Abstract
Gene expression is one of the most critical cellular processes. It is controlled by complex mechanisms at the genomic, epigenomic, transcriptomic, and proteomic levels. Any aberration in these mechanisms can lead to dysregulated gene expression. One recently discovered process that controls gene expression includes chemical modifications of RNA molecules by RNA-modifying proteins, a field known as epitranscriptomics. Epitranscriptomics can regulate mRNA splicing, nuclear export, stabilization, translation, or induce degradation of target RNA molecules. Dysregulation in RNA-modifying proteins has been found to contribute to many pathological conditions, such as cancer, diabetes, obesity, cardiovascular diseases, and neurological diseases, among others. This article reviews the role of epitranscriptomics in the pathogenesis and progression of renal cell carcinoma. It summarizes the molecular function of RNA-modifying proteins in the pathogenesis of renal cell carcinoma.
Similar content being viewed by others
Facts
-
Epitranscriptomics play a central role in controlling gene expression essential to the pathogenesis of renal cell carcinoma.
-
Aberrant RNA modifications have been found to contribute to the development of renal cell carcinoma and other types of cancer.
-
An improved understanding of RNA modifications in renal cell carcinoma would undoubtedly contribute to developing new diagnostic and therapeutic strategies.
Open questions
-
What is the effect of aberrant RNA modifications on renal cell carcinoma progression?
-
Are there any potential clinical applications of epitranscriptomics that can assist renal cell carcinoma patients?
-
What are the consequences of the crosstalk between epitranscriptomics and epigenetics at the molecular level?
Introduction
Kidney cancer had an incidence of more than 400,000 cases and more than 179,000 deaths in 2020 [1]. Around 25-30% of kidney cancer cases have metastatic disease at diagnosis time and, therefore, have a limited survival rate [2]. The most prevalent type of kidney cancer is renal cell carcinoma (RCC), accounting for 90% of kidney cancer cases [3]. RCC is a heterogeneous group of epithelial tumors with more than ten histological subtypes [4]. Clear cell RCC (ccRCC) is the most common class of RCC, accounting for 70-80%. Other common types of RCC include papillary RCC (pRCC) and chromophobe RCC (chRCC) [3]. The most commonly mutated genes in ccRCC include the von Hippel–Lindau (VHL) gene and chromatin-remodeling genes such as breast cancer 1 (BRCA1) associated-protein 1 (BAP1), SET domain-containing 2 (SETD2), and polybromo 1 (PBRM1) [5]. VHL regulates hypoxia-inducible factor (HIF) protein. Loss of VHL leads to HIF accumulation, inducing signaling pathways that lead to tumor progression [6]. PBRM1, SETD2, and BAP1 play an essential role in chromatin remodeling and are considered co-drivers of tumor progression [7]. Bromodomain proteins (BRDs), such as bromodomain‑containing protein 9 (BRD9) and bromodomain PHD finger transcription factor (BPTF), also mediate chromatin remodeling in RCC [8].
RCC is characterized by several hallmarks, such as uncontrolled cell growth, apoptosis evasion, angiogenesis, and metabolic reprogramming. Cyclin-dependent protein kinase 2 (CDK2) contributes to impaired cell cycle regulation in RCC [9]. Tumor necrosis factor receptor-associated factor 1 (TRAF1) has antiapoptotic roles [10], and its expression was significantly decreased in RCC [11]. Integrin β4 (ITGB4) is a transmembrane protein that plays a crucial role in promoting angiogenesis [12], thus enhancing tumor invasion and metastasis. In RCC, ITGB4 was overexpressed at advanced stages [13]. Lactate Dehydrogenase A (LDHA) plays a critical role in RCC metabolism and predicts poor prognosis [14]. Other vital players in RCC metabolic reprogramming include solute carrier proteins (SLCs) [15], such as SLC1A5, which has been significantly associated with poor prognosis of ccRCC [16].
Early-stage RCC is managed through nephrectomy, while advanced stages require systemic therapies such as tyrosine kinase inhibitors (TKIs) and immune checkpoint inhibitors (ICIs) [17]. Recently, the combination of cabozantinib, a TKI, and the ICIs nivolumab and ipilimumab significantly increased the progression-free survival of advanced ccRCC patients compared with patients treated with nivolumab and ipilimumab [18]. Although combination therapy showed more effective outcomes, it may induce added toxicity compared to monotherapy [19]. Primary or acquired treatment resistances remain significant clinical challenges [20, 21]. Given these challenges, it is crucial to look for novel therapeutic targets and biomarkers of RCC.
Epitranscriptomics is the study of chemical modifications of RNA molecules occurring after RNA synthesis [22, 23]. Chemical modifications of RNA include N6-methyladenosine (m6A), 5-methylcytosine (m5C), pseudouridine (Ψ), 5-hydroxymethylcytosine (hm5C), and N1-methyladenosine (m1A) [24]. RNA is modified by several types of proteins grouped into three main categories: writers, readers, and erasers. Writer enzymes deposit the chemical modification to the RNA molecule, while erasers remove them. Reader proteins specifically detect and bind to chemically modified RNA molecules [25]. Additionally, RNA is modified by base editing, such as adenosine-to-inosine (A-to-I) and cytosine-to-uridine (C-to-U) editing [23]. Epitranscriptomic modifications were found to be involved in many diseases, including cancer [26], cardiovascular diseases [27], diabetes [28, 29], obesity [28, 30], and major depressive disorder [31].
In this review, RNA modifications related to RCC have been outlined. This review briefly discussed the mode of action of each modification and its role in cancer development with a focus on RCC. Furthermore, the crosstalk between epitranscriptomics and epigenetics, along with the modifications of noncoding RNA in the context of RCC, were reviewed.
Aberrant RNA modifications in RCC
Aberration in RNA modifications leads to RCC progression by induction of cancer hallmarks (Fig. 1). This section reviews different types of RNA modifications and their effects on cancer progression, specifically RCC. The findings of studies conducted to analyze the effects of aberrant RNA modifications on RCC are summarized in Table 1.
N6-methyladenosine
Methylation of RNA at the N6 position of adenosine resulting in m6A is the most characterized and abundant RNA methylation. It is a dynamic modification that involves several proteins acting as writers, readers, and erasers of m6A [26] (Fig. 2). This section will discuss recent findings about proteins involved in m6A deposition, detection, and removal and their roles in RCC progression.
Modification of adenosine to N6-methyladenosine is a dynamic process mediated by the writer enzymes, including METTL3, METTL14, and WTAP. Methylation is catalyzed by SAM as a cofactor and methyl group donor. FTO and ALKBH reverse the process along with the conversion of α-ketoglutarate to succinate. YTHDC1 reads N6-methyladenosine and mediates mRNA splicing and nuclear export. In the cytoplasm, YTHDF1, YTHDF3, and YTHDC2 stabilize the mRNA molecule and mediate its translation, while YTHDF2 mediates mRNA degradation. M6A N6-methyladenosine, METTL methyltransferase-like protein, WTAP Wilms’ tumor 1-associating protein, SAM S-adenosylmethionine, SAH S-adenosylhomocysteine, FTO fat mass and obesity-associated protein, ALKBH ALKB homolog, YTHDC YTH domain-containing protein, YTHDF YTH domain-family protein, IGF2BP Insulin-like growth factor 2 binding protein.
Writers
M6A methyltransferases are proteins that modify RNA with m6A. This reaction is catalyzed by multicomponent methyltransferase complexes. Most of the m6A modifications of RNA molecules occur by methyltransferase like 3 (METTL3), METTL14, and Wilms’ tumor 1-associating protein (WTAP) complex that catalyzes methylation of internal adenosine residues to form m6A [32]. The methylation reaction catalyzed by this complex needs S-adenosylmethionine (SAM) as a cofactor and methyl-group donor. Upon adenosine methylation, SAM is converted to S-adenosylhomocysteine (SAH). METTL3 was identified as the catalytic subunit of this complex, catalyzing the methylation reaction and crosslinking with SAM. The substrate RNA is recognized by the METTL14 molecule, which also plays a role in stabilizing the complex. WTAP provides a scaffolding function and localizes the complex at the nuclear speckles [33, 34]. M6A deposition is guided by histone H3 trimethylation at lysine 36 (H3K36me3). METTL14 binds to H3K36me3 and deposits m6A co-transcriptionally, indicating complex crosstalk between histone modifications and RNA methylation [35].
Depending on the type of cancer, the m6A methyltransferase complex can either suppress or promote tumor growth. For example, METTL3 and METTL14 exhibited an oncogenic role in acute myeloid leukemia (AML), in which they were highly expressed and correlated with shorter survival [36]. Additionally, WTAP was found to play an oncogenic role in AML [37]. In contrast, METTL3 and METTL14 showed a tumor-suppressive role in glioblastoma stem cells [38]. The functions of the components within the m6A writing complex are connected. For example, METTL3 was found to regulate WTAP homeostasis [39]. Interestingly, although WTAP forms a complex with METTL3, many unique target RNAs are associated with only one of the proteins in the complex [34]. Other methyltransferases play a role in m6A methylation. METTL16 methylates mRNAs involved in the SAM synthesis [40]. METTL7B has a tumorigenesis role by inducing cancer cell proliferation in non-small cell lung carcinoma (NSCLC) [41].
