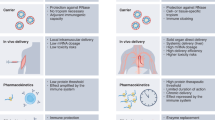
Abstract
mRNA formulated with lipid nanoparticles is a transformative technology that has enabled the rapid development and administration of billions of coronavirus disease 2019 (COVID-19) vaccine doses worldwide. However, avoiding unacceptable toxicity with mRNA drugs and vaccines presents challenges. Lipid nanoparticle structural components, production methods, route of administration and proteins produced from complexed mRNAs all present toxicity concerns. Here, we discuss these concerns, specifically how cell tropism and tissue distribution of mRNA and lipid nanoparticles can lead to toxicity, and their possible reactogenicity. We focus on adverse events from mRNA applications for protein replacement and gene editing therapies as well as vaccines, tracing common biochemical and cellular pathways. The potential and limitations of existing models and tools used to screen for on-target efficacy and de-risk off-target toxicity, including in vivo and next-generation in vitro models, are also discussed.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Barouch, D. H. Covid-19 vaccines - immunity, variants, boosters. N. Engl. J. Med. 387, 1011–1020 (2022).
El Sahly, H. M. et al. Efficacy of the mRNA-1273 SARS-CoV-2 vaccine at completion of blinded phase. N. Engl. J. Med. 385, 1774–1785 (2021).
Thomas, S. J. et al. Efficacy and safety of the BNT162b2 mRNA COVID-19 vaccine in participants with a history of cancer: subgroup analysis of a global phase 3 randomized clinical trial. Vaccine 40, 1483–1492 (2022).
Rouf, N. Z., Biswas, S., Tarannum, N., Oishee, L. M. & Muna, M. M. Demystifying mRNA vaccines: an emerging platform at the forefront of cryptic diseases. RNA Biol. 19, 386–410 (2022).
Chalkias, S. et al. A bivalent omicron-containing booster vaccine against Covid-19. N. Engl. J. Med. 387, 1279–1291 (2022).
Verma, M. et al. The landscape for lipid-nanoparticle-based genomic medicines. Nat. Rev. Drug. Discov. 22, 349–350 (2023).
CONGRESS.GOV. S.5002 - FDA Modernization Act 2.0. https://congress.gov/bill117th-congress/senate-bill/5002 (2022).
Ingber, D. E. Human organs-on-chips for disease modelling, drug development and personalized medicine. Nat. Rev. Genet. 23, 467–491 (2022).
Granot, Y. & Peer, D. Delivering the right message: challenges and opportunities in lipid nanoparticles-mediated modified mRNA therapeutics-an innate immune system standpoint. Semin. Immunol. 34, 68–77 (2017).
Duan, Q. et al. How far are the new wave of mRNA drugs from us? mRNA product current perspective and future development. Front. Immunol. 13, 974433 (2022).
Igyarto, B. Z., Jacobsen, S. & Ndeupen, S. Future considerations for the mRNA-lipid nanoparticle vaccine platform. Curr. Opin. Virol. 48, 65–72 (2021).
Moghimi, S. M. & Simberg, D. Pro-inflammatory concerns with lipid nanoparticles. Mol. Ther. 30, 2109–2110 (2022).
Kenjo, E. et al. Low immunogenicity of LNP allows repeated administrations of CRISPR–Cas9 mRNA into skeletal muscle in mice. Nat. Commun. 12, 7101 (2021).
Guerrini, G. et al. Monitoring anti-PEG antibodies level upon repeated lipid nanoparticle-based COVID-19 vaccine administration. Int. J. Mol. Sci. 23, 8838 (2022).
Verbeke, R., Hogan, M. J., Lore, K. & Pardi, N. Innate immune mechanisms of mRNA vaccines. Immunity 55, 1993–2005 (2022). This article describes the current understanding of how both mRNA and its LNP carrier contribute to the immune response elicited by mRNA vaccines, with the LNP acting as a potent adjuvant.
Tenchov, R., Bird, R., Curtze, A. E. & Zhou, Q. Lipid nanoparticles ─ from liposomes to mRNA vaccine delivery, a landscape of research diversity and advancement. ACS Nano 15, 16982–17015 (2021).
Li, D. et al. Messenger RNA-based therapeutics and vaccines: what’s beyond COVID-19? ACS Pharmacol. Transl. Sci. 6, 943–969 (2023).
Huang, X. et al. The landscape of mRNA nanomedicine. Nat. Med. 28, 2273–2287 (2022).
Alexopoulou, L., Holt, A. C., Medzhitov, R. & Flavell, R. A. Recognition of double-stranded RNA and activation of NF-κB by Toll-like receptor 3. Nature 413, 732–738 (2001).
Kariko, K., Ni, H., Capodici, J., Lamphier, M. & Weissman, D. mRNA is an endogenous ligand for Toll-like receptor 3. J. Biol. Chem. 279, 12542–12550 (2004).
Panda, D. et al. IRF1 maintains optimal constitutive expression of antiviral genes and regulates the early antiviral response. Front. Immunol. 10, 1019 (2019).
Kobayashi, K. et al. IRAK-M is a negative regulator of Toll-like receptor signaling. Cell 110, 191–202 (2002).
Cheng, K., Wang, X. & Yin, H. Small-molecule inhibitors of the TLR3/dsRNA complex. J. Am. Chem. Soc. 133, 3764–3767 (2011).
Bernard, J. J. et al. Ultraviolet radiation damages self noncoding RNA and is detected by TLR3. Nat. Med. 18, 1286–1290 (2012).
Hornung, V., Barchet, W., Schlee, M. & Hartmann, G. in Toll-like receptors (TLRs) and Innate Immunity (eds Bauer, S. & Hartmann, G.) 71–86. Handbook of Experimental Pharmacology series https://doi.org/10.1007/978-3-540-72167-3_4 (Springer, 2008).
Weissman, D. et al. HIV gag mRNA transfection of dendritic cells (DC) delivers encoded antigen to MHC class I and II molecules, causes DC maturation, and induces a potent human in vitro primary immune response. J. Immunol. 165, 4710–4717 (2000).
Koski, G. K. et al. Cutting edge: innate immune system discriminates between RNA containing bacterial versus eukaryotic structural features that prime for high-level IL-12 secretion by dendritic cells. J. Immunol. 172, 3989–3993 (2004).
Diebold, S. S., Kaisho, T., Hemmi, H., Akira, S. & Reis e Sousa, C. Innate antiviral responses by means of TLR7-mediated recognition of single-stranded RNA. Science 303, 1529–1531 (2004).
Heil, F. et al. Species-specific recognition of single-stranded RNA via toll-like receptor 7 and 8. Science 303, 1526–1529 (2004).
Zhang, Z. et al. Structural analysis reveals that Toll-like receptor 7 is a dual receptor for guanosine and single-stranded RNA. Immunity 45, 737–748 (2016).
Tanji, H. et al. Toll-like receptor 8 senses degradation products of single-stranded RNA. Nat. Struct. Mol. Biol. 22, 109–115 (2015).
Guo, S. et al. Size, shape, and sequence-dependent immunogenicity of RNA nanoparticles. Mol. Ther. Nucleic Acids 9, 399–408 (2017).
