Abstract
The aim of this study was to explore the association between genetically predicted circulating levels of immunity and inflammation, and the risk of Alzheimer’s disease (AD) and hippocampal volume, by conducting a two-sample Mendelian Randomization Study. We identified 12 markers of immune cells and derived ratios (platelet count, eosinophil count, neutrophil count, basophil count, monocyte count, lymphocyte count, platelet-to-lymphocyte ratio, monocyte-to-lymphocyte ratio, CD4 count, CD8 count, CD4-to-CD8 ratio, and CD56) and 5 signaling molecules (IL-6, fibrinogen, CRP, and Lp-PLA2 activity and mass) as primary exposures of interest. Other genetically available immune biomarkers with a weaker a priori link to AD were considered secondary exposures. Associations with AD were evaluated in The International Genomics of Alzheimer’s Project (IGAP) GWAS dataset (21,982 cases; 41,944 controls of European ancestry). For hippocampal volume, we extracted data from a GWAS meta-analysis on 33,536 participants of European ancestry. None of the primary or secondary exposures showed statistically significant associations with AD or with hippocampal volume following P-value correction for multiple comparisons using false discovery rate < 5% (Q-value < 0.05). CD4 count showed the strongest suggestive association with AD (odds ratio 1.32, P < 0.01, Q > 0.05). There was evidence for heterogeneity in the MR inverse variance-weighted meta-analyses as measured by Cochran Q, and weighted median and weighted mode for multiple exposures. Further cluster analyses did not reveal clusters of variants that could influence the risk factor in distinct ways. This study suggests that genetically predicted circulating biomarkers of immunity and inflammation are not associated with AD risk or hippocampal volume. Future studies should assess competing risk, explore in more depth the role of adaptive immunity in AD, in particular T cells and the CD4 subtype, and confirm these findings in other ethnicities.
Similar content being viewed by others
Introduction
The immune system is increasingly recognized to be involved in the pathogenesis of Alzheimer’s disease (AD)1,2. Recent genome-wide association studies (GWASs) have established AD risk loci within or near genes that are expressed in microglia3. This led to the concept of the innate immune system being involved in the early steps of the disease and, thus, much effort has been dedicated in studying innate immunity in relation to AD. A recent meta-analysis of observational studies revealed that the immune-related signaling molecules C-reactive protein (CRP), interleukin (IL)-6, α1-antichymotrypsin, lipoprotein-associated phospholipase A2 (Lp-PLA2) activity, and fibrinogen were each associated with risk of all-cause dementia4. Less is known about the contribution of the adaptive immune system in relation to AD, but a recent observational study discovered clonally expanded CD8+ T-effector memory cells in the cerebrospinal fluid of AD patients, revealing an adaptive immune response in AD5. Moreover, we previously found that higher levels of innate immune cells lead to a higher dementia risk, whereas higher levels of adaptive immune cells are protective for developing dementia6. Given the observational design of the majority of available studies and the difficulty of studying the effect of the immune system on AD in a trial, it is uncertain whether the observed associations are causal, i.e., independent of other risk factors, and not biased by reverse causation7.
Mendelian randomization (MR) exploits genetic variants influencing the exposure of interest as unbiased proxies for the exposure, i.e., instruments, to infer causality8. To our knowledge, there are only few MR studies performed where the association between circulating biomarkers of immunity and inflammation, and dementia, was studied9,10,11,12,13. Moreover, a large GWAS meta-analysis on hippocampal volume14 allows exploration of these biomarkers with hippocampal volume as imaging endophenotype of AD, which, to date, has not been performed. Furthermore, as GWAS studies are increasing in size, the number of instruments that can be used to estimate the causal effect of a risk factor on an outcome also increases. This could lead to more heterogeneity among the causal estimates obtained from multiple genetic variants, pointing to a possible violation of the necessary instrumental variable assumptions, but also to a scenario in which causal estimates based on each variant in turn differ more strongly than expected by chance alone. These variants could then be divided into distinct clusters, such that all variants in the cluster have similar causal estimates. There are now novel techniques available, which allow for cluster analyses of variants, which can capture distinct causal mechanisms by which a risk factor influences an outcome with different magnitudes of causal effect15.