METTL14 showed a tumor-suppressive effect in RCC and was downregulated compared to normal controls. Higher expression of METTL14 was associated with better survival [42]. Upon METTL14 downregulation in ccRCC, ITGB4 was overexpressed and promoted invasiveness and metastasis [43]. METTL14 was found to suppress the NUF2 component of NDC80 kinetochore complex (NUF2), cell division cycle associated 3 (CDCA3), and kinesin family member 14 (KIF14), leading to ccRCC progression upon downregulation of METTL14 [44]. Similarly, downregulated METTL14 induced an accumulation of BPTF [45] and P2X purinoceptor 6 (P2RX6), an ATP receptor [46], driving metastasis and invasion in ccRCC. Therefore, methylation induced by METTL14 suppressed BPTF and P2RX6 expression. In contrast to the previous studies, METTL14 stabilized PTEN, leading to tumor progression inhibition and suggesting a potential therapeutic target [47].
METTL7B was found to be upregulated in ccRCC compared to normal controls. Its expression was significantly associated with tumor size, lymph node metastasis, and poor prognosis [48]. In contrast, METTL7A expression was considerably lower in renal cancer than in normal tissues, and its low expression was associated with poor prognosis [49]. METTL3 played a tumor-suppressive role in the RCC, and its downregulation was associated with larger tumor size, higher histological grade, and poor survival [50, 51]. In RCC, WTAP was found to maintain the stability of CDK2 mRNA, which in turn promotes the proliferation of RCC cells [52]. Additionally, WTAP mediated m6A modification of sphingosine-1-phosphate receptor 3 (S1PR3). S1PR3 was found to induce RCC proliferation and metastasis by activating the phosphatidylinositol 3‑kinase (PI3K)/protein kinase B (AKT) pathway [53].
Readers
Reader proteins detect m6A and create a biological signal. Mainly, these proteins belong to 4 families: YT521-B homology (YTH) domain-containing proteins, insulin-like growth factor 2 binding protein (IGF2BP) family, heterogeneous nuclear ribonucleoprotein (hnRNP), and proline-rich and coiled-coil-containing protein 2 A (PRRC2A) [26]. Most of the studies that investigated RCC were conducted on YTH domain-containing proteins, IGF2BP, and hnRNP, which will be the focus of this section.
YTH domain-containing proteins
The YTH domain-containing proteins include YTH domain family proteins (YTHDFs) and YTH domain-containing proteins (YTHDCs). In eukaryotes, these proteins are expressed by five genes: YTHDF1, YTHDF2, YTHDF3, YTHDC1, and YTHDC2. After recognizing m6A modification of target genes, YTH domain-containing proteins recruit different complexes to regulate several processes, such as RNA translation, RNA decay, RNA splicing, and nuclear export. Furthermore, they play various roles in cancer development and progression [54].
YTHDF2 was the first characterized member of YTH domain-containing proteins. It was found to promote target decay. YTHDF2 was upregulated in lung adenocarcinoma and was found to degrade axis inhibition protein 1 (AXIN1), which encodes a negative regulator of the Wnt/β-catenin signaling, eventually leading to Wnt/β-catenin activation and tumor progression [55]. In contrast, YTHDF1 and YTHDF3 were found to stabilize their targets and promote translation. Upregulation of YTHDF1 was found to be associated with poor prognosis of ovarian cancer. Mechanistically, YTHDF1 was found to augment the translation of eukaryotic translation initiation factor 3 subunit C (EIF3C) in an m6A-dependent manner. Thus promoting the overall translation output, which in turn initiates ovarian cancer tumorigenesis and metastasis [56]. Moreover, YTHDF3 overexpression was clinically correlated with brain metastasis of breast cancer by promoting the translation of crucial brain metastatic genes ST6 N-acetylgalactosaminide alpha-2,6-sialyltransferase 5 (ST6GALNAC5) and gap junction protein alpha 1 (GJA1) [57]. YTHDC1 was found to play roles in mRNA splicing and nuclear export, while YTHDC2 was found to promote translation [54]. In triple-negative breast cancer, YTHDC1 had an oncogenic role by promoting SMAD3 mRNA nuclear export and expression to augment the transforming growth factor-β (TGF-β) signaling cascade [58]. YTHDC2 promoted gastric cancer progression via increasing the translation of yes-associated protein (YAP) oncogene [59].
In ccRCC, the expression of YTHDF1-3 and YTHDC1 were significantly down-regulated compared to the normal tissue [60]. YTHDF1 played a role in stabilizing m6A-modified PTEN and was found to suppress tumor progression in ccRCC by inhibiting the activation of the PI3K/AKT signaling pathway [47]. Moreover, YTHDF2 was found to be downregulated in ccRCC and acted by promoting the decay of ITGB4 mRNA [43]. Additionally, in NONO-TFE3 translocation renal cell carcinoma (NONO-TFE3 tRCC), a subtype of RCC associated with Xp11.2 translocation/TFE3 gene fusions RCC (Xp11.2 tRCCs), YTHDF2 played a tumor-suppressive role by suppressing poly(ADP-ribose) polymerase 1 (PARP1) expression. The downregulation of YTHDF2 promoted cell proliferation, invasion, and migration [61].
Insulin-like growth factor 2 binding proteins
IGF2BPs, also known as insulin mRNA binding proteins (IMPs), are another group of m6A readers, consisting of IGF2BP1, IGF2BP2, and IGF2BP3. IGF2BPs are expressed in most embryonic tissues and play crucial roles in embryogenesis by controlling RNA localization, stability, and translation [62, 63]. IGF2BPs recognize m6A-modified RNAs through their K homology (KH) domain and function by stabilizing their target RNA [64]. In adults, IGF2BP2 is widely expressed in different tissues. In contrast, the expression levels of IGF2BP1 and IGF2BP3 are negligible in adults except in reproductive organs [65]. Therefore, they are considered oncofetal proteins since they are severely upregulated in various tumors.
IGF2BPs were found to be overexpressed and play oncogenic roles in different tumor types. They promote cancer progression by stabilizing methylated mRNAs of oncogenic targets [66]. In breast cancer, IGF2BP1 was found to maintain the stability of m6A-modified c-Myc mRNA in vivo [67]. IGF2BP2 and IGF2BP3 enhanced the stability of methylated ephrin type-A receptor 2 (EPHA2) and vascular endothelial growth factor A (VEGFA) mRNAs, respectively, in colorectal cancer cells. EPHA2 and VEGFA activate both PI3K/AKT and the extracellular signal-regulated kinase 1/2 (ERK1/2) signaling pathways to induce tumor progression by increasing tumor cell proliferation and vasculogenic mimicry [68]. Vasculogenic mimicry occurs when tumor cells form microvascular channels that provide blood supply in aggressive tumors independently of tumor angiogenic mechanisms [69].
Overexpression of IGF2BP3 in ccRCC was associated with advanced stage and grade of primary tumors, coagulative tumor necrosis, and sarcomatoid differentiation [70]. IGF2BP3 was significantly overexpressed in metastatic RCC and primary RCC that were likely to develop metastasis. It was suggested as a diagnostic marker to identify patients with a high risk of developing metastasis [70,71,72,73,74]. Similarly, patients with localized pRCC and chRCC overexpressing IGF2BP3 were over ten times more likely to develop metastasis than patients with low IGF2BP3 expression [75]. Moreover, high levels of circulating IGF2BP3 were detected in RCC patients with high-grade and more aggressive tumors and were independently associated with poor survival. Hence, it suggests its potential application as a minimally invasive sampling method that can improve therapeutic planning [76].
In RCC, the expression of IGF2BPs could be increased by early growth response 2 (EGR2) transcription factor. IGF2BPs, in turn, enhance the stability of the S1PR3 mRNA [53]. IGF2BP1 was upregulated in ccRCC cell lines and facilitated tumor energy metabolism by promoting glycolysis [77]. Moreover, the upregulation of IGF2BP3 activated the nuclear factor kappa B (NF-кB) pathway, which promoted RCC cell migration and invasion [78]. Furthermore, IGF2BP2 was involved in stabilizing TRAF1 by m6A RNA methylation via METTL14 in sunitinib-resistant RCC cells. Ultimately, this led to suppressing the apoptotic and antiangiogenic effects of sunitinib [79]. Overall, these studies suggest the oncogenic role of IGF2BPs in RCC, and due to their oncofetal transcription, they are considered a potential therapeutic target.