Mu, X., Greenwald, E., Ahmad, S. & Hur, S. An origin of the immunogenicity of in vitro transcribed RNA. Nucleic Acids Res. 46, 5239–5249 (2018).
Bui, T. M., Wiesolek, H. L. & Sumagin, R. ICAM-1: a master regulator of cellular responses in inflammation, injury resolution, and tumorigenesis. J. Leukoc. Biol. 108, 787–799 (2020).
Mu, X. & Hur, S. Immunogenicity of in vitro-transcribed RNA. Acc. Chem. Res. 54, 4012–4023 (2021).
Schlake, T., Thess, A., Fotin-Mleczek, M. & Kallen, K. J. Developing mRNA-vaccine technologies. RNA Biol. 9, 1319–1330 (2012).
Verbeke, R., Lentacker, I., De Smedt, S. C. & Dewitte, H. Three decades of messenger RNA vaccine development. Nano Today 28, 100766 (2019).
Vierbuchen, T., Stein, K. & Heine, H. RNA is taking its toll: impact of RNA-specific Toll-like receptors on health and disease. Allergy 74, 223–235 (2019).
Kariko, K., Buckstein, M., Ni, H. & Weissman, D. Suppression of RNA recognition by Toll-like receptors: the impact of nucleoside modification and the evolutionary origin of RNA. Immunity 23, 165–175 (2005). In this seminal article, it is shown that incorporation of modified nucleosides or pseudouridine in IVT mRNA ablates their immunogenicity against dendritic cells and other TLR-presenting cells.
Kormann, M. S. et al. Expression of therapeutic proteins after delivery of chemically modified mRNA in mice. Nat. Biotechnol. 29, 154–157 (2011).
Kariko, K., Muramatsu, H., Ludwig, J. & Weissman, D. Generating the optimal mRNA for therapy: HPLC purification eliminates immune activation and improves translation of nucleoside-modified, protein-encoding mRNA. Nucleic Acids Res. 39, e142 (2011).
Kariko, K., Muramatsu, H., Keller, J. M. & Weissman, D. Increased erythropoiesis in mice injected with submicrogram quantities of pseudouridine-containing mRNA encoding erythropoietin. Mol. Ther. 20, 948–953 (2012).
Weissman, D. mRNA transcript therapy. Expert Rev. Vaccines 14, 265–281 (2015).
Holtkamp, S. et al. Modification of antigen-encoding RNA increases stability, translational efficacy, and T-cell stimulatory capacity of dendritic cells. Blood 108, 4009–4017 (2006).
Schuberth-Wagner, C. et al. A conserved histidine in the RNA sensor RIG-I controls immune tolerance to N1-2′ O-methylated self RNA. Immunity 43, 41–51 (2015).
Abbas, Y. M. et al. Structure of human IFIT1 with capped RNA reveals adaptable mRNA binding and mechanisms for sensing N1 and N2 ribose 2′-O methylations. Proc. Natl Acad. Sci. USA 114, E2106–E2115 (2017).
Andries, O. et al. N1-methylpseudouridine-incorporated mRNA outperforms pseudouridine-incorporated mRNA by providing enhanced protein expression and reduced immunogenicity in mammalian cell lines and mice. J. Control. Release 217, 337–344 (2015).
Baiersdörfer, M. et al. A facile method for the removal of dsRNA contaminant from in vitro-transcribed mRNA. Mol. Ther. Nucleic Acids 15, 26–35 (2019).
Dousis, A., Ravichandran, K., Hobert, E. M., Moore, M. J. & Rabideau, A. E. An engineered T7 RNA polymerase that produces mRNA free of immunostimulatory byproducts. Nat. Biotechnol. 41, 560–568 (2022).
Nelson, J. et al. Impact of mRNA chemistry and manufacturing process on innate immune activation. Sci. Adv. 6, eaaz6893 (2020).
Qin, S. et al. mRNA-based therapeutics: powerful and versatile tools to combat diseases. Signal Transduct. Target. Ther. 7, 166 (2022). This review provides an in-depth description of the principles of action of mRNA-based drugs at the molecular level.
Pekker, M. & Shneider, M. The surface charge of a cell lipid membrane. Preprint at https://doi.org/10.48550/arXiv.1401.4707 (2014).
Houseley, J. & Tollervey, D. The many pathways of RNA degradation. Cell 136, 763–776 (2009).
Kiaie, S. H. et al. Recent advances in mRNA-LNP therapeutics: immunological and pharmacological aspects. J. Nanobiotechnol. 20, 276 (2022).
Ickenstein, L. M. & Garidel, P. Lipid-based nanoparticle formulations for small molecules and RNA drugs. Expert Opin. Drug Deliv. 16, 1205–1226 (2019).
Nakamura, T. et al. Extrahepatic targeting of lipid nanoparticles in vivo with intracellular targeting for future nanomedicines. Adv. Drug Deliv. Rev. 188, 114417 (2022).
Zhang, Y., Sun, C., Wang, C., Jankovic, K. E. & Dong, Y. Lipids and lipid derivatives for RNA delivery. Chem. Rev. 121, 12181–12277 (2021).
Han, X. et al. An ionizable lipid toolbox for RNA delivery. Nat. Commun. 12, 7233 (2021).
Suk, J. S., Xu, Q., Kim, N., Hanes, J. & Ensign, L. M. PEGylation as a strategy for improving nanoparticle-based drug and gene delivery. Adv. Drug Deliv. Rev. 99, 28–51 (2016).
Paunovska, K. et al. Nanoparticles containing oxidized cholesterol deliver mRNA to the liver microenvironment at clinically relevant doses. Adv. Mater. 31, e1807748 (2019).
Cheng, X. & Lee, R. J. The role of helper lipids in lipid nanoparticles (LNPs) designed for oligonucleotide delivery. Adv. Drug Deliv. Rev. 99, 129–137 (2016).
Zhang, R. et al. Helper lipid structure influences protein adsorption and delivery of lipid nanoparticles to spleen and liver. Biomater. Sci. 9, 1449–1463 (2021).
Weng, J., Yang, M., Wang, W., Xu, X. & Tian, Z. Revealing thermodynamics and kinetics of lipid self-assembly by markov state model analysis. J. Am. Chem. Soc. 142, 21344–21352 (2020).
Cornebise, M. et al. Discovery of a novel amino lipid that improves lipid nanoparticle performance through specific interactions with mRNA. Adv. Funct. Mater. 32, 2106727 (2021).
Dolgin, E. Startups set off new wave of mRNA therapeutics. Nat. Biotechnol. 39, 1029–1031 (2021).
Rohner, E., Yang, R., Foo, K. S., Goedel, A. & Chien, K. R. Unlocking the promise of mRNA therapeutics. Nat. Biotechnol. 40, 1586–1600 (2022).
Jeong, M., Lee, Y., Park, J., Jung, H. & Lee, H. Lipid nanoparticles (LNPs) for in vivo RNA delivery and their breakthrough technology for future applications. Adv. Drug Deliv. Rev. 200, 114990 (2023). This review summarizes recent advances of RNA therapeutics and provides a comprehensive description of their pharmacokinetic and pharmacodynamic challenges.
