Abstract
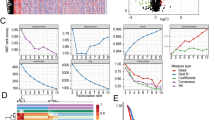
Increasing evidence imputes cancer progression and resistance to therapy to intra-tumor molecular heterogeneity set off by cancer cell plasticity. Re-activation of developmental programs strictly linked to epithelial-to-mesenchymal transition and gaining of stem cells properties are crucial in this setting. Many biological processes involved in cancer onset and progression show rhythmic fluctuations driven by the circadian clock circuitry. Novel cancer patient stratification tools taking into account the temporal dimension of these biological processes are definitely needed. Lung cancer and colorectal cancer (CRC) are the leading causes of cancer death worldwide. Here, by developing an innovative computational approach we named Phase-Finder, we show that the molecular heterogeneity characterizing the two deadliest cancers, CRC and lung adenocarcinoma (LUAD), rather than a merely stochastic event is the readout of specific cancer molecular states which correlate with time-qualified patterns of gene expression. We performed time-course transcriptome analysis of CRC and LUAD cell lines and upon computing circadian genes expression-based correlation matrices we derived pseudo-time points to infer time-qualified patterns in the transcriptomic analysis of real-world data (RWD) from large cohorts of CRC and LUAD patients. Our temporal classification of CRC and LUAD cohorts was able to effectively render time-specific patterns in cancer phenotype switching determining dynamical distribution of molecular subtypes impacting patient prognosis.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout


Similar content being viewed by others
References
Sulli G, Lam MTY, Panda S. Interplay between circadian clock and cancer: new frontiers for cancer treatment. Trends Cancer. 2019;5:475–94.
Shafi AA, Knudsen KE. Cancer and the circadian clock. Cancer Res. 2019;79:3806–14.
Papagiannakopoulos T, Bauer MR, Davidson SM, Heimann M, Subbaraj L, Bhutkar A, et al. Circadian rhythm disruption promotes lung tumorigenesis. Cell Metab. 2016;24:324–31.
Chun SK, Fortin BM, Fellows RC, Habowski AN, Verlande A, Song WA, et al. Disruption of the circadian clock drives Apc loss of heterozygosity to accelerate colorectal cancer. Sci Adv. 2022;8:eabo2389.
Gerlinger M, Rowan AJ, Horswell S, Larkin J, Endesfelder D, Gronroos E, et al. Intratumor heterogeneity and branched evolution revealed by multiregion sequencing. N. Engl J Med. 2012;366:883–92.
Maynard A, McCoach CE, Rotow JK, Harris L, Haderk F, Kerr DL, et al. Therapy-induced evolution of human lung cancer revealed by single-cell RNA sequencing. Cell. 2020;182:1232–1251.e22.
de Bruin EC, McGranahan N, Mitter R, Salm M, Wedge DC, Yates L, et al. Spatial and temporal diversity in genomic instability processes defines lung cancer evolution. Science. 2014;346:251–6.
Cancer Genome Atlas Research N. Comprehensive molecular profiling of lung adenocarcinoma. Nature. 2014;511:543–50.
Guinney J, Dienstmann R, Wang X, de Reyniès A, Schlicker A, Soneson C, et al. The consensus molecular subtypes of colorectal cancer. Nat Med. 2015;21:1350–6.
Marisa L, Blum Y, Taieb J, Ayadi M, Pilati C, Le Malicot K, et al. Intratumor CMS heterogeneity impacts patient prognosis in localized colon cancer. Clin Cancer Res. 2021;27:4768–80.
Siravegna G, Mussolin B, Buscarino M, Corti G, Cassingena A, Crisafulli G, et al. Clonal evolution and resistance to EGFR blockade in the blood of colorectal cancer patients. Nat Med. 2015;21:795–801.
Dawson S-J, Tsui DWY, Murtaza M, Biggs H, Rueda OM, Chin S-F, et al. Analysis of circulating tumor DNA to monitor metastatic breast cancer. N Engl J Med. 2013;368:1199–209.
Marjanovic ND, Hofree M, Chan JE, Canner D, Wu K, Trakala M, et al. Emergence of a high-plasticity cell state during lung cancer evolution. Cancer Cell. 2020;38:229–246.e13.
Zhang Y, Donaher JL, Das S, Li X, Reinhardt F, Krall JA, et al. Genome-wide CRISPR screen identifies PRC2 and KMT2D-COMPASS as regulators of distinct EMT trajectories that contribute differentially to metastasis. Nat Cell Biol. 2022;24:554–64.
Colangelo T, Carbone A, Mazzarelli F, Cuttano R, Dama E, Nittoli T, et al. Loss of circadian gene Timeless induces EMT and tumor progression in colorectal cancer via Zeb1-dependent mechanism. Cell Death Differ. 2022. https://doi.org/10.1038/s41418-022-00935-y.
