Gi complex in nanodiscs. c, Local refinement maps of VFT–CRD, CRD–7TM and Gαiβγ for the CINA-bound CaSR–Gi complex in nanodiscs with corresponding Fourier shell correlation (FSC) curves indicating nominal resolutions using the FSC = 0.143 criterion. d, Angular distribution heat map of particle projections for the global reconstruction of the CINA-bound CaSR–Gi complex in nanodiscs. e, Cryo-EM data processing workflow for the PAM-free CaSR–Gi complex in nanodiscs. f, Representative cryo-EM micrograph (out of 31047 similar micrographs) and 2D averages for the PAM-free CaSR–Gi complex in nanodiscs. g, Local refinement maps of VFT–CRD, CRD–7TM and Gαiβγ for the PAM-free CaSR–Gi complex in nanodiscs with corresponding Fourier shell correlation (FSC) curves indicating nominal resolutions using the FSC = 0.143 criterion. h, Angular distribution heat map of particle projections for the global reconstruction of the PAM-free CaSR–Gi complex in nanodiscs. i, Local refinement maps of VFT–CRD, CRD–7TM for the PAM-free active-state CaSR in nanodiscs with corresponding FSC curves indicating nominal resolutions using the FSC = 0.143 criterion. j, Angular distribution heat map of particle projections for global reconstruction of the PAM-free active-state CaSR in nanodiscs.
Extended Data Fig. 4 Cryo-EM processing and reconstruction of the CINA-bound CaSR–Gq complex in lipid nanodiscs.
a, Cryo-EM data processing workflow for the CINA-bound CaSR–Gq complex in nanodiscs. b, Representative cryo-EM micrograph (out of 47434 similar micrographs) and 2D averages for the CINA-bound CaSR–Gq complex in nanodiscs. c, Local refinement maps of VFT–CRD, CRD–7TM and Gαqβγ for the CINA-bound CaSR–Gq complex in nanodiscs with corresponding FSC curves indicating nominal resolutions using the FSC = 0.143 criterion. d, Angular distribution heat map of particle projections for global reconstruction of the CINA-bound CaSR–Gq complex in nanodiscs. e, Local refinement maps of VFT–CRD, CRD–7TM for the CINA-bound active-state CaSR in nanodiscs with corresponding FSC curves indicating nominal resolutions using the FSC = 0.143 criterion. f, Angular distribution heat map of particle projections for reconstruction of the CINA-bound active-state CaSR in nanodiscs. g, Comparison of the maps of the CINA-bound CaSR–Gq and CaSR–Gi complexes in nanodiscs. The angles of the Gα αN helix relative to the lipid membrane are highlighted by dashed lines.
Extended Data Fig. 5 Agreement between cryo-EM density and model.
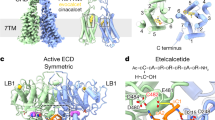
a, EM density and model for the 7TM with the extended CINA (at map thresholds of 0.3–0.35), and cinacalcet (at a map threshold of 0.3) in the CINA-bound active-state CaSR. b, EM density and model for the 7TM and H8 (residues 869 to 877) of the Gi-bound protomer (at map thresholds of 1.1–1.7), cinacalcet (at a map threshold of 1.35), and lipids (using the unsharpened map at thresholds of 0.4–0.6) in the CINA-bound CaSR–Gi complex. c, EM density and model for the 7TM and the extended H8 (residues 869 to 885) of the Gq-bound protomer (at map thresholds of 0.9–1.0), cinacalcet (at a map threshold of 0.85), and lipids (at a map threshold of 0.82) in the CINA-bound CaSR–Gq complex. Residues 878 to 885 in the C terminus are highlighted in turquoise as in Fig. 4b. d, EM density and model for the 7TM and H8 (residues 869 to 877) of the Gi-bound protomer (at map thresholds of 0.41–0.46), the 7TM Trp (at a map threshold of 0.4), and lipids (at map thresholds of 0.31–0.4) in the PAM-free CaSR–Gi complex.