Barbier, A. J., Jiang, A. Y., Zhang, P., Wooster, R. & Anderson, D. G. The clinical progress of mRNA vaccines and immunotherapies. Nat. Biotechnol. 40, 840–854 (2022).
Pardi, N. et al. Expression kinetics of nucleoside-modified mRNA delivered in lipid nanoparticles to mice by various routes. J. Control. Release 217, 345–351 (2015).
Dilliard, S. A., Cheng, Q. & Siegwart, D. J. On the mechanism of tissue-specific mRNA delivery by selective organ targeting nanoparticles. Proc. Natl Acad. Sci. USA 118, e2019256118 (2021).
John, S. et al. Multi-antigenic human cytomegalovirus mRNA vaccines that elicit potent humoral and cell-mediated immunity. Vaccine 36, 1689–1699 (2018).
Deering, R. P., Kommareddy, S., Ulmer, J. B., Brito, L. A. & Geall, A. J. Nucleic acid vaccines: prospects for non-viral delivery of mRNA vaccines. Expert Opin. Drug Deliv. 11, 885–899 (2014).
Liang, F. et al. Efficient targeting and activation of antigen-presenting cells in vivo after modified mRNA vaccine administration in rhesus macaques. Mol. Ther. 25, 2635–2647 (2017).
Ols, S. et al. Route of vaccine administration alters antigen trafficking but not innate or adaptive immunity. Cell Rep. 30, 3964–3971.e3967 (2020).
Giannotta, G. & Giannotta, N. mRNA COVID-19 vaccines and long-lived plasma cells: a complicated relationship. Vaccines 9, 1503 (2021).
Li, C. et al. Mechanisms of innate and adaptive immunity to the Pfizer-BioNTech BNT162b2 vaccine. Nat. Immunol. 23, 543–555 (2022).
Manolova, V. et al. Nanoparticles target distinct dendritic cell populations according to their size. Eur. J. Immunol. 38, 1404–1413 (2008).
Pardi, N. et al. Nucleoside-modified mRNA vaccines induce potent T follicular helper and germinal center B cell responses. J. Exp. Med. 215, 1571–1588 (2018).
Roltgen, K. et al. Immune imprinting, breadth of variant recognition, and germinal center response in human SARS-CoV-2 infection and vaccination. Cell 185, 1025–1040.e1014 (2022).
Zhang, Z. et al. Humoral and cellular immune memory to four COVID-19 vaccines. Cell 185, 2434–2451.e2417 (2022).
Alameh, M. G. et al. Lipid nanoparticles enhance the efficacy of mRNA and protein subunit vaccines by inducing robust T follicular helper cell and humoral responses. Immunity 54, 2877–2892 e2877 (2021). This article shows that the LNP component of the mRNA-based vaccines against SARS-CoV-2 has inherent adjuvanticity, leading to long-lived plasma cell and memory B cell responses.
Lorentzen, C. L., Haanen, J. B., Met, O. & Svane, I. M. Clinical advances and ongoing trials on mRNA vaccines for cancer treatment. Lancet Oncol. 23, e450–e458 (2022).
Jung, H. N., Lee, S. Y., Lee, S., Youn, H. & Im, H. J. Lipid nanoparticles for delivery of RNA therapeutics: current status and the role of in vivo imaging. Theranostics 12, 7509–7531 (2022).
Zehrung, D., Jarrahian, C. & Wales, A. Intradermal delivery for vaccine dose sparing: overview of current issues. Vaccine 31, 3392–3395 (2013).
Shi, J. et al. Delivery of mRNA for regulating functions of immune cells. J. Control. Release 345, 494–511 (2022).
Huysmans, H. et al. Expression kinetics and innate immune response after electroporation and LNP-mediated delivery of a self-amplifying mRNA in the skin. Mol. Ther. Nucleic Acids 17, 867–878 (2019).
Szoke, D. et al. Nucleoside-modified VEGFC mRNA induces organ-specific lymphatic growth and reverses experimental lymphedema. Nat. Commun. 12, 3460 (2021).
Anderluzzi, G. et al. The role of nanoparticle format and route of administration on self-amplifying mRNA vaccine potency. J. Control. Release 342, 388–399 (2022).
Oberli, M. A. et al. Lipid nanoparticle assisted mRNA delivery for potent cancer immunotherapy. Nano Lett. 17, 1326–1335 (2017).
Zadory, M., Lopez, E., Babity, S., Gravel, S. P. & Brambilla, D. Current knowledge on the tissue distribution of mRNA nanocarriers for therapeutic protein expression. Biomater. Sci. 10, 6077–6115 (2022).
Ngo, W. et al. Identifying cell receptors for the nanoparticle protein corona using genome screens. Nat. Chem. Biol. 18, 1023–1031 (2022).
Sato, Y., Kinami, Y., Hashiba, K. & Harashima, H. Different kinetics for the hepatic uptake of lipid nanoparticles between the apolipoprotein E/low density lipoprotein receptor and the N-acetyl-d-galactosamine/asialoglycoprotein receptor pathway. J. Control. Release 322, 217–226 (2020). This article provides novel insight into the mechanism and kinetics of LNP uptake by hepatocytes.
Chen, D., Parayath, N., Ganesh, S., Wang, W. & Amiji, M. The role of apolipoprotein- and vitronectin-enriched protein corona on lipid nanoparticles for in vivo targeted delivery and transfection of oligonucleotides in murine tumor models. Nanoscale 11, 18806–18824 (2019).
Akinc, A. et al. Targeted delivery of RNAi therapeutics with endogenous and exogenous ligand-based mechanisms. Mol. Ther. 18, 1357–1364 (2010).
Zhang, A. et al. Absorption, distribution, metabolism, and excretion of nanocarriers in vivo and their influences. Adv. Colloid Interface Sci. 284, 102261 (2020).
Eipel, C., Abshagen, K. & Vollmar, B. Regulation of hepatic blood flow: the hepatic arterial buffer response revisited. World J. Gastroenterol. 16, 6046–6057 (2010).
Tsoi, K. M. et al. Mechanism of hard-nanomaterial clearance by the liver. Nat. Mater. 15, 1212–1221 (2016).
Li, J., Chen, C. & Xia, T. Understanding nanomaterial–liver interactions to facilitate the development of safer nanoapplications. Adv. Mater. 34, e2016456 (2022).
Ramaswamy, S. et al. Systemic delivery of factor IX messenger RNA for protein replacement therapy. Proc. Natl Acad. Sci. USA 114, E1941–E1950 (2017).
Sedic, M. et al. Safety evaluation of lipid nanoparticle-formulated modified mRNA in the Sprague-Dawley rat and cynomolgus monkey. Vet. Pathol. 55, 341–354 (2018).
Cao, J. et al. mRNA therapy restores euglycemia and prevents liver tumors in murine model of glycogen storage disease. Nat. Commun. 12, 3090 (2021).