Melocchi V, Dama E, Mazzarelli F, Cuttano R, Colangelo T, Di Candia L, et al. Aggressive early-stage lung adenocarcinoma is characterized by epithelial cell plasticity with acquirement of stem-like traits and immune evasion phenotype. Oncogene. 2021;40:4980–91.
Brady JJ, Chuang C-H, Greenside PG, Rogers ZN, Murray CW, Caswell DR, et al. An Arntl2-driven secretome enables lung adenocarcinoma metastatic self-sufficiency. Cancer Cell. 2016;29:697–710.
El-Athman R, Fuhr L, Relógio A. A systems-level analysis reveals circadian regulation of splicing in colorectal cancer. EBioMedicine. 2018;33:68–81.
Nakata A, Yoshida R, Yamaguchi R, Yamauchi M, Tamada Y, Fujita A, et al. Elevated β-catenin pathway as a novel target for patients with resistance to EGF receptor targeting drugs. Sci Rep. 2015;5:13076.
Cancer Genome Atlas N. Comprehensive molecular characterization of human colon and rectal cancer. Nature. 2012;487:330–7.
Gao J, Aksoy BA, Dogrusoz U, Dresdner G, Gross B, Sumer SO, et al. Integrative analysis of complex cancer genomics and clinical profiles using the cBioPortal. Sci Signal. 2013;6:pl1.
Wu G, Anafi RC, Hughes ME, Kornacker K, Hogenesch JB. MetaCycle: an integrated R package to evaluate periodicity in large scale data. Bioinformatics. 2016;32:3351–3.
Corsello SM, Nagari RT, Spangler RD, Rossen J, Kocak M, Bryan JG, et al. Discovering the anticancer potential of non-oncology drugs by systematic viability profiling. Nat Cancer. 2020;1:235–48.
Subramanian A, Tamayo P, Mootha VK, Mukherjee S, Ebert BL, Gillette MA, et al. Gene set enrichment analysis: a knowledge-based approach for interpreting genome-wide expression profiles. Proc Natl Acad Sci USA. 2005;102:15545–50.
Fuhr L, Abreu M, Carbone A, El-Athman R, Bianchi F, Laukkanen MO, et al. The interplay between colon cancer cells and tumour-associated stromal cells impacts the biological clock and enhances malignant phenotypes. Cancers. 2019;11. https://doi.org/10.3390/cancers11070988.
Zhang R, Podtelezhnikov AA, Hogenesch JB, Anafi RC. Discovering biology in periodic data through phase set enrichment analysis (PSEA). J Biol Rhythms. 2016;31:244–57.
Zhan T, Rindtorff N, Boutros M. Wnt signaling in cancer. Oncogene. 2017;36:1461–73.
Roh W, Geffen Y, Cha H, Miller M, Anand S, Kim J, et al. High-resolution profiling of lung adenocarcinoma identifies expression subtypes with specific biomarkers and clinically relevant vulnerabilities. Cancer Res. 2022;82:3917–31.
Fuhr L, El-Athman R, Scrima R, Cela O, Carbone A, Knoop H, et al. The circadian clock regulates metabolic phenotype rewiring via HKDC1 and modulates tumor progression and drug response in colorectal cancer. EBioMedicine. 2018;33:105–21.
Parascandolo A, Bonavita R, Astaburuaga R, Sciuto A, Reggio S, Barra E, et al. Effect of naive and cancer-educated fibroblasts on colon cancer cell circadian growth rhythm. Cell Death Dis. 2020;11:289.
Stevens RG, Hansen J, Costa G, Haus E, Kauppinen T, Aronson KJ, et al. Considerations of circadian impact for defining ‘shift work’ in cancer studies: IARC Working Group Report. Occup Environ Med. 2011;68:154–62.
Tivey A, Church M, Rothwell D, Dive C, Cook N. Circulating tumour DNA — looking beyond the blood. Nat Rev Clin Oncol. 2022;19:600–12.
Acknowledgements
This work was supported in part by the Italian Minister of Health [Ricerca Corrente program 2022-2024 to FB and GM; Ricerca Finalizzata GR-2016-02363975, RF-2021-12372433 and CLEARLY to FB], by the Associazione Italiana Ricerca sul Cancro (AIRC) [IG-22827 to FB]. The study funders had no role in the design of the study, the collection, analysis, and interpretation of the data, the writing of the manuscript, and the decision to submit the manuscript for publication. All authors gave their consent to publication.
Author information
Authors and Affiliations
Contributions
Conception and design: GM, FB. Statistical and bioinformatics analyses: VM, RC, EM, FB. Interpretation of data: VM, RC, GM, FB. Writing, review of the manuscript: GM, FB. All authors approved the final version of the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Melocchi, V., Cuttano, R., Murgo, E. et al. The circadian clock circuitry deconvolutes colorectal cancer and lung adenocarcinoma heterogeneity in a dynamic time-related framework. Cancer Gene Ther 30, 1323–1329 (2023). https://doi.org/10.1038/s41417-023-00646-7
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41417-023-00646-7