Extended Data Fig. 6 Asymmetric dimer interface of CaSR in the active state and in G-protein-coupled states.
a, Structural comparison between active-state CaSR in detergent (PDB ID: 7M3F) and lipid nanodiscs. The ECDs in the two structures show different tilting angles relative to the 7TMs. b, Structural comparison between the CINA-bound CaSR–Gi and CaSR–Gq complexes in nanodiscs. The 7TMs (except the cytoplasmic regions) and the bound ligands are well aligned. c, The VFT of the NGC protomer of the CINA-bound CaSR–Gi or CaSR–Gq complex is aligned to the VFT of the GC protomer. d, The asymmetric TM6–TM6 interface in the structure of CINA-bound active-state CaSR in nanodiscs. e,f, The asymmetric 7TM interface in the structure of the CINA-bound CaSR–Gi in nanodiscs shown in two views, highlighting the interface between TM6 of 7TMNGC and TM6 of 7TMGC (e), and the interface between TM5 and TM6 of 7TMGC and TM7 of 7TMNGC (f). The residues that are involved in dimer interactions, and the models of phospholipids (DOPC) and cholesterol are shown. g, Functional responses of CaSR WT and mutants to Ca2+, for Gi3 and Gq activation, categorized based on interactions with the lower (F776A, K805A, F806A and F809A) and upper (I822A, Y825A, Y829A and V833A) phospholipids.
Extended Data Fig. 7 Polyamine-binding sites on CaSR.
a, Mass spectrometry for spermidine and spermine standards, and sample of purified CINA-bound CaSR–Gi complex in lipid nanodiscs. No significant amount of spermidine was detected. b, The spermine-binding site in the CRD–ECL regions of the CINA-bound CaSR–Gq complex in nanodiscs. The model and cryo-EM density of the bound spermine (at a map threshold of 1.13), a water molecule (in red sphere), the upper phospholipid (using the unsharpened map at a map threshold of 0.51) are shown. c, Electrostatic surface distribution of the modelled surface of CaSR in the CINA-bound CaSR–Gi complex in nanodiscs. The three spermine-binding sites are highlighted with dashed boxes. d, IP1-accumulation assays monitoring the functional responses of full-length CaSR to Ca2+, spermine or spermine in presence of NAM NPS 2143. Pre-incubated NPS 2143 inhibited the spermine response, whereas increasing concentrations of the PAM etelcalcetide potentiated the spermine response. e, G-protein-activation and IP1-accumulation assays monitoring the functional responses of full-length CaSR to Ca2+, spermine or spermine with 3 mM Ca2+, in the presence/absence of 20 mM (G-protein-activation assay) or 10 mM (IP1 accumulation) EDTA. Functional responses by Gi3 and Gq activation were measured 20 seconds after ligand stimulation to capture the initial activation by Gi3 and Gq, as well as to circumvent any potential long-term cellular effects of EDTA, albeit responses were similar in the IP1-accumulation assay. Pharmacological analysis across G-protein-activation and IP1-accumulation assays indicate that spermine (dark blue) may function like an ago-PAM, which is potentiated by Ca2+ as evident from its left-shifted potency with 3 mM Ca2+ (light green), and right-shifted potency in the presence of EDTA (light blue, dark green). Similar data were obtained for CaSR Δ894 WT (data not shown).
Extended Data Fig. 8 Conformational changes of CaSR associated with Gi and Gq coupling, and distinct Gi and Gq binding interfaces.
a,b, Superposition of the structures of the CINA-bound active-state CaSR and CaSR–Gi complex (a) or CaSR–Gq complex (b) based on VFT alignment. Conformational differences between the superimposed structures are indicated by arrows. c, Comparison of the two 7TMs in the CINA-bound active-state CaSR structure. d, Comparison between 7TMNGC in the CINA-bound CaSR–Gi complex and the 7TM in the active-state CaSR. e,f, Comparison of 7TMGC and 7TMNGC in the CINA-bound CaSR–Gi (e) and CaSR–Gq (f) complexes. g,h, Interactions between ICL2 of CaSR and the N-terminal regions of Gi (g) and Gq (h). i,j, Superposition of the structures of the CINA-bound CaSR–Gi and CaSR–Gq complexes by aligning the two 7TMs in top (i) and side (j) views. Conformational differences of Gi and Gq are indicated by arrows. k,l, The CaSR C terminus selectively recognizes Gi and Gq. The structures of the CINA-bound CaSR–Gi and CaSR–Gq complexes are overlaid based on the alignment of 7TMGC showing only the Gq-bound protomer (k) or the Gi-bound protomer (l). Potential clashes are represented by yellow stars. m, Sequence alignment of the α5 helix in the Gαq/11 class. The strictly conserved residues are highlighted in red.