Jiang, L. et al. Dual mRNA therapy restores metabolic function in long-term studies in mice with propionic acidemia. Nat. Commun. 11, 5339 (2020).
Zhu, X. et al. Systemic mRNA therapy for the treatment of Fabry disease: preclinical studies in wild-type mice, Fabry mouse model, and wild-type non-human primates. Am. J. Hum. Genet. 104, 625–637 (2019).
Ishida, T. & Kiwada, H. Accelerated blood clearance (ABC) phenomenon upon repeated injection of PEGylated liposomes. Int. J. Pharm. 354, 56–62 (2008).
Yang, Q. & Lai, S. K. Anti-PEG immunity: emergence, characteristics, and unaddressed questions. Wiley Interdiscip. Rev. Nanomed. Nanobiotechnol. 7, 655–677 (2015).
El Sayed, M. M. et al. Hepatosplenic phagocytic cells indirectly contribute to anti-PEG IgM production in the accelerated blood clearance (ABC) phenomenon against PEGylated liposomes: appearance of an unexplained mechanism in the ABC phenomenon. J. Control. Release 323, 102–109 (2020).
Shi, D. et al. To PEGylate or not to PEGylate: immunological properties of nanomedicine’s most popular component, polyethylene glycol and its alternatives. Adv. Drug Deliv. Rev. 180, 114079 (2022).
Estape Senti, M. et al. Anti-PEG antibodies compromise the integrity of PEGylated lipid-based nanoparticles via complement. J. Control. Release 341, 475–486 (2022). Ths study shows that anti-PEG antibodies can compromise the integrity of LNP-mRNA formulations, causing early mRNA release, and trigger the release of complement activation products.
Suzuki, T. et al. Accelerated blood clearance of PEGylated liposomes containing doxorubicin upon repeated administration to dogs. Int. J. Pharm. 436, 636–643 (2012).
Abramson, A. et al. Oral mRNA delivery using capsule-mediated gastrointestinal tissue injections. Matter 5, 975–987 (2022).
O’Driscoll, C. M., Bernkop-Schnurch, A., Friedl, J. D., Preat, V. & Jannin, V. Oral delivery of non-viral nucleic acid-based therapeutics - do we have the guts for this? Eur. J. Pharm. Sci. 133, 190–204 (2019).
Tenchov, R., Sasso, J. M. & Zhou, Q. A. PEGylated lipid nanoparticle formulations: immunological safety and efficiency perspective. Bioconjug. Chem. 34, 941–960 (2023).
An, D. et al. Long-term efficacy and safety of mRNA therapy in two murine models of methylmalonic acidemia. EBioMedicine 45, 519–528 (2019).
Truong, B. et al. Lipid nanoparticle-targeted mRNA therapy as a treatment for the inherited metabolic liver disorder arginase deficiency. Proc. Natl Acad. Sci. USA 116, 21150–21159 (2019).
Broudic, K. et al. Nonclinical safety evaluation of a novel ionizable lipid for mRNA delivery. Toxicol. Appl. Pharmacol. 451, 116143 (2022).
Szebeni, J. et al. Applying lessons learned from nanomedicines to understand rare hypersensitivity reactions to mRNA-based SARS-CoV-2 vaccines. Nat. Nanotechnol. 17, 337–346 (2022). This review summarizes known reactogenicity concerns from past nanomedicine infusions and draws parallels to understand potential risks with SARS-CoV-2 LNP-mRNA vaccines.
Lacy, P. & Stow, J. L. Cytokine release from innate immune cells: association with diverse membrane trafficking pathways. Blood 118, 9–18 (2011).
Elsabahy, M. & Wooley, K. L. Cytokines as biomarkers of nanoparticle immunotoxicity. Chem. Soc. Rev. 42, 5552–5576 (2013).
Parhiz, H. et al. Added to pre-existing inflammation, mRNA-lipid nanoparticles induce inflammation exacerbation (IE). J. Control. Release 344, 50–61 (2022).
Hammel, M. et al. Correlating the structure and gene silencing activity of oligonucleotide-loaded lipid nanoparticles using small-angle X-ray scattering. ACS Nano 17, 11454–11465 (2023).
Cárdenas, M., Campbell, R. A., Arteta, M. Y., Lawrence, M. J. & Sebastiani, F. Review of structural design guiding the development of lipid nanoparticles for nucleic acid delivery. Curr. Opin. Colloid Interface Sci. 66, 101705 (2023).
Lokugamage, M. P. et al. Mild innate immune activation overrides efficient nanoparticle-mediated RNA delivery. Adv. Mater. 32, e1904905 (2020).
Ndeupen, S. et al. The mRNA-LNP platform’s lipid nanoparticle component used in preclinical vaccine studies is highly inflammatory. iScience 24, 103479 (2021).
Kauffman, K. J. et al. Efficacy and immunogenicity of unmodified and pseudouridine-modified mRNA delivered systemically with lipid nanoparticles in vivo. Biomaterials 109, 78–87 (2016).
Unterberger, S., Davies, K. A., Rambhatla, S. B. & Sacre, S. Contribution of Toll-like receptors and the NLRP3 inflammasome in rheumatoid arthritis pathophysiology. Immunotargets Ther. 10, 285–298 (2021).
Yu, P. et al. Pyroptosis: mechanisms and diseases. Signal Transduct. Target. Ther. 6, 128 (2021).
Forster Iii, J., Nandi, D. & Kulkarni, A. mRNA-carrying lipid nanoparticles that induce lysosomal rupture activate NLRP3 inflammasome and reduce mRNA transfection efficiency. Biomater. Sci. 10, 5566–5582 (2022). In this study it is shown that the ionizable cationic lipids, and cholesterol impact the endosomal rupture capabilities of LNP-mRNA formulations and lead to NLRP3 inflammasome activation.
Chevriaux, A. et al. Cathepsin B is required for NLRP3 inflammasome activation in macrophages, through NLRP3 interaction. Front. Cell Dev. Biol. 8, 167 (2020).
Tahtinen, S. et al. IL-1 and IL-1ra are key regulators of the inflammatory response to RNA vaccines. Nat. Immunol. 23, 532–542 (2022). This study shows that the ionizable lipid in LNP-mRNA vaccines triggers the IL-1 pathway, TLR signalling and cytokine release more in human than in murine leukocytes.
Liu, T., Zhang, L., Joo, D. & Sun, S. C. NF-kappaB signaling in inflammation. Signal Transduct. Target. Ther. 2, 17023 (2017).
Dunkelberger, J. R. & Song, W. C. Complement and its role in innate and adaptive immune responses. Cell Res. 20, 34–50 (2010).
Song, W. C. Crosstalk between complement and toll-like receptors. Toxicol. Pathol. 40, 174–182 (2012).
Holers, V. M. Phenotypes of complement knockouts. Immunopharmacology 49, 125–131 (2000).
Xu, Y. et al. Complement activation in factor D-deficient mice. Proc. Natl Acad. Sci. USA 98, 14577–14582 (2001).
Batista, A. F. et al. Complement C3 lowering in adult inducible conditional knockout mice: long‐lasting effects. Alzheimer’s Dement. 18, e068094 (2022).