Heterogeneous nuclear ribonucleoproteins
Another group of m6A reader proteins includes hnRNPs, which are responsible for pre-mRNA maturation into functional mRNA, mRNA stabilization, and translocation. Up to date, twenty members of the hnRNP family have been identified and termed hnRNP A-U [80]. HnRNPs were found to be involved in apoptosis, epithelial-mesenchymal transition (EMT), and cancer angiogenesis [81]. HnRNPC was upregulated in pRCC, promoting cell proliferation and migration in vitro [82]. More extensive studies are needed to investigate the role of hnRNP proteins in RCC pathogenesis.
Erasers
M6A erasers remove methyl groups from RNA bases. This group of RNA-modifying proteins includes members of the AlkB homolog (ALKBH) family, which consists of ALKBH 1-8 and fat mass and obesity-associated protein (FTO). They catalyze the demethylation of nucleic acid bases by oxidation reaction dependent on iron (II) and α-ketoglutarate [83]. The demethylation reaction converts α-ketoglutarate to succinate, which is accompanied by the release of formaldehyde and carbon dioxide [84].
The expression level of ALKBH genes varies in different cancers. A tumor-suppressive effect of ALKBH5 was detected in pancreatic cancer by preventing its progression by activating period circadian regulator 1 (PER1), which in turn inhibited cell growth [85]. Moreover, ALKBH5 inhibited the metastasis of colon cancer [86]. On the contrary, ALKBH5 promoted invasion and metastasis in gastric cancer [87]. In addition, it inhibited autophagy of epithelial ovarian cancer by enhancing the stability of BCL2 apoptosis regulator (BCL2), which has an antiapoptotic role [88]. Additionally, ALKBH5 promoted NSCLC progression by repressing tissue inhibitors of metalloproteinase 3 (TIMP-3) [89].
ALKBH5 was differentially expressed between RCC subtypes and oncocytomas, suggesting its potential application as a diagnostic marker [90]. Studies have shown inconsistent results regarding its expression in RCC and adjacent normal tissue. Some studies suggested that the expression of ALKBH5 was significantly downregulated in ccRCC compared to normal tissue. Additionally, it was correlated with shortened overall and cancer-specific survival [91]. On the other hand, other studies suggested an oncogenic role of ALKBH5, which was overexpressed in RCC. Moreover, elevated expression of ALKBH5 was correlated with larger tumor volume, higher TNM staging, and worse prognosis. Mechanistically, ALKBH5 expression was upregulated by HIF induced by hypoxia. ALKBH5 stabilized aurora kinase B (AURKB), promoting RCC cell proliferation [92]. Additionally, ALKBH5 was found to promote cell proliferation and migration through cell cycle alteration and EMT [93]. A similar oncogenic effect was detected in ALKBH3 in RCC patients, in which its expression was positively correlated with advanced TNM staging and poor prognosis [94]. Moreover, ALKBH1 had an oncogenic role in RCC cell lines and promoted cell migration and viability [95]. Overall, further studies are required to elucidate the molecular mechanisms behind the different roles of ALKBH in RCC.
Like ALKBH, FTO also showed conflicting effects, with studies reporting tumor-suppressive effects while others reported oncogenic effects. Jeschke et al. found that FTO presented a tumor-suppressive role in many epithelial tumors, including breast, prostate, cervical, liver, and lung cancers. This was shown, upon FTO depletion, by the induction of Wnt signaling and EMT transition [96]. Similarly, FTO exhibited a suppressive effect on prostate cancer proliferation and metastasis by stabilizing chloride intracellular channel 4 (CLIC4), which encodes a protein that can inhibit cell proliferation through the TGF-β pathway [97].
On the contrary, a recent study utilized epitranscriptomic landscape mapping and revealed that positive FTO expression was associated with poor survival of breast cancer [98]. FTO was elevated in gastric cancer and showed an oncogenic effect by promoting cell proliferation and metastasis via inducing the degradation of caveolin-1 (CAV1) mRNA by demethylation. As a result of CAV1 degradation, mitochondrial dynamics are altered, leading to an elevated ATP level and thereby favoring cancer growth [99]. Another example of the oncogenic role of FTO was observed in pancreatic cancer, where FTO induced cancer progression via stabilizing the mRNA of platelet-derived growth factor C (PDGFC). Eventually, this led to the reactivation of the PI3K/AKT signaling pathway that promoted cell growth [100].
The dichotomous effect of FTO was also observed in RCC. Decreased FTO level correlated with increased tumor severity and poor overall and cancer-specific survival following nephrectomy, suggesting a tumor-suppressive role of FTO [91, 101]. Moreover, in VHL-deficient ccRCC cells with ectopic FTO expression, the expression of peroxisome proliferator-activated receptor γ coactivator 1 α (PGC-1α) increased due to decreased m6A levels in its transcript. PGC-1α restored mitochondrial activity, which was revealed by an elevated ATP level and induced oxidative stress and ROS production. Consequently, this resulted in impaired tumor growth [101]. This finding contradicts what was found in gastric cancer, where the ATP reduction induced by FTO depletion restricted cancer growth [99]. This could be due to the opposing effects of ATP within the tumor microenvironment, where it can promote or inhibit cancer growth depending on its concentration, receptors expressed by cancer and immune cells, and the expression of ectonucleotidase enzymes that hydrolyze ATP [102, 103].
Besides its tumor-suppressive effect in RCC, FTO also exhibited an oncogenic effect. It modulated EMT and cell cycle, promoting cell proliferation and migration in RCC cell lines [93]. A study conducted by Xiao et al. found that FTO was overexpressed in ccRCC tumors with VHL deletions or mutations compared to adjacent normal tissue. Additionally, they detected a synthetic lethal interaction between FTO and VHL. FTO inhibition selectively reduced the growth of VHL-deficient cells in vitro and in vivo in a HIF-independent manner. Furthermore, they identified the glutamine transporter SLC1A5 as a target of FTO, leading to the metabolic reprogramming and survival of VHL-deficient cells [104]. Moreover, FTO showed an oncogenic role in HIF2αlow/− ccRCC by stabilizing the mRNA of BRD9. Inhibition of BRD9 in BALB/c mice bearing HIF2αlow/− ccRCC cell line–derived xenografts and patient-derived tumor xenografts led to tumor growth inhibition and prolonged survival with greater efficacy than sunitinib [105]. This indicates that cells with different genetic backgrounds respond to FTO differently. The context-dependent role of FTO as oncogenic or tumor-suppressive is complex and requires further investigation of its underlying molecular mechanisms.
5-Methylcytosine (m5C)
Methylation at position 5 of cytidine residue can occur in mRNA, rRNA, tRNA, and other noncoding RNA molecules [26]. Similar to m6A modifications, there is a group of m5C writers, readers, and erasers. M5C writers include seven members of the NOL1/NOP2/SUN domain family member (NSUN) family, NSUN1 to NSUN7, and DNA methyltransferase-2 (DNMT2). Reader proteins that bind to m5C include YTHDF2, Aly/REF export factor (ALYREF), and Y-box binding protein 1 (YBX1) [106]. ALYREF mediates the nuclear export of mRNA, while YBX1 regulates mRNA stability in the cytoplasm [107]. M5C demethylation occurs via the eraser molecule ten-eleven translocation (TET) [106] (Fig. 3).
Modification of cytosine to 5-methylcytosine is mediated by NSUN proteins. This process is reversed by TET. ALYREF is a reader protein that mediates the nuclear export of mRNA, while YBX1 regulates mRNA stability in the cytoplasm. M5C 5-methylcytosine, NSUN NOL1/NOP2/SUN domain family member, TET ten-eleven translocation, ALYREF Aly/REF export factor, YBX1 Y-box binding protein 1.
M5C was found to alter the progression of different types of cancers. In breast cancer, the overexpression of NSUN2 was correlated with increased metastasis and invasion [108]. ALYREF and NSUN5 were overexpressed at the metastasis stage of head and neck squamous cell carcinoma [109]. In ccRCC, NSUN1, NSUN2, NSUN5, and NSUN6 were upregulated, while NSUN4 and TET2 were downregulated [110, 111]. NSUN5 promoted ccRCC progression through the Warburg effect and increased cell growth through stabilizing enolase 3 (ENO3) mRNA [112]. Warburg effect is the phenomenon of cells undergoing anaerobic glycolysis for energy production even under normal oxygen concentration [113]. Another study on ccRCC revealed the role of YBX1 in stabilizing phosphatidylethanolamine binding protein 1 (PEBP1). YBX1 is recruited to m5C-containing PEBP1 mRNA by phosphatidylethanolamine binding protein 1 pseudogene 2 (PEBP1P2). The low expression of PEBP1P2 is correlated with poor prognosis and advanced stages of the ccRCC [114]. Further studies are required to elaborate on the molecular mechanism of m5C regulators and how they affect RCC progression.