Extended Data Fig. 9 Functional characterization and relative expression levels of wild-type CaSR and CaSR mutants.
a, IP1-accumulation assays monitoring functional responses of CaSR WT (with or without pretreatment with PTX) and CaSR mutants to Ca2+, categorized based on different structural domains/regions. b, Selected mutants identical or complementary to the mutants tested in the G-protein-activation assays in Fig. 4. c, [35S]-GTPγS binding activity assays monitoring functional responses of an extracellular domain truncated CaSR mutant ΔECD CaSR (CaSR residues 599–894) to Trp and cinacalcet in the presence and absence of 10 mM Ca2+. d, IP1-accumulation assay monitoring the functional response of ΔECD CaSR to cinacalcet and Trp. e,f, Surface expression levels of CaSR WT and mutants for Gi3 and Gq activation assays (e), and surface expression levels of CaSR WT and mutants for IP1-accumulation assays (f).
Extended Data Fig. 10 The 7TM modulatory sites in the presence and absence of cinacalcet.
a, The PAM-binding pocket of 7TMGC in the CINA-bound CaSR–Gi complex. b, The PAM-binding pocket of 7TMNGC in the CINA-bound CaSR–Gi complex. c, Overall structure of the PAM-free CaSR–Gi complex. The 7TM Trp, phospholipids (DOPC), and cholesterol are shown as spheres. d, The PAM-binding pocket of 7TMGC in the PAM-free CaSR–Gi complex. e, The PAM-binding pocket of 7TMNGC in the PAM-free CaSR–Gi complex. f, Trp and cinacalcet occupy a similar position in 7TMGC in the CINA-bound and in the PAM-free CaSR–Gi complexes. The binding-pocket residues share almost identical conformations in the two structures. g, The model and density of the two W818 rotamers in 7TMGC in the PAM-free CaSR–Gi complex. h, Comparison of 7TMNGC in the PAM-free and CINA-bound CaSR–Gi complexes. i, The two 7TMs in the CINA-bound active-state CaSR show different tilt angles in the lipid bilayer, and the protomer with the extended CINA has a better resolved C terminus. j,k, Modelling based on the structure of the CINA-bound CaSR–Gi complex shows how the C781W/I822W mutations would not efficiently occlude the bent PAM conformation (j), and how the L773W mutation would occlude the bent PAM conformation (k). A potential clash is represented by the yellow star. l, Superposition of the 7TMs in the CINA-bound CaSR–Gi complex with those in the active-state CaSR (CaSR*) based on the poses (bent or extended) of cinacalcet.
Supplementary information
Supplementary Information
Supplementary Figs 1 and 2.
Supplementary Video 1
Conformational dynamics of 7TMGC relative to 7TMNGC in the structure of the CINA-bound CaSR–Gi complex. Morph between the CINA-bound 7TMNGC and the CINA-bound 7TMGC. The models are aligned as shown in Fig. 5c.
Supplementary Video 2
Conformational dynamics of 7TMGC relative to 7TMNGC in the structure of the PAM-free CaSR–Gi complex. Morph between the PAM-free 7TMNGC and the Trp-bound 7TMGC. The models are aligned as shown in Fig. 5b.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
He, F., Wu, CG., Gao, Y. et al. Allosteric modulation and G-protein selectivity of the Ca2+-sensing receptor. Nature 626, 1141–1148 (2024). https://doi.org/10.1038/s41586-024-07055-2
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41586-024-07055-2
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.