Shiraishi, K. et al. Exploring the relationship between anti-PEG IgM behaviors and PEGylated nanoparticles and its significance for accelerated blood clearance. J. Control. Release 234, 59–67 (2016).
Suzuki, T. et al. PEG shedding-rate-dependent blood clearance of PEGylated lipid nanoparticles in mice: faster PEG shedding attenuates anti-PEG IgM production. Int. J. Pharm. 588, 119792 (2020).
Vu, V. P. et al. Immunoglobulin deposition on biomolecule corona determines complement opsonization efficiency of preclinical and clinical nanoparticles. Nat. Nanotechnol. 14, 260–268 (2019).
Shimabukuro, T. & Nair, N. Allergic reactions including anaphylaxis after receipt of the first dose of Pfizer–BioNTech COVID-19 vaccine. JAMA 325, 780–781 (2021).
Lim, X. R. et al. Anaphylatoxin complement 5a in Pfizer BNT162b2-induced immediate-type vaccine hypersensitivity reactions. Vaccines 11, 1020 (2023).
Padin-Gonzalez, E. et al. Understanding the role and impact of poly (ethylene glycol) (PEG) on nanoparticle formulation: implications for COVID-19 vaccines. Front. Bioeng. Biotechnol. 10, 882363 (2022).
Kozma, G. T. et al. Pseudo-anaphylaxis to polyethylene glycol (PEG)-coated liposomes: roles of anti-PEG IgM and complement activation in a porcine model of human infusion reactions. ACS Nano 13, 9315–9324 (2019).
Chonn, A., Cullis, P. R. & Devine, D. V. The role of surface charge in the activation of the classical and alternative pathways of complement by liposomes. J. Immunol. 146, 4234–4241 (1991).
Holley, C. K. & Dobrovolskaia, M. A. Innate immunity modulating impurities and the immunotoxicity of nanobiotechnology-based drug products. Molecules 26, 7308 (2021).
Descotes, J. & Choquet-Kastylevsky, G. Gell and Coombs’s classification: is it still valid? Toxicology 158, 43–49 (2001).
Dobrovolskaia, M. A. Lessons learned from immunological characterization of nanomaterials at the Nanotechnology Characterization Laboratory. Front. Immunol. 13, 984252 (2022).
Zolnik, B. S. & Sadrieh, N. Regulatory perspective on the importance of ADME assessment of nanoscale material containing drugs. Adv. Drug Deliv. Rev. 61, 422–427 (2009).
Patel, S. et al. Boosting intracellular delivery of lipid nanoparticle-encapsulated mRNA. Nano Lett. 17, 5711–5718 (2017).
Magtanong, L. et al. Exogenous monounsaturated fatty acids promote a ferroptosis-resistant cell state. Cell Chem. Biol. 26, 420–432 e429 (2019).
Das, U. N. Saturated fatty acids, MUFAs and PUFAs regulate ferroptosis. Cell Chem. Biol. 26, 309–311 (2019).
Young, R. S. E. et al. Apocryphal FADS2 activity promotes fatty acid diversification in cancer. Cell Rep. 34, 108738 (2021).
Spickett, C. M. The lipid peroxidation product 4-hydroxy-2-nonenal: advances in chemistry and analysis. Redox Biol. 1, 145–152 (2013).
Ioannou, M. S. et al. Neuron-astrocyte metabolic coupling protects against activity-induced fatty acid toxicity. Cell 177, 1522–1535 e1514 (2019).
Bosma, M. et al. Sequestration of fatty acids in triglycerides prevents endoplasmic reticulum stress in an in vitro model of cardiomyocyte lipotoxicity. Biochim. Biophys. Acta 1841, 1648–1655 (2014).
Ding, J. et al. The peroxisomal enzyme L-PBE is required to prevent the dietary toxicity of medium-chain fatty acids. Cell Rep. 5, 248–258 (2013).
Sato, Y. et al. Highly specific delivery of siRNA to hepatocytes circumvents endothelial cell-mediated lipid nanoparticle-associated toxicity leading to the safe and efficacious decrease in the hepatitis B virus. J. Control. Release 266, 216–225 (2017).
Kauffman, K. J. et al. Optimization of lipid nanoparticle formulations for mRNA delivery in vivo with fractional factorial and definitive screening designs. Nano Lett. 15, 7300–7306 (2015).
Dilliard, S. A. & Siegwart, D. J. Passive, active and endogenous organ-targeted lipid and polymer nanoparticles for delivery of genetic drugs. Nat. Rev. Mater. 8, 282–300 (2023). This review presents the passive, active and endogenous targeting mechanisms that mediate the delivery of LNP-formulated nucleic acids to cells and tissues.
Verhoef, J. J. & Anchordoquy, T. J. Questioning the use of PEGylation for drug delivery. Drug Deliv. Transl. Res. 3, 499–503 (2013).
Bao, Y. et al. Effect of PEGylation on biodistribution and gene silencing of siRNA/lipid nanoparticle complexes. Pharm. Res. 30, 342–351 (2013).
Harvie, P., Wong, F. M. & Bally, M. B. Use of poly(ethylene glycol)–lipid conjugates to regulate the surface attributes and transfection activity of lipid–DNA particles. J. Pharm. Sci. 89, 652–663 (2000).
Besin, G. et al. Accelerated blood clearance of lipid nanoparticles entails a biphasic humoral response of B-1 followed by B-2 lymphocytes to distinct antigenic moieties. Immunohorizons 3, 282–293 (2019). This article proposes a mechanism for the increased blood clearance of PEGylated LNP-mRNA formulations and the connection between their innate and adaptive immune responses.
Wardemann, H., Boehm, T., Dear, N. & Carsetti, R. B-1a B cells that link the innate and adaptive immune responses are lacking in the absence of the spleen. J. Exp. Med. 195, 771–780 (2002).
Mohamed, M. et al. PEGylated liposomes: immunological responses. Sci. Technol. Adv. Mater. 20, 710–724 (2019).
Wang, X. et al. Preparation of selective organ-targeting (SORT) lipid nanoparticles (LNPs) using multiple technical methods for tissue-specific mRNA delivery. Nat. Protoc. 18, 265–291 (2022).
Chander, N., Basha, G., Cheng, M. H. Y., Witzigmann, D. & Cullis, P. R. Lipid nanoparticle mRNA systems containing high levels of sphingomyelin engender enhanced protein expression in hepatic and extra-hepatic tissues. Mol. Ther. Methods Clin. Dev. 30, 235–245 (2023).
Patel, S. et al. Naturally-occurring cholesterol analogues in lipid nanoparticles induce polymorphic shape and enhance intracellular delivery of mRNA. Nat. Commun. 11, 983 (2020).
Kedmi, R., Ben-Arie, N. & Peer, D. The systemic toxicity of positively charged lipid nanoparticles and the role of Toll-like receptor 4 in immune activation. Biomaterials 31, 6867–6875 (2010).
Audouy, S. A., de Leij, L. F., Hoekstra, D. & Molema, G. In vivo characteristics of cationic liposomes as delivery vectors for gene therapy. Pharm. Res. 19, 1599–1605 (2002).