Here, by leveraging data from large-scale genomic studies on circulating biomarkers of immunity and inflammation, and the large AD dataset from The International Genomics of Alzheimer’s Project (IGAP) GWAS3 and hippocampal volume GWAS14, we implemented a two-sample MR study to (1) explore the associations between genetic predisposition to higher or lower circulating levels of immune cells12,16 and signaling molecules13,17,18,19 with risk of AD; (2) explore the associations between genetic predisposition to these biomarkers with hippocampal volume; (3) explore the associations between genetic predisposition to circulating biomarkers of immunity and inflammation with limited a priori evidence with AD and hippocampal volume; and (4) explore whether different genetic variants influence the exposures, and thus AD and hippocampal volume in distinct ways by performing cluster analyses.
Methods
Study design, data sources, and genetic instrument selection
Table 1 summarizes the data sources used in the current MR study. The genetic instruments were taken from publicly available summary statistics. For each of the circulating immunity traits, we selected single-nucleotide polymorphisms (SNPs) associated with their circulating levels at a genome-wide threshold of significance (P < 5 × 10−8). After extracting the summary statistics for significant SNPs, we pruned all SNPs in linkage disequilibrium (LD) (r2 < 0.01 in the European 1000 Genomes Project reference panel), retaining SNPs with the lowest P-value as an independent instrument. For some exposures (IL-6, CRP, Lp-PLA2, 23 cytokines, and IL-1), we used previously selected instruments (Table 1). We identified independent SNPs significantly associated with circulating biomarker levels of immunity and inflammation, and merged these with the outcome datasets; the SNPs that were also available in the outcome datasets were used as final instruments for analysis. As all analyses are based on publicly available summary statistics and not individual-level data, no ethical approval from an institutional review board was required.
Primary exposures (biomarkers of immunity and inflammation with strong a priori evidence)
To minimize weak instrument bias and maximize power, we carefully selected our primary exposures prior to data analysis based on the underlying GWAS size, population characteristics, and a priori evidence for the associations with AD (Table 1)20. We identified 12 immune cells and derived ratios (platelet count, eosinophil count, neutrophil count, basophil count, monocyte count, lymphocyte count, platelet-to-lymphocyte (PLR) ratio, monocyte-to-lymphocyte ratio, CD4 count, CD8 count, CD4 : CD8 ratio, and CD56) and 5 signaling molecules (IL-6, fibrinogen, CRP, and Lp-PLA2 activity and Lp-PLA2 mass) as primary exposures of interest.
Secondary exposures (biomarkers of immunity and inflammation with limited a priori evidence)
Other immune-related exposures, for which there are less validated biomarkers of immunity and inflammation or less valid instruments available, were selected as secondary exposures (Table 1). More specifically, a GWAS identified multiple common genetic variants that influence circulating levels of 41 cytokines and growth factors21, of which we used pre-selected instruments for 23 cytokines or growth factors that were not in LD and not associated with levels of >1 cytokine22. These instruments have a weaker a priori link to AD and were therefore selected as secondary exposures. Furthermore, we used genetic instruments for IL-1, intercellular adhesion molecule 1 (ICAM-1), and P-selectin as additional secondary exposures due to smaller powered underlying GWASs.
Outcomes
The primary outcome for this study was AD defined by clinical diagnosis of AD. In addition, we looked at hippocampal volume as an imaging AD endophenotype as hippocampal degeneration is one of the pathological hallmarks of AD. We extracted estimates for the associations of the selected instruments with AD from IGAP GWAS dataset (21,982 cases; 41,944 controls of European ancestry)3. For hippocampal volume, we extracted data from publicly available summary statistics of the Cohorts for Heart and Aging Research in Genomic Epidemiology–Enhancing Neuro Imaging Genetics through Meta Analysis GWAS meta-analysis on 33,536 participants of European ancestry14.
Our power calculations23 revealed that based on the sample size of IGAP, we had sufficient power for most biomarkers of immunity and inflammation to detect meaningful effect sizes. Specifically, we had >80% power to detect significant associations with AD for 8 out of 12 immune cells and for 3 out of 5 signaling molecules at an effect size (odds ratio [OR]) of 1.10 or 0.90 (Supplementary Table S1). All markers were analyzed, even when potentially underpowered, to guide future research.
Statistical analysis
We first extracted data and harmonized the effect alleles across GWASs. The MR association between each immune cell or signaling molecule and AD or hippocampal volume was then estimated using the Wald estimator and the delta method after pooling individual SNP MR estimates using inverse variance-weighted (IVW) meta-analysis24. Fixed-effect IVW was used in the absence of heterogeneity and random effects in the presence of heterogeneity (Cochran Q-derived P < 0.05). Statistical significance for the MR associations with AD and hippocampal volume were set at a P-value corrected for multiple comparisons using false discovery rate (FDR) < 5%. A P < 0.05, but above the FDR-corrected threshold, was considered as suggestive for an association. These analyses were repeated for the secondary exposures with AD and hippocampal volume, and we set a separate corrected P-value for multiple comparisons of secondary exposures using FDR < 5%.