Pseudouridine
Uridine can be converted to pseudouridine (Ψ), a C5-glycoside isomer of uridine, by Ψ synthase. Ψ was initially detected in rRNA and tRNA. Later, it was also detected in mRNA, long noncoding RNA (lncRNA), and small nuclear RNA (snRNA) molecules [115]. It is involved in several physiological roles depending on the modified RNA molecule. It influences rRNA folding and ribosome assembly in rRNA and alters tRNA interaction with rRNA and mRNA [116]. In mRNA, Ψ affects stability, promotes pre-mRNA splicing, and mediates translation [117]. In eukaryotes, dyskerin pseudouridine synthase 1 (DKC1) and several proteins belonging to the pseudouridine synthase (PUS) family function as Ψ writers (Fig. 4) [26]. In breast cancer, DKC1 overexpression predicted poor prognosis [118]. Furthermore, PUS7 was upregulated in ovarian cancer and has been suggested as a potential biomarker [119]. In ccRCC, DKC1 was found to have oncogenic roles by promoting cell proliferation, migration, and invasion via the NF-κB pathway [120].
RNA editing
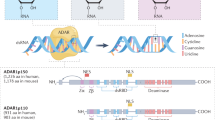
In addition to the chemical modifications of RNA bases that do not affect RNA sequence, RNA bases can be modified by deamination, resulting in the conversion of the RNA base type. There are two types of RNA editing: A-to-I editing and C-to-U editing (Fig. 5) [23].
Adenosine to inosine editing
A-to-I editing is the most prevalent RNA editing in vertebrates. It is mediated by adenosine deaminase acting on dsRNA 1 (ADAR1) and ADAR2. ADARs were found to be associated with splicing factors, indicating their potential roles in alternative splicing and transcriptional control [121]. Additionally, ADARs target RNA viruses by inducing A-to-I hypermutations. They have both pro- and anti-viral effects [122]. Moreover, ADARs were found to play roles in cancer progression. In breast cancer, upregulation of ADAR1 promoted cancer progression through cell cycle regulation and controlling DNA damage response [123]. Based on data from the Cancer Genome Atlas (TCGA), there was no significant difference in ADAR1 expression in ccRCC compared to normal renal tissue, and it did not have an impact on patients’ survival [124].
Cytosine to uridine editing
Apolipoprotein B mRNA-editing cytosine deaminase (APOBEC) catalyzes cytosine deamination mediating C-to-U conversion. APOBEC family includes activation-induced cytidine deaminase (AID), APOBEC1 (A1), APOBEC2 (A2), APOBEC3 (A3), and APOBEC4 (A4). A3 comprises seven members– A3A, A3B, A3C, A3D, A3F, A3G, A3H [125]. Members of the APOBEC family play various physiological roles, and all of them have C-to-U deaminase activity except A2 and A4 [126]. A1, A3A, and A3G were found to act on single-stranded RNA as a substrate [127]. A1 regulates lipid metabolism by mediating dietary lipid uptake from the intestine [128]. A3 deaminates viral DNA or RNA, leading to viral restriction, which is the degradation of the viral genome when the mutational load is so high that the genome cannot function properly [127, 129]. Deamination of A3 can support viral evolution provided it does not lead to viral restriction and the resulting mutations are fixed in the viral genome [127].
Besides its role in viral restriction and evolution, APOBEC has a similar effect on cancer progression, where it can lead to tumor restriction or tumor evolution. Deamination induced by APOBEC initiates mutagenesis in cancer cells, leading to autonomous lethality and tumor restriction. Mutations introduced by APOBEC can lead to chromosomal instability. Pecori et al. hypothesized that the inflammatory microenvironment of cancer leads to elevated expression of A3 that induces localized hypermutation attempting to kill malignant cells [127].
APOBEC proteins were also involved in generating tumor heterogeneity when the level of the introduced mutations was low. This will eventually lead to tumor progression [127]. For example, inhibition of lncRNA H19 and its target A3G due to sulforaphane treatment inhibited pancreatic cancer progression in vitro and in vivo by inhibiting TGF-β-induced SMAD2 phosphorylation [130]. This indicates that despite the mutation surge induced by APOBEC that promotes the evolution of more aggressive clones, it also provides potential targets for cytotoxic and immunotherapies [131]. Furthermore, A3G was found to be an unfavorable prognostic marker in ccRCC patients. The expression of A3G was positively correlated with the expression of several immunoinhibitors and the presence of immunosuppressive cells [132].
The crosstalk between epigenetics and RNA modifications
Recently, many studies have shown an integration between epitranscriptomic and epigenetic modifications, such as histone modifications [133] and DNA methylation [134]. This integration has been observed in many physiological and pathogenic processes and can influence chromatin accessibility and transcription regulation. METTL3 has been found to play a role in mammalian development by regulating the heterochromatin of mouse embryonic stem cells [135]. In glioblastoma, METTL3 modified and promoted the expression of genes involved in histone modifications in an m6A-dependent manner [136]. Moreover, METTL3 was found to facilitate the demethylation of nearby genomic DNA in an m6A-dependent manner in cancer and normal cells. Upon RNA methylation by METTL3, fragile-X mental retardation autosomal 1 (FXR1) protein recognizes m6A and recruits TET1 protein to demethylate DNA in a process called RNA methylation-coupled DNA demethylation [134].
An example of the crosstalk between epigenetic and epitranscriptomic regulations in ccRCC is the interaction between IGF2BP3 and a lncRNA cofactor called DNA methylation–deregulated and RNA m6A reader–cooperating lncRNA (DMDRMR). DNA hypomethylation of its promoter region induces DMDRMR expression. IGF2BP3 and DMDRMR stabilized target mRNAs, such as cyclin-dependent kinase 4 (CDK4) and three extracellular matrix components: collagen type VI alpha 1 chain (COL6A1), laminin subunit alpha 5 (LAMA5), and fibronectin 1 (FN1). Consequently, activation of CDK4 led to accelerating ccRCC cell proliferation, and FN1 partially promoted invasion and metastasis. Moreover, the elevated expression of IGF2BP3 and DMDRMR was associated with poor overall survival [137].
Another component of the epigenetic machinery includes noncoding RNA molecules. Initially, scientists thought that RNA modifications occur only in mRNA molecules. With the advance in detection technologies, recent studies detected modifications in noncoding RNA, such as rRNAs, tRNAs, microRNAs (miRNAs), lncRNAs, and small nucleolar RNAs [32]. Additionally, noncoding RNAs can regulate the expression of RNA-modifying proteins. In RCC, several studies detected noncoding RNAs as regulators or targets of RNA-modifying proteins, eventually controlling the expression of downstream genes involved in tumorigenesis (Table 2).
Noncoding RNAs as regulators
WTAP is targeted and silenced by miR-501-3p, which was found to be downregulated in RCC. The overexpression of miR-501-3p inhibited disease progression [138]. Another inhibitory effect of miRNA was found against IGF2BP1, which was inhibited by miR-372 in RCC by direct interaction with its putative binding site at 3’-UTR. In RCC cell lines and tissue samples, miRNA-372 was down-regulated, and when miRNA-372 was overexpressed, it inhibited RCC cell proliferation and invasion. This suggests that miRNA-372 has therapeutic potential in the treatment of RCC [139]. Moreover, ccRCC cell lines that were treated with miR-155 exhibited an inhibition of FTO, resulting in increased tumor cell proliferation [140].
The lncRNA TRAF3IP2 antisense RNA 1 (TRAF3IP2-AS1) functions by binding to PARP1 mRNA and recruiting m6A methyltransferase complex consisting of METTL3, METTL14, and WTAP, leading to PARP1 degradation. In NONO-TFE3 tRCC, TRAF3IP2-AS1 was downregulated, which resulted in PARP1 accumulation and promoted tumorigenesis [61].
Noncoding RNAs as targets
In ccRCC, IGF2BP3 was found to stabilize a lncRNA called cyclin-dependent kinase inhibitor 2B antisense 1 (CDKN2B-AS1). CDKN2B-AS1 was significantly upregulated in ccRCC and participated in epigenetic activation of NUF2. Thus, it enhanced NUF2 transcription, which promoted tumor cell growth and metastasis both in vitro and in vivo. Patients with elevated IGF2BP3, CDKN2B-AS1, and NUF2 showed reduced survival time [141].
Furthermore, epithelial splicing regulatory protein 2 (ESRP2) inhibits metastasis and is ubiquitinated by lnc-LSG1, causing ESRP2 protein degradation. METTL14 was found to target lnc-LSG1 by m6A modification, which inhibits its binding to ESRP2 via YTHDC1. As a result, ESRP2 will be protected from degradation and inhibit metastasis. In ccRCC, METTL14 is downregulated, leading to the binding of ESRP2 to lnc-LSG1 and ESRP2 degradation [142].
Moreover, METTL14 was found to downregulate the expression of the oncogenic long noncoding RNA nuclear enriched abundant transcript 1_1 (NEAT1_1) in an m6A-dependent manner. YTHDF2 detected m6A modification on NEAT1_1, leading to NEAT1_1 degradation. Downregulation of METTL14 in ccRCC will lead to the stabilization of the NEAT1_1, resulting in a malignant phenotype [143].