Xue, H. Y., Liu, S. & Wong, H. L. Nanotoxicity: a key obstacle to clinical translation of siRNA-based nanomedicine. Nanomedicine 9, 295–312 (2014).
Maldonado-Pereira, L., Schweiss, M., Barnaba, C. & Medina-Meza, I. G. The role of cholesterol oxidation products in food toxicity. Food Chem. Toxicol. 118, 908–939 (2018).
Song, Y., Liu, J., Zhao, K., Gao, L. & Zhao, J. Cholesterol-induced toxicity: an integrated view of the role of cholesterol in multiple diseases. Cell Metab. 33, 1911–1925 (2021).
Austin, L. A. et al. Split-dose administration enhances immune responses elicited by a mRNA/lipid nanoparticle vaccine expressing respiratory syncytial virus F protein. Mol. Pharm. 20, 279–289 (2022).
Qin, M., Du, G. & Sun, X. Recent advances in the noninvasive delivery of mRNA. Acc. Chem. Res. 54, 4262–4271 (2021).
Koh, K. J. et al. Formulation, characterization and evaluation of mRNA-loaded dissolvable polymeric microneedles (RNApatch). Sci. Rep. 8, 11842 (2018).
Golombek, S. et al. Intradermal delivery of synthetic mRNA using hollow microneedles for efficient and rapid production of exogenous proteins in skin. Mol. Ther. Nucleic Acids 11, 382–392 (2018).
Barz, M. et al. RNA particles comprising polysarcosine. WO2020069718 (2020).
Nogueira, S. S. et al. Polysarcosine-functionalized lipid nanoparticles for therapeutic mRNA delivery. ACS Appl. Nano Mater. 3, 10634–10645 (2020).
Evers, M. J. W. et al. Delivery of modified mRNA to damaged myocardium by systemic administration of lipid nanoparticles. J. Control. Release 343, 207–216 (2022).
Magadum, A. et al. Pkm2 regulates cardiomyocyte cell cycle and promotes cardiac regeneration. Circulation 141, 1249–1265 (2020).
Algarni, A. et al. In vivo delivery of plasmid DNA by lipid nanoparticles: the influence of ionizable cationic lipids on organ-selective gene expression. Biomater. Sci. 10, 2940–2952 (2022).
Hashiba, K. et al. Branching ionizable lipids can enhance the stability, fusogenicity, and functional delivery of mRNA. Small Sci. 3, 2200071 (2022).
Akinc, A. et al. A combinatorial library of lipid-like materials for delivery of RNAi therapeutics. Nat. Biotechnol. 26, 561–569 (2008).
Zaslavsky, J., Bannigan, P. & Allen, C. Re-envisioning the design of nanomedicines: harnessing automation and artificial intelligence. Expert Opin. Drug Deliv. 50, 241–257 (2023).
Azarnezhad, A., Samadian, H., Jaymand, M., Sobhani, M. & Ahmadi, A. Toxicological profile of lipid-based nanostructures: are they considered as completely safe nanocarriers? Crit. Rev. Toxicol. 50, 148–176 (2020).
Swingle, K. L. et al. Amniotic fluid stabilized lipid nanoparticles for in utero intra-amniotic mRNA delivery. J. Control. Release 341, 616–633 (2022).
Miao, L., Zhang, Y. & Huang, L. mRNA vaccine for cancer immunotherapy. Mol. Cancer 20, 41 (2021).
Liu, J. Q. et al. Intratumoral delivery of IL-12 and IL-27 mRNA using lipid nanoparticles for cancer immunotherapy. J. Control. Release 345, 306–313 (2022).
Liu, L. et al. Combination immunotherapy of MUC1 mRNA nano-vaccine and CTLA-4 blockade effectively inhibits growth of triple negative breast cancer. Mol. Ther. 26, 45–55 (2018).
Xiao, Y. et al. Combining p53 mRNA nanotherapy with immune checkpoint blockade reprograms the immune microenvironment for effective cancer therapy. Nat. Commun. 13, 758 (2022).
Gillmore, J. D. et al. CRISPR-Cas9 in vivo gene editing for transthyretin amyloidosis. N. Engl. J. Med. 385, 493–502 (2021).
Tao, W. et al. Mechanistically probing lipid-siRNA nanoparticle-associated toxicities identifies Jak inhibitors effective in mitigating multifaceted toxic responses. Mol. Ther. 19, 567–575 (2011).
Terada, T., Kulkarni, J. A., Huynh, A., Tam, Y. Y. C. & Cullis, P. Protective effect of edaravone against cationic lipid-mediated oxidative stress and apoptosis. Biol. Pharm. Bull. 44, 144–149 (2021).
O’Brien, J., Hayder, H., Zayed, Y. & Peng, C. Overview of MicroRNA biogenesis, mechanisms of actions, and circulation. Front. Endocrinol. 9, 402 (2018).
Elhanati, S. et al. Reciprocal regulation between SIRT6 and miR-122 controls liver metabolism and predicts hepatocarcinoma prognosis. Cell Rep. 14, 234–242 (2016).
Davis, A. M., Scott, T. A. & Morris, K. V. Harnessing rift valley fever virus NSs gene for cancer gene therapy. Cancer Gene Ther. 29, 1477–1486 (2022).
Jain, R. et al. MicroRNAs enable mRNA therapeutics to selectively program cancer cells to self-destruct. Nucleic Acid. Ther. 28, 285–296 (2018). This study demostrates how inclusion of miRNAs in mRNA therapeutics can allow for the selective expression of therapeutic payloads in diseased cells in vivo.
Loughrey, D. & Dahlman, J. E. Non-liver mRNA delivery. Acc. Chem. Res. 55, 13–23 (2022).
Cheng, Q. et al. Selective organ targeting (SORT) nanoparticles for tissue-specific mRNA delivery and CRISPR-Cas gene editing. Nat. Nanotechnol. 15, 313–320 (2020). In this study, it was shown that the inclusion of permanently charged lipids in an LNP formulation can faciliate the targeted in vivo biodistribution and expression of mRNA payloads.
Lopez, C. et al. Loading of lutein in egg-sphingomyelin vesicles as lipid carriers: thermotropic phase behaviour, structure of sphingosome membranes and lutein crystals. Food Res. Int. 138, 109770 (2020).
Sato, Y., Hatakeyama, H., Hyodo, M. & Harashima, H. Relationship between the physicochemical properties of lipid nanoparticles and the quality of siRNA delivery to liver cells. Mol. Ther. 24, 788–795 (2016).
Muller, P. Y. & Milton, M. N. The determination and interpretation of the therapeutic index in drug development. Nat. Rev. Drug. Discov. 11, 751–761 (2012).
Zuberi, A. & Lutz, C. Mouse models for drug discovery. can new tools and technology improve translational power? ILAR J. 57, 178–185 (2016).
Li, M., Al-Jamal, K. T., Kostarelos, K. & Reineke, J. Physiologically based pharmacokinetic modeling of nanoparticles. ACS Nano 4, 6303–6317 (2010).
Moghimi, S. M. & Simberg, D. Translational gaps in animal models of human infusion reactions to nanomedicines. Nanomedicine 13, 973–975 (2018).
Hatit, M. Z. C. et al. Species-dependent in vivo mRNA delivery and cellular responses to nanoparticles. Nat. Nanotechnol. 17, 310–318 (2022).
Blais, E. M. et al. Reconciled rat and human metabolic networks for comparative toxicogenomics and biomarker predictions. Nat. Commun. 8, 14250 (2017).
Whitebread, S., Hamon, J., Bojanic, D. & Urban, L. Keynote review: in vitro safety pharmacology profiling: an essential tool for successful drug development. Drug Discov. Today 10, 1421–1433 (2005).
Niles, A. L., Moravec, R. A. & Riss, T. L. Update on in vitro cytotoxicity assays for drug development. Expert Opin. Drug Discov. 3, 655–669 (2008).
Stamatiadis, P. et al. Comparative analysis of mouse and human preimplantation development following POU5F1 CRISPR/Cas9 targeting reveals interspecies differences. Hum. Reprod. 36, 1242–1252 (2021).
Moreb, E. A. & Lynch, M. D. Genome dependent Cas9/gRNA search time underlies sequence dependent gRNA activity. Nat. Commun. 12, 5034 (2021).
Rogers, M. A. & Aikawa, E. Cardiovascular calcification: artificial intelligence and big data accelerate mechanistic discovery. Nat. Rev. Cardiol. 16, 261–274 (2019).
Malik, N. & Rao, M. S. A review of the methods for human iPSC derivation. Methods Mol. Biol. 997, 23–33 (2013).
Ware, B. R., Berger, D. R. & Khetani, S. R. Prediction of drug-induced liver injury in micropatterned co-cultures containing iPSC-derived human hepatocytes. Toxicol. Sci. 145, 252–262 (2015).
Kopljar, I. et al. Development of a human iPSC cardiomyocyte-based scoring system for cardiac hazard identification in early drug safety de-risking. Stem Cell Rep. 11, 1365–1377 (2018).
Macedo, M. H., Araujo, F., Martinez, E., Barrias, C. & Sarmento, B. iPSC-derived enterocyte-like cells for drug absorption and metabolism studies. Trends Mol. Med. 24, 696–708 (2018).
Gutbier, S. et al. Large-scale production of human iPSC-derived macrophages for drug screening. Int. J. Mol. Sci. 21, 4808 (2020).
Sharma, A., Sances, S., Workman, M. J. & Svendsen, C. N. Multi-lineage human iPSC-derived platforms for disease modeling and drug discovery. Cell Stem Cell 26, 309–329 (2020).
Liu, C., Oikonomopoulos, A., Sayed, N. & Wu, J. C. Modeling human diseases with induced pluripotent stem cells: from 2D to 3D and beyond. Development 145, dev156166 (2018).
Broutier, L. et al. Human primary liver cancer-derived organoid cultures for disease modeling and drug screening. Nat. Med. 23, 1424–1435 (2017).
Berical, A. et al. A multimodal iPSC platform for cystic fibrosis drug testing. Nat. Commun. 13, 4270 (2022).
Czerniecki, S. M. et al. High-throughput screening enhances kidney organoid differentiation from human pluripotent stem cells and enables automated multidimensional phenotyping. Cell Stem Cell 22, 929–940 e924 (2018).
Rowe, R. G. & Daley, G. Q. Induced pluripotent stem cells in disease modelling and drug discovery. Nat. Rev. Genet. 20, 377–388 (2019).
Yanez Arteta, M. et al. Successful reprogramming of cellular protein production through mRNA delivered by functionalized lipid nanoparticles. Proc. Natl Acad. Sci. USA 115, E3351–E3360 (2018).
Astolfi, M. et al. Micro-dissected tumor tissues on chip: an ex vivo method for drug testing and personalized therapy. Lab. Chip 16, 312–325 (2016).
Workman, M. J. et al. Engineered human pluripotent-stem-cell-derived intestinal tissues with a functional enteric nervous system. Nat. Med. 23, 49–59 (2017).
Ronaldson-Bouchard, K. & Vunjak-Novakovic, G. Organs-on-a-chip: a fast track for engineered human tissues in drug development. Cell Stem Cell 22, 310–324 (2018).
Theobald, J. et al. Liver-kidney-on-chip to study toxicity of drug metabolites. ACS Biomater. Sci. Eng. 4, 78–89 (2018).
Conant, G. et al. High-content assessment of cardiac function using heart-on-a-chip devices as drug screening model. Stem Cell Rev. Rep. 13, 335–346 (2017).
Jalili-Firoozinezhad, S. et al. Modeling radiation injury-induced cell death and countermeasure drug responses in a human gut-on-a-chip. Cell Death Dis. 9, 223 (2018).
Yin, F. et al. A 3D human placenta-on-a-chip model to probe nanoparticle exposure at the placental barrier. Toxicol. Vitr. 54, 105–113 (2019).
Cohen, A. et al. Mechanism and reversal of drug-induced nephrotoxicity on a chip. Sci. Transl. Med. 13, eabd6299 (2021).
Si, L. et al. A human-airway-on-a-chip for the rapid identification of candidate antiviral therapeutics and prophylactics. Nat. Biomed. Eng. 5, 815–829 (2021).
Goyal, G. et al. Ectopic lymphoid follicle formation and human seasonal influenza vaccination responses recapitulated in an organ-on-a-chip. Adv. Sci. 9, e2103241 (2022).
Skardal, A., Shupe, T. & Atala, A. Organoid-on-a-chip and body-on-a-chip systems for drug screening and disease modeling. Drug Discov. Today 21, 1399–1411 (2016).
Ozkan, A., Ghousifam, N., Hoopes, P. J., Yankeelov, T. E. & Rylander, M. N. In vitro vascularized liver and tumor tissue microenvironments on a chip for dynamic determination of nanoparticle transport and toxicity. Biotechnol. Bioeng. 116, 1201–1219 (2019).
Nashimoto, Y. et al. Vascularized cancer on a chip: the effect of perfusion on growth and drug delivery of tumor spheroid. Biomaterials 229, 119547 (2020).
Ewart, L. et al. Performance assessment and economic analysis of a human liver-chip for predictive toxicology. Commun. Med. 2, 154 (2022).
Jang, K. J. et al. Reproducing human and cross-species drug toxicities using a liver-chip. Sci. Transl. Med. 11, eaax5516 (2019). This article demonstrates the capacity of liver-on-chip platforms to identify drug-induced liver injuries from small molecules.
Lu, R. X. Z. & Radisic, M. Organ-on-a-chip platforms for evaluation of environmental nanoparticle toxicity. Bioact. Mater. 6, 2801–2819 (2021).
Shanti, A., Teo, J. & Stefanini, C. In vitro immune organs-on-chip for drug development: a review. Pharmaceutics 10, 278 (2018).
Baysoy, A., Bai, Z., Satija, R. & Fan, R. The technological landscape and applications of single-cell multi-omics. Nat. Rev. Mol. Cell Biol. 24, 695–713 (2023).