Conclusion and future perspectives
Several decades following the discovery of RNA modifications, the role of RNA modifications in cancer was not fully understood [144]. Recently, the advancement of detection methods of RNA modifications led to a dramatic burst in identifying their functional roles in health and disease. The varying functions of some RNA-modifying proteins, which can either promote or inhibit RCC progression, provide a better understanding of the heterogenous RCC environment. The aberrant expression of RNA-modifying proteins has been found to induce cancer hallmarks in RCC, such as cell proliferation, metabolic programming, angiogenesis, invasion, and metastasis. Moreover, RNA modifications play an essential role in developing several physiological conditions that can increase the risk of RCC, such as obesity, diabetes, and inflammatory responses. Furthermore, epitranscriptomics modifications are dynamic processes; therefore, manipulating modifications could lead to tumor reversion, offering a promising therapeutic strategy [98]. In conclusion, an improved understanding of RNA modifications in cancer may contribute to the development of new diagnostic and therapeutic strategies for RCC.
Reporting summary
Further information on research design is available in the Nature Research Reporting Summary linked to this article.
References
Kidney cancer statistics: World Cancer Research Fund International; [Available from: https://www.wcrf.org/cancer-trends/kidney-cancer-statistics/.
Huang J, Leung DKW, Chan EOT, Lok V, Leung S, Wong I, et al. A global trend analysis of kidney cancer incidence and mortality and their associations with smoking, alcohol consumption, and metabolic syndrome. Eur Urol Focus. 2022;8:200–9.
Braga EA, Fridman MV, Loginov VI, Dmitriev AA, Morozov SG. Molecular mechanisms in clear cell renal cell carcinoma: role of miRNAs and hypermethylated miRNA genes in crucial oncogenic pathways and processes. Front Genet. 2019;10:320.
Feng C, Huang X, Li X, Mao J. The roles of base modifications in kidney cancer. Front Oncol. 2020;10:580018.
Dizman N, Philip EJ, Pal SK. Genomic profiling in renal cell carcinoma. Nat Rev Nephrol. 2020;16:435–51.
Shen C, Kaelin WG. The VHL/HIF axis in clear cell renal carcinoma. Semin Cancer Biol. 2013;23:18–25.
Walton J, Lawson K, Prinos P, Finelli A, Arrowsmith C, Ailles L. PBRM1, SETD2 and BAP1—the trinity of 3p in clear cell renal cell carcinoma. Nat Rev Urol. 2023;20:96–115.
Wen Q, Liu H, Lou K, Zhang X, Chao W, Xin J, et al. Essential role of bromodomain proteins in renal cell carcinoma (Review). Mol Med Rep. 2023;28:139.
Hongo FMDPD, Takaha NMDPD, Oishi MMDPD, Ueda TMDPD, Nakamura TMDPD, Naitoh YMDPD, et al. CDK1 and CDK2 activity is a strong predictor of renal cell carcinoma recurrence. Urol Oncol. 2014;32:1240–6.
Edilova MI, Abdul-Sater AA, Watts TH. TRAF1 signaling in human health and disease. Front Immunol. 2018;9:2969.
Rajandram R, Bennett NC, Wang Z, Perry-Keene J, Vesey DA, Johnson DW, et al. Patient samples of renal cell carcinoma show reduced expression of TRAF1 compared with normal kidney and functional studies in vitro indicate TRAF1 promotes apoptosis: potential for targeted therapy. Pathology. 2012;44:453–9.
Nikolopoulos SN, Blaikie P, Yoshioka T, Guo W, Giancotti FG. Integrin β4 signaling promotes tumor angiogenesis. Cancer Cell. 2004;6:471–83.
Huang W, Fan L, Tang Y, Chi Y, Li J. A pan-cancer analysis of the oncogenic role of integrin Beta4 (ITGB4) in Human Tumors. Int J Gen Med. 2021;14:9629–45.
Girgis H, Masui O, White NM, Scorilas A, Rotondo F, Seivwright A, et al. Lactate Dehydrogenase A is a potential prognostic marker in clear cell renal cell carcinoma. Mol Cancer. 2014;13:101.
El-Gebali S, Bentz S, Hediger MA, Anderle P. Solute carriers (SLCs) in cancer. Mol Asp Med. 2013;34:719–34.
Liu Y, Yang L, An H, Chang Y, Zhang W, Zhu Y, et al. High expression of Solute Carrier Family 1, member 5 (SLC1A5) is associated with poor prognosis in clear-cell renal cell carcinoma. Sci Rep. 2015;5:16954.
Yang J, Wang K, Yang Z. Treatment strategies for clear cell renal cell carcinoma: past, present and future. Front Oncol. 2023;13:1133832.
Choueiri TK, Powles T, Albiges L, Burotto M, Szczylik C, Zurawski B, et al. Cabozantinib plus Nivolumab and Ipilimumab in Renal-Cell Carcinoma. NEJM. 2023;388:1767–78.
Patel H, Shinder B, Srinivasan R, Singer E. Challenges and opportunities in the management of metastatic renal cell carcinoma: combination therapy and the role of cytoreductive surgery. Curr Opin Oncol. 2020;32:240–9.
Elgendy M, Fusco JP, Segura V, Lozano MD, Minucci S, Echeveste JI, et al. Identification of mutations associated with acquired resistance to sunitinib in renal cell cancer. Int J Cancer. 2019;145:1991–2001.
Numakura K, Sekine Y, Hatakeyama S, Muto Y, Sobu R, Kobayashi M, et al. Primary resistance to nivolumab plus ipilimumab therapy in patients with metastatic renal cell carcinoma. Cancer Med. 2023;12:16837–45.
Kan RL, Chen J, Sallam T. Crosstalk between epitranscriptomic and epigenetic mechanisms in gene regulation. Trends Genet. 2022;38:182–93.
Christofi T, Zaravinos A. RNA editing in the forefront of epitranscriptomics and human health. J Transl Med. 2019;17:319.
Song J, Yi C. Chemical modifications to RNA: a new layer of gene expression regulation. ACS Chem Biol. 2017;12:316–25.
Esteve-Puig R, Bueno-Costa A, Esteller M. Writers, readers and erasers of RNA modifications in cancer. Cancer Lett. 2020;474:127–37.
Barbieri I, Kouzarides T. Role of RNA modifications in cancer. Nat Rev Cancer. 2020;20:303–22.
Leptidis S, Papakonstantinou E, Diakou KI, Pierouli K, Mitsis T, Dragoumani K, et al. Epitranscriptomics of cardiovascular diseases. Int J Mol Med. 2022;49:1–21.
Kung C-P, Maggi LB Jr, Weber JD. The role of RNA editing in cancer development and metabolic disorders. Front Endocrinol. 2018;9:762.
Geng X, Li Z, Yang Y. Emerging role of epitranscriptomics in diabetes mellitus and its complications. Front Endocrinol. 2022;13:907060.
Sanoudou D, Gkouskou KK, Eliopoulos AG, Mantzoros CS. Epitranscriptomic challenges and promises in metabolic diseases. Metab Clin Exp. 2022;132:155219.
Blaze J, Plaza-Jennings A, Turecki G, Haghighi F, Akbarian S. Epitranscriptomic and metabolomic signatures involved in translational mechanisms of suicide and depression. Biol Psychiatry. 2022;91:S73–S74.
Zhao W, Qi X, Liu L, Ma S, Liu J, Wu J. Epigenetic regulation of m6A modifications in human cancer. Mol Ther Nucleic Acids. 2020;19:405–12.
Wang X, Huang J, Zou T, Yin P. Human m6A writers: two subunits, 2 roles. RNA Biol. 2017;14:300–4.
Ping X-L, Sun B-F, Wang L, Xiao W, Yang X, Wang W-J, et al. Mammalian WTAP is a regulatory subunit of the RNA N6-methyladenosine methyltransferase. Cell Res. 2014;24:177–89.
Huang H, Weng H, Zhou K, Wu T, Zhao BS, Sun M, et al. Histone H3 trimethylation at lysine 36 guides m6A RNA modification co-transcriptionally. Nature. 2019;567:414–9.
Sang L, Wu X, Yan T, Naren D, Liu X, Zheng X, et al. The m6A RNA methyltransferase METTL3/METTL14 promotes leukemogenesis through the mdm2/p53 pathway in acute myeloid leukemia. J Cancer. 2022;13:1019–30.
Bansal H, Yihua Q, Iyer SP, Ganapathy S, Proia DA, Proia D, et al. WTAP is a novel oncogenic protein in acute myeloid leukemia. Leukemia. 2014;28:1171–4.
Cui Q, Shi H, Ye P, Li L, Qu Q, Sun G, et al. m6A RNA methylation regulates the self-renewal and tumorigenesis of glioblastoma stem cells. Cell Rep. 2017;18:2622–34.
Sorci M, Ianniello Z, Cruciani S, Larivera S, Ginistrelli LC, Capuano E, et al. METTL3 regulates WTAP protein homeostasis. Cell Death Dis. 2018;9:796.
Pendleton KE, Chen B, Liu K, Hunter OV, Xie Y, Tu BP, et al. The U6 snRNA m6A methyltransferase METTL16 regulates SAM synthetase intron retention. Cell. 2017;169:824–35.e814.
Liu D, Li W, Zhong F, Yin J, Zhou W, Li S, et al. METTL7B is required for cancer cell proliferation and tumorigenesis in non-small cell lung cancer. Front Pharm. 2020;11:178.
Wang Y, Cong R, Liu S, Zhu B, Wang X, Xing Q. Decreased expression of METTL14 predicts poor prognosis and construction of a prognostic signature for clear cell renal cell carcinoma. Cancer Cell Int. 2021;21:46.
Liu Z, Sun T, Piao C, Zhang Z, Kong C. METTL14-mediated N6-methyladenosine modification of ITGB4 mRNA inhibits metastasis of clear cell renal cell carcinoma. Cell Commun Signal. 2022;20:36.
Yang Z, Peng B, Pan Y, Gu Y. Analysis and verification of N6-methyladenosine-modified genes as novel biomarkers for clear cell renal cell carcinoma. Bioengineered. 2021;12:9473–83.
Zhang C, Chen L, Liu Y, Huang J, Liu A, Xu Y, et al. Downregulated METTL14 accumulates BPTF that reinforces super-enhancers and distal lung metastasis via glycolytic reprogramming in renal cell carcinoma. Theranostics. 2021;11:3676–93.
Gong D, Zhang J, Chen Y, Xu Y, Ma J, Hu G, et al. The m6A-suppressed P2RX6 activation promotes renal cancer cells migration and invasion through ATP-induced Ca2+ influx modulating ERK1/2 phosphorylation and MMP9 signaling pathway. J Exp Clin Cancer Res. 2019;38:233.
Zhang L, Luo X, Qiao S. METTL14-mediated N6-methyladenosine modification of Pten mRNA inhibits tumour progression in clear-cell renal cell carcinoma. Br J Cancer. 2022;127:30–42.
Li W, Xu S, Peng N, Zhang Z, He H, Chen R, et al. Downregulation of METTL7B Inhibits Proliferation of Human Clear Cell Renal Cancer Cells In Vivo and In Vitro. Front Oncol. 2021;11:634542.
Yang Z, Zhang W, Li L, Hu N, Dong X, Chen Y, et al. The novel putative methyltransferase METTL7A as one prognostic biomarker potentially associated with immune infiltration in human renal cancer. Heliyon. 2023;9:e15371.
Li X, Tang J, Huang W, Wang F, Li P, Qin C, et al. The M6A methyltransferase METTL3: acting as a tumor suppressor in renal cell carcinoma. Oncotarget. 2017;8:96103–16.
Gundert L, Strick A, Hagen F, Schmidt D, Klümper N, Tolkach Y, et al. Systematic expression analysis of m6A RNA methyltransferases in clear cell renal cell carcinoma. BJUI Compass. 2021;2:402–11.
Tang J, Wang F, Cheng G, Si S, Sun X, Han J, et al. Wilms’ tumor 1-associating protein promotes renal cell carcinoma proliferation by regulating CDK2 mRNA stability. J Exp Clin Cancer Res. 2018;37:40.
Ying Y, Ma X, Fang J, Chen S, Wang W, Li J, et al. EGR2-mediated regulation of m6A reader IGF2BP proteins drive RCC tumorigenesis and metastasis via enhancing S1PR3 mRNA stabilization. Cell Death Dis. 2021;12:750.
Liu S, Li G, Li Q, Zhang Q, Zhuo L, Chen X, et al. The roles and mechanisms of YTH domain-containing proteins in cancer development and progression. Am J Cancer Res. 2020;10:1068.
Li Y, Sheng H, Ma F, Wu Q, Huang J, Chen Q, et al. RNA m6A reader YTHDF2 facilitates lung adenocarcinoma cell proliferation and metastasis by targeting the AXIN1/Wnt/β-catenin signaling. Cell Death Dis. 2021;12:479.
Liu T, Wei Q, Jin J, Luo Q, Liu Y, Yang Y, et al. The m6A reader YTHDF1 promotes ovarian cancer progression via augmenting EIF3C translation. Nucleic Acids Res. 2020;48:3816–31.
Chang G, Shi L, Ye Y, Shi H, Zeng L, Tiwary S, et al. YTHDF3 induces the translation of m6a-enriched gene transcripts to promote breast cancer brain metastasis. Cancer Cell. 2020;38:857–71.e857.
Tan B, Zhou K, Liu W, Prince E, Qing Y, Li Y, et al. RNA N6-methyladenosine reader YTHDC1 is essential for TGF-beta-mediated metastasis of triple negative breast cancer. Theranostics. 2022;12:5727.
Yuan W, Chen S, Li B, Han X, Meng B, Zou Y, et al. The N6-methyladenosine reader protein YTHDC2 promotes gastric cancer progression via enhancing YAP mRNA translation. Transl Oncol. 2022;16:101308.
Hagen F, Gundert L, Strick A, Klümper N, Schmidt D, Kristiansen G, et al. N6‐Methyladenosine (m6A) readers are dysregulated in renal cell carcinoma. Mol Carcinog. 2021;60:354–62.
Yang L, Chen Y, Liu N, Shi Q, Han X, Gan W, et al. Low expression of TRAF3IP2-AS1 promotes progression of NONO-TFE3 translocation renal cell carcinoma by stimulating N6-methyladenosine of PARP1 mRNA and downregulating PTEN. J Hematol Oncol. 2021;14:46.
Liu HB, Muhammad T, Guo Y, Li MJ, Sha QQ, Zhang CX, et al. RNA‐binding protein IGF2BP2/IMP2 is a critical maternal activator in early zygotic genome activation. Adv Sci. 2019;6:1900295.
Wang J, Chen L, Qiang P. The role of IGF2BP2, an m6A reader gene, in human metabolic diseases and cancers. Cancer Cell Int. 2021;21:99.
Huang H, Weng H, Sun W, Qin X, Shi H, Wu H, et al. Recognition of RNA N6-methyladenosine by IGF2BP proteins enhances mRNA stability and translation. Nat Cell Biol. 2018;20:285–95.
Hammer NA, Byskov AG, Rajpert-De Meyts E, Grøndahl ML, Bredkjær HE, Wewer UM, et al. Expression of IGF-II mRNA-binding proteins (IMPs) in gonads and testicular cancer. Reproduction. 2005;130:203–12.
Sun C-Y, Cao D, Du B-B, Chen C-W, Liu D. The role of Insulin-like growth factor 2 mRNA-binding proteins (IGF2BPs) as m6A readers in cancer. Int J Biol Sci. 2022;18:2744.
Zhu P, He F, Hou Y, Tu G, Li Q, Jin T, et al. A novel hypoxic long noncoding RNA KB-1980E6. 3 maintains breast cancer stem cell stemness via interacting with IGF2BP1 to facilitate c-Myc mRNA stability. Oncogene. 2021;40:1609–27.
Liu X, He H, Zhang F, Hu X, Bi F, Li K, et al. m6A methylated EphA2 and VEGFA through IGF2BP2/3 regulation promotes vasculogenic mimicry in colorectal cancer via PI3K/AKT and ERK1/2 signaling. Cell Death Dis. 2022;13:483.
Maniotis AJ, Folberg R, Hess A, Seftor EA, Gardner LMG, Pe’er J, et al. Vascular channel formation by human melanoma cells in vivo and in vitro: vasculogenic mimicry. Am J Pathol. 1999;155:739–52.
Hoffmann NE, Sheinin Y, Lohse CM, Parker AS, Leibovich BC, Jiang Z, et al. External validation of IMP3 expression as an independent prognostic marker for metastatic progression and death for patients with clear cell renal cell carcinoma. Cancer. 2008;112:1471–9.
Jiang Z, Chu PG, Woda BA, Rock KL, Liu Q, Hsieh C-C, et al. Analysis of RNA-binding protein IMP3 to predict metastasis and prognosis of renal-cell carcinoma: a retrospective study. Lancet Oncol. 2006;7:556–64.
Jiang Z, Lohse CM, Chu PG, Wu CL, Woda BA, Rock KL, et al. Oncofetal protein IMP3: a novel molecular marker that predicts metastasis of papillary and chromophobe renal cell carcinomas. Cancer. 2008;112:2676–82.
Park JY, Choe M, Kang Y, Lee SS. IMP3, a promising prognostic marker in clear cell renal cell carcinoma. J Pathol Transl Med. 2014;48:108–16.