Dobrowolski, C. et al. Nanoparticle single-cell multiomic readouts reveal that cell heterogeneity influences lipid nanoparticle-mediated messenger RNA delivery. Nat. Nanotechnol. 17, 871–879 (2022). This study reveals that cell subsets have distinct responses to LNPs that may affect mRNA therapies.
Kackos, C. M. et al. mRNA vaccine mitigates SARS-CoV-2 infections and COVID-19. Microbiol. Spectr. 11, e04240–e04222 (2023).
Kulkarni, J. A. et al. The current landscape of nucleic acid therapeutics. Nat. Nanotechnol. 16, 630–643 (2021).
Chaudhary, N., Weissman, D. & Whitehead, K. A. mRNA vaccines for infectious diseases: principles, delivery and clinical translation. Nat. Rev. Drug Discov. 20, 817–838 (2021).
Munchel, S. E., Shultzaberger, R. K., Takizawa, N. & Weis, K. Dynamic profiling of mRNA turnover reveals gene-specific and system-wide regulation of mRNA decay. Mol. Biol. Cell 22, 2787–2795 (2011).
Mathieson, T. et al. Systematic analysis of protein turnover in primary cells. Nat. Commun. 9, 689 (2018).
Deal, C. E., Carfi, A. & Plante, O. J. Advancements in mRNA encoded antibodies for passive immunotherapy. Vaccines 9, 108 (2021).
Van Tendeloo, V. F., Ponsaerts, P. & Berneman, Z. N. mRNA-based gene transfer as a tool for gene and cell therapy. Curr. Opin. Mol. Ther. 9, 423–431 (2007).
Gu, Y., Duan, J., Yang, N., Yang, Y. & Zhao, X. mRNA vaccines in the prevention and treatment of diseases. MedComm 3, e167 (2022).
Sanger, H. L., Klotz, G., Riesner, D., Gross, H. J. & Kleinschmidt, A. K. Viroids are single-stranded covalently closed circular RNA molecules existing as highly base-paired rod-like structures. Proc. Natl Acad. Sci. USA 73, 3852–3856 (1976).
Salzman, J. Circular RNA expression: its potential regulation and function. Trends Genet. 32, 309–316 (2016).
Chen, Y. G. et al. Sensing self and foreign circular RNAs by intron identity. Mol. Cell 67, 228–238.e225 (2017).
Chen, Y. G. et al. N6-methyladenosine modification controls circular RNA immunity. Mol. Cell 76, 96–109.e109 (2019).
Wesselhoeft, R. A. et al. RNA circularization diminishes immunogenicity and can extend translation duration in vivo. Mol. Cell 74, 508–520.e504 (2019).
Loan Young, T., Chang Wang, K., James Varley, A. & Li, B. Clinical delivery of circular RNA: lessons learned from RNA drug development. Adv. Drug. Deliv. Rev. 197, 114826 (2023).
Bai, Y. et al. Research progress on circular RNA vaccines. Front. Immunol. 13, 1091797 (2022).
Munson, M. J. et al. A high-throughput Galectin-9 imaging assay for quantifying nanoparticle uptake, endosomal escape and functional RNA delivery. Commun. Biol. 4, 211 (2021).
Robinson, M. W., Harmon, C. & O’Farrelly, C. Liver immunology and its role in inflammation and homeostasis. Cell Mol. Immunol. 13, 267–276 (2016).
Malik, A. et al. “Complimenting the complement”: mechanistic insights and opportunities for therapeutics in hepatocellular carcinoma. Front. Oncol. 10, 627701 (2021).
Hillebrandt, S. et al. Complement factor 5 is a quantitative trait gene that modifies liver fibrogenesis in mice and humans. Nat. Genet. 37, 835–843 (2005).
Francia, V., Schiffelers, R. M., Cullis, P. R. & Witzigmann, D. The biomolecular corona of lipid nanoparticles for gene therapy. Bioconjug. Chem. 31, 2046–2059 (2020).
Packer, M., Gyawali, D., Yerabolu, R., Schariter, J. & White, P. A novel mechanism for the loss of mRNA activity in lipid nanoparticle delivery systems. Nat. Commun. 12, 6777 (2021). This work describes previously unknown lipid–mRNA reactions that may hamper the translatability and therapeutic effect of mRNA payloads.
Suzuki, Y. & Ishihara, H. Difference in the lipid nanoparticle technology employed in three approved siRNA (Patisiran) and mRNA (COVID-19 vaccine) drugs. Drug. Metab. Pharmacokinet. 41, 100424 (2021).
Sebastiani, F. et al. Apolipoprotein E binding drives structural and compositional rearrangement of mRNA-containing lipid nanoparticles. ACS Nano 15, 6709–6722 (2021). This study shows that binding of ApoE on the surface of LNPs can change their internal structure and cause release of mRNA.
Moghimi, S. M., Haroon, H. B., Yaghmur, A., Simberg, D. & Trohopoulos, P. N. Nanometer- and Ångstrom-scale characteristics that modulate complement responses to nanoparticles. J. Control. Release 351, 432–443 (2022).
Francia, V. et al. Corona composition can affect the mechanisms cells use to internalize nanoparticles. ACS Nano 13, 11107–11121 (2019).
DeLoid, G. M., Cohen, J. M., Pyrgiotakis, G. & Demokritou, P. Preparation, characterization, and in vitro dosimetry of dispersed, engineered nanomaterials. Nat. Protoc. 12, 355–371 (2017).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
E.J. is employed by Moderna, Inc. D.B. is a Northeastern University post-doctoral fellow with a Moderna, Inc.-sponsored fellowship. M.A.R. is currently affiliated with Intellia Therapeutics, but completed this review while working at Moderna.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Glossary
- Adjuvanticity
-
The property of certain substances to enhance the immune response against an antigen, thereby improving the effectiveness of vaccines.
- Biomolecular corona
-
A layer of proteins, lipids and small molecules that forms on the surface of nanoparticles when they interact with biological fluids, influencing their biological identity and activity.
- Drug-induced liver injury
-
Liver damage caused by medications or other xenobiotics, which ranges from small abnormalities in liver tests to severe liver dysfunction or failure.
- Ionizable lipids
-
Lipids that remain neutral at physiological pH but are protonated at low pH and are commonly used in the formulation of lipid nanoparticles for RNA delivery.
- Lipid nanoparticles
-
Nanoparticles made of ionizable and other types of lipid, often used as delivery vehicles for genetic material.
- MicroRNAs
-
Small, non-coding, endogenous RNA molecules that regulate protein synthesis by binding to and destroying specific mRNA, thus inhibiting its translation.
- Pattern recognition receptors
-
Proteins that recognize molecules found in pathogens or released because of cellular damage and that can regulate the innate immune response of cells.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Bitounis, D., Jacquinet, E., Rogers, M.A. et al. Strategies to reduce the risks of mRNA drug and vaccine toxicity. Nat Rev Drug Discov 23, 281–300 (2024). https://doi.org/10.1038/s41573-023-00859-3
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41573-023-00859-3