Xie C, Li Y, Li Q, Chen Y, Yao J, Yin G, et al. Increased insulin mRNA binding protein-3 expression correlates with vascular enhancement of renal cell carcinoma by intravenous contrast-CT and is associated with bone metastasis. J Bone Oncol. 2015;4:69–76.
Jiang Z, Chu PG, Woda BA, Liu Q, Balaji KC, Rock KL, et al. Combination of quantitative IMP3 and tumor stage: a new system to predict metastasis for patients with localized renal cell carcinomas. Clin Cancer Res. 2008;14:5579–84.
Tschirdewahn S, Panic A, Püllen L, Harke NN, Hadaschik B, Riesz P, et al. Circulating and tissue IMP3 levels are correlated with poor survival in renal cell carcinoma. Int J Cancer. 2019;145:531–9.
Yuan B, Zhou J. N6-methyladenosine (m6A) reader IGF2BP1 facilitates clear-cell renal cell carcinoma aerobic glycolysis. PeerJ (San Francisco, CA). 2023;11:e14591.
Pei X, Li M, Zhan J, Yu Y, Wei X, Guan L, et al. Enhanced IMP3 expression activates NF-кB pathway and promotes renal cell carcinoma progression. PLoS ONE. 2015;10:e0124338.
Chen Y, Lu Z, Qi C, Yu C, Li Y, Huan W, et al. N6-methyladenosine-modified TRAF1 promotes sunitinib resistance by regulating apoptosis and angiogenesis in a METTL14-dependent manner in renal cell carcinoma. Mol Cancer. 2022;21:1–111.
Li H, Liu J, Shen S, Dai D, Cheng S, Dong X, et al. Pan‐cancer analysis of alternative splicing regulator heterogeneous nuclear ribonucleoproteins (hnRNPs) family and their prognostic potential. J Cell Mol Med. 2020;24:11111–9.
Han N, Li W, Zhang M. The function of the RNA-binding protein hnRNP in cancer metastasis. J Cancer Res Ther. 2013;9:129–34.
Wu J, Wei Y, Miao C, Wang S, Wang X, Wang Z. Essential m6A methylation regulator HNRNPC serves as a targetable biomarker for papillary renal cell carcinoma. J Oncol. 2022;2022:1–29.
Perry GS, Das M, Woon ECY. Inhibition of AlkB nucleic acid demethylases: promising new epigenetic targets. J Med Chem. 2021;64:16974–7003.
Gutierrez R, O’Connor TR. DNA direct reversal repair and alkylating agent drug resistance. Cancer Drug Resist. 2021;4:414–23.
Guo X, Li K, Jiang W, Hu Y, Xiao W, Huang Y, et al. RNA demethylase ALKBH5 prevents pancreatic cancer progression by posttranscriptional activation of PER1 in an m6A-YTHDF2-dependent manner. Mol Cancer. 2020;19:91.
Yang P, Wang Q, Liu A, Zhu J, Feng J. ALKBH5 holds prognostic values and inhibits the metastasis of colon cancer. Pathol Oncol Res. 2019;26:1615–23.
Zhang J, Guo S, Piao H-Y, Wang Y, Wu Y, Meng X-Y, et al. ALKBH5 promotes invasion and metastasis of gastric cancer by decreasing methylation of the lncRNA NEAT1. J Physiol Biochem. 2019;75:379–89.
Zhu H, Gan X, Jiang X, Diao S, Wu H, Hu J. ALKBH5 inhibited autophagy of epithelial ovarian cancer through miR-7 and BCL-2. J Exp Clin Cancer Res. 2019;38:163.
Zhu Z, Qian Q, Zhao X, Ma L, Chen P. N6-methyladenosine ALKBH5 promotes non-small cell lung cancer progress by regulating TIMP3 stability. Gene. 2020;731:144348.
Guimarães-Teixeira C, Barros-Silva D, Lobo J, Soares-Fernandes D, Constâncio V, Leite-Silva P, et al. Deregulation of N6-methyladenosine RNA modification and its erasers FTO/ALKBH5 among the main renal cell tumor subtypes. J Pers Med. 2021;11:996.
Strick A, Hagen F, Gundert L, Klümper N, Tolkach Y, Schmidt D, et al. The N6‐methyladenosine (m6A) erasers alkylation repair homologue 5 (ALKBH5) and fat mass and obesity‐associated protein (FTO) are prognostic biomarkers in patients with clear cell renal carcinoma. BJU Int. 2020;125:617–24.
Zhang X, Wang F, Wang Z, Yang X, Yu H, Si S, et al. ALKBH5 promotes the proliferation of renal cell carcinoma by regulating AURKB expression in an m6A-dependent manner. Ann Transl Med. 2020;8:646.
Hu W, Klümper N, Schmidt D, Ritter M, Ellinger J, Hauser S. Depletion of the m6A demethylases FTO and ALKBH5 impairs growth and metastatic capacity through EMT phenotype change in clear cell renal cell carcinoma. Am J Transl Res. 2023;15:1744.
Hotta K, Sho M, Fujimoto K, Shimada K, Yamato I, Anai S, et al. Clinical significance and therapeutic potential of prostate cancer antigen-1/ALKBH3 in human renal cell carcinoma. Oncol Rep. 2015;34:648–54.
Li L, Zhu C, Xu Q, Xu S, Ye J, Xu D, et al. ALKBH1 contributes to renal cell carcinoma progression by reducing N6-methyladenine of GPR137. Eur J Clin Invest. 2023;53:e13986.
Jeschke J, Collignon E, Al Wardi C, Krayem M, Bizet M, Jia Y, et al. Downregulation of the FTO m6A RNA demethylase promotes EMT-mediated progression of epithelial tumors and sensitivity to Wnt inhibitors. Nat Cancer. 2021;2:611–28.
Zou L, Chen W, Zhou X, Yang T, Luo J, Long Z, et al. N6-methyladenosine demethylase FTO suppressed prostate cancer progression by maintaining CLIC4 mRNA stability. Cell Death Discov. 2022;8:184.
Keelan S, Ola M, Charmsaz S, Cocchiglia S, Ottaviani D, Hickey S, et al. Dynamic epi-transcriptomic landscape mapping with disease progression in estrogen receptor-positive breast cancer. Cancer Commun. 2023;43:615.
Zhou Y, Wang Q, Deng H, Xu B, Zhou Y, Liu J, et al. N6-methyladenosine demethylase FTO promotes growth and metastasis of gastric cancer via m6A modification of caveolin-1 and metabolic regulation of mitochondrial dynamics. Cell Death Dis. 2022;13:72.
Tan Z, Shi S, Xu J, Liu X, Lei Y, Zhang B, et al. RNA N6-methyladenosine demethylase FTO promotes pancreatic cancer progression by inducing the autocrine activity of PDGFC in an m6A-YTHDF2-dependent manner. Oncogene. 2022;41:2860–72.
Zhuang C, Zhuang C, Luo X, Huang X, Yao L, Li J, et al. N6‐methyladenosine demethylase FTO suppresses clear cell renal cell carcinoma through a novel FTO‐PGC‐1α signalling axis. J Cell Mol Med. 2019;23:2163–73.
Vultaggio-Poma V, Sarti AC, Di Virgilio F. Extracellular ATP: a feasible target for cancer therapy. Cells (Basel, Switz). 2020;9:2496.
Jiang JX, Riquelme MA, Zhou JZ. ATP, a double-edged sword in cancer. Oncoscience. 2015;2:673.
Xiao Y, Thakkar KN, Zhao H, Broughton J, Li Y, Seoane JA, et al. The m6A RNA demethylase FTO is a HIF-independent synthetic lethal partner with the VHL tumor suppressor. Proc Natl Acad Sci. 2020;117:21441–9.
Zhang C, Chen L, Lou W, Su J, Huang J, Liu A, et al. Aberrant activation of m6A demethylase FTO renders HIF2αlow/− clear cell renal cell carcinoma sensitive to BRD9 inhibitors. Sci Transl Med. 2021;13:eabf6045.
Zhang Q, Liu F, Chen W, Miao H, Liang H, Liao Z, et al. The role of RNA m5C modification in cancer metastasis. Int J Biol Sci. 2021;17:3369–80.
Xue C, Zhao Y, Li L. Advances in RNA cytosine-5 methylation: detection, regulatory mechanisms, biological functions and links to cancer. Biomark Res. 2020;8:1–43.
Yi J, Gao R, Chen Y, Yang Z, Han P, Zhang H, et al. Overexpression of NSUN2 by DNA hypomethylation is associated with metastatic progression in human breast cancer. Oncotarget. 2017;8:20751–65.
Xue M, Shi Q, Zheng L, Li Q, Yang L, Zhang Y. Gene signatures of m5C regulators may predict prognoses of patients with head and neck squamous cell carcinoma. Am J Transl Res. 2020;12:6841–52.
Wu J, Hou C, Wang Y, Wang Z, Li P, Wang Z. Comprehensive analysis of m5C RNA methylation regulator genes in clear cell renal cell carcinoma. Int J Genomics. 2021;2021:1–24.
Li H, Jiang H, Huang Z, Chen Z, Chen N. Prognostic value of an m5C RNA methylation regulator-related signature for clear cell renal cell carcinoma. Cancer Manag Res. 2021;13:6673–87.
Wang J, Ju HJ, Zhang F, Tian H, Wang WG, Ma YL, et al. A novel NSUN5/ENO3 pathway promotes the Warburg effect and cell growth in clear cell renal cell carcinoma by 5-methylcytosine-stabilized ENO3 mRNA. Am J Transl Res. 2023;15:878.
Wettersten HI, Aboud OA, Lara JPN, Weiss RH. Metabolic reprogramming in clear cell renal cell carcinoma. Nat Rev Nephrol. 2017;13:410–9.
Yang L, Yin H, Chen Y, Pan C, Hang H, Lu Y, et al. Low expression of PEBP1P2 promotes metastasis of clear cell renal cell carcinoma by post-transcriptional regulation of PEBP1 and KLF13 mRNA. Exp Hematol Oncol. 2022;11:1–87.
Tusup M, Kundig T, Pascolo S. Epitranscriptomics of cancer. World J Clin Oncol. 2018;9:42.
Charette M, Gray MW. Pseudouridine in RNA: what, where, how, and why. IUBMB life. 2000;49:341–52.
Xue C, Chu Q, Zheng Q, Jiang S, Bao Z, Su Y, et al. Role of main RNA modifications in cancer: N6-methyladenosine, 5-methylcytosine, and pseudouridine. Signal Transduct Target Ther. 2022;7:142.
Elsharawy KA, Mohammed OJ, Aleskandarany MA, Hyder A, El-Gammal HL, Abou-Dobara MI, et al. The nucleolar-related protein Dyskerin pseudouridine synthase 1 (DKC1) predicts poor prognosis in breast cancer. Br J Cancer. 2020;123:1543–52.
Li H, Chen L, Han Y, Zhang F, Wang Y, Han Y, et al. The identification of RNA modification Gene PUS7 as a potential biomarker of ovarian cancer. Biol (Basel, Switz). 2021;10:1130.
Zhang M, Pan Y, Jiang R, Hou P, Shan H, Chen F, et al. DKC1 serves as a potential prognostic biomarker for human clear cell renal cell carcinoma and promotes its proliferation, migration and invasion via the NF‑κB pathway. Oncol Rep. 2018;40:968–78.
Raitskin O, Cho DSC, Sperling J, Nishikura K, Sperling R. RNA editing activity is associated with splicing factors in lnRNP particles: the nuclear pre-mRNA processing machinery. Proc Natl Acad Sci. 2001;98:6571–6.
Samuel CE. Adenosine deaminases acting on RNA (ADARs) are both antiviral and proviral. Virology. 2010;411:180–93.
Sagredo EA, Sagredo AI, Blanco A, Rojas De Santiago P, Rivas S, Assar R, et al. ADAR1 Transcriptome editing promotes breast cancer progression through the regulation of cell cycle and DNA damage response. Biochim Biophys Acta. 2020;1867:118716.
Li W, Yang F-Q, Sun C-M, Huang J-H, Zhang H-M, Li X, et al. circPRRC2A promotes angiogenesis and metastasis through epithelial-mesenchymal transition and upregulates TRPM3 in renal cell carcinoma. Theranostics. 2020;10:4395–409.
Cervantes-Gracia K, Gramalla-Schmitz A, Weischedel J, Chahwan R. APOBECs orchestrate genomic and epigenomic editing across health and disease. Trends Genet. 2021;37:1028–43.
Lerner T, Papavasiliou FN, Pecori R. RNA editors, cofactors, and mRNA targets: an overview of the C-to-U RNA editing machinery and its implication in human disease. Genes. 2018;10:13.
Pecori R, Di Giorgio S, Paulo Lorenzo J, Nina Papavasiliou F. Functions and consequences of AID/APOBEC-mediated DNA and RNA deamination. Nat Rev Genet. 2022;23:505–18.
Navaratnam N, Sarwar R. An overview of cytidine deaminases. Int J Hematol. 2006;83:195–200.
Imahashi M, Nakashima M, Iwatani Y. Antiviral mechanism and biochemical basis of the human APOBEC3 Family. Front Microbiol. 2012;3:250.
Luo Y, Yan B, Liu L, Yin L, Ji H, An X, et al. Sulforaphane inhibits the expression of long noncoding RNA H19 and its target APOBEC3G and thereby pancreatic cancer progression. Cancers. 2021;13:827.
Vile RG, Melcher A, Pandha H, Harrington KJ, Pulido JS. APOBEC and cancer viroimmunotherapy: thinking the unthinkable. Clin Cancer Res. 2021;27:3280–90.
Peng T, Liu B, Lin S, Cao C, Wu P, Zhi W, et al. APOBEC3G expression correlates with unfavorable prognosis and immune infiltration in kidney renal clear cell carcinoma. Heliyon. 2022;8:e12191.
Li Y, Xia L, Tan K, Ye X, Zuo Z, Li M, et al. N 6-methyladenosine co-transcriptionally directs the demethylation of histone H3K9me2. Nat Genet. 2020;52:870–7.
Deng S, Zhang J, Su J, Zuo Z, Zeng L, Liu K, et al. RNA m6A regulates transcription via DNA demethylation and chromatin accessibility. Nat Genet. 2022;54:1427–37.
Xu W, Li J, He C, Wen J, Ma H, Rong B, et al. METTL3 regulates heterochromatin in mouse embryonic stem cells. Nat (Lond). 2021;591:317–21.
Li F, Chen S, Yu J, Gao Z, Sun Z, Yi Y, et al. Interplay of m6A and histone modifications contributes to temozolomide resistance in glioblastoma. Clin Transl Med. 2021;11:e553.
Gu Y, Niu S, Wang Y, Duan L, Pan Y, Tong Z, et al. DMDRMR-mediated regulation of m6A-modified CDK4 by m6A reader IGF2BP3 drives ccRCC progression. Cancer Res. 2021;81:923–34.
He L, Chen S, Ying Y, Xie H, Li J, Ma X, et al. MicroRNA‐501‐3p inhibits the proliferation of kidney cancer cells by targeting WTAP. Cancer Med. 2021;10:7222–32.
Huang X, Huang M, Kong L, Li Y. miR-372 suppresses tumour proliferation and invasion by targeting IGF2BP1 in renal cell carcinoma. Cell Prolif. 2015;48:593–9.
Yang W, Xie L, Wang P, Zhuang C. MiR-155 regulates m6A level and cell progression by targeting FTO in clear cell renal cell carcinoma. Cell Signal. 2022;91:110217.
Xie X, Lin J, Fan X, Zhong Y, Chen Y, Liu K, et al. LncRNA CDKN2B-AS1 stabilized by IGF2BP3 drives the malignancy of renal clear cell carcinoma through epigenetically activating NUF2 transcription. Cell Death Dis. 2021;12:201.
Shen D, Ding L, Lu Z, Wang R, Yu C, Wang H, et al. METTL14-mediated Lnc-LSG1 m6A modification inhibits clear cell renal cell carcinoma metastasis via regulating ESRP2 ubiquitination. Mol Ther Nucleic Acids. 2022;27:547–61.
Liu T, Wang H, Fu Z, Wang Z, Wang J, Gan X, et al. Methyltransferase‐like 14 suppresses growth and metastasis of renal cell carcinoma by decreasing long noncoding RNA NEAT1. Cancer Sci. 2022;113:446–58.
Grosjean H. RNA modification: the Golden Period 1995–2015. RNA. 2015;21:625–6.
Acknowledgements
Figures of this article were created using Inkscape 1.2.1 (9c6d41e410, 2022-07-14). The figures were created in part using Servier Medical Art, made available by Servier, and are licensed under a Creative Commons Attribution 3.0 Unported License.
Funding
RH is supported by ASPIRE, the technology program management pillar of Abu Dhabi’s Advanced Technology Research Council (ATRC), via the ASPIRE Precision Medicine Research Institute Abu Dhabi (ASPIREPMRIAD) award grant number VRI-20-10. I.T. and R.H. are supported by the University of Sharjah grant (code number: 200-109021-02).
Author information
Authors and Affiliations
Contributions
MAA, KB, IMT, and RH conceptualized the manuscript. MAA collected the references and wrote the first draft of the manuscript. KB, IMT, and RH reviewed and edited the manuscript. IMT and RH acquired funding. MAA designed the figures and the tables. All authors approved the final version of the article.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Consent for publication
Not applicable.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Edited by Professor Boris Zhivotovsky
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Alhammadi, M.A., Bajbouj, K., Talaat, I.M. et al. The role of RNA-modifying proteins in renal cell carcinoma. Cell Death Dis 15, 227 (2024). https://doi.org/10.1038/s41419-024-06479-y
Received:
Revised:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41419-024-06479-y