Cochran’s Q-derived P < 0.05 from the IVW MR was used as indicator of possible horizontal pleiotropy25. For markers with >2 SNPs showing either significant or suggestive associations or significant heterogeneity in the primary IVW MR analysis, we conducted additional sensitivity analyses that vary in their underlying assumptions regarding the presence of pleiotropic genetic variants that may be associated with the outcome independently of the exposure. In particular, we used the weighted median method, which requires that at least half of the information for the MR analysis comes from valid instruments26. We also used the weighted mode approach, which requires that the largest number of similar (identical in infinite samples) individual-instrument causal effect estimates comes from valid instruments, even if the majority of instruments are invalid27. For consistency with other literature and to further relax the IVW assumptions, we used MR-Egger regression, which provides a consistent estimate of the causal effect, under a weaker assumption—the InSIDE (INstrument Strength Independent of Direct Effect) assumption28. In addition, we used the contamination mixture method, which is implemented by constructing a likelihood function based on the variant-specific causal estimates. If a genetic variant is a valid instrument, then its causal estimate will be normally distributed about the true value of the causal effect, but if a genetic variant is not a valid instrument then its causal estimate will be normally distributed about some other value29. We also tested for outlier SNPs using MR-Pleiotropy Residual Sum and Outlier30.
Finally, as it is possible that different genetic variants influence the risk factor in distinct ways, e.g., via distinct biological mechanisms, we further examined heterogeneity by performing cluster analyses using the MR-Clust package15. As recommended, we implemented this method conservatively, i.e., only assigning a variant to a cluster if the conditional probability of cluster assignment is ≥0.8 and only reporting a cluster if at least 4 variants satisfy this criterion. Variants that do not satisfy these criteria and that do not fit into a null cluster will be assigned to a “junk” cluster. Immune cells or signaling molecules that showed suggestive associations and for which more genome-wide significant SNPs were available were also explored for potential clustering of variants.
Statistical analyses were conducted in RStudio (R version 3.6.3).
Results
Primary exposures with AD
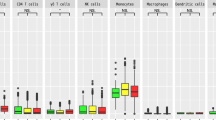
The primary results of the MR analyses for the genetic variants of immune cells and signaling molecules with AD are presented in Fig. 1. Following P-value correction for multiple comparisons using FDR < 5% (Q-value < 0.05), none of the immune cells or signaling molecules showed statistically significant associations with AD. CD4 count showed the strongest suggestive association with AD by an OR of 1.32, P = 0.005, Q = 0.170 (P < 0.01, Q > 0.05) with the next strongest suggestive association being between CRP and AD with P = 0.029 (P < 0.05, Q > 0.05).
Shown are the results derived from the primary inverse variance-weighted meta-analysis. None of the immune cells or signaling molecules survived the multiple testing threshold of false discovery rate < 5% (q < 0.05). CD, cluster of differentiation; CRP, C-reactive protein; IL, interleukin; Lp-PLA2, Lipoprotein-associated phospholipase A2; MLR, monocyte-to-lymphocyte ratio; PLR, platelet-to-lymphocyte ratio.
Primary exposures with hippocampal volume
The primary results of the MR analyses for the immune cells and signaling molecules with hippocampal volume are presented in Fig. 1 and Supplementary Table S1. None of the immune cells or signaling molecules showed statistically significant associations with hippocampal volume following P-value correction for multiple comparisons. Only PLR ratio showed a suggestive P = 0.037 (P < 0.05, Q > 0.05) association with hippocampal volume.
Secondary exposures with AD and hippocampal volume
The secondary results of the MR analyses are presented in Fig. 2 and Supplementary Table S2. Similarly, none of the biomarkers of immunity and inflammation showed statistically significant associations with AD or hippocampal volume following P-value correction for multiple comparisons. MIP-1b showed a suggestive association with AD, with P = 0.024 (P-value < 0.05, Q > 0.05), whereas stem cell factor (P = 0.031) and ICAM-1 (P = 0.016) showed suggestive associations with hippocampal volume (P < 0.05 level, Q > 0.05).
Shown are the results derived from the secondary inverse variance-weighted meta-analysis. None of the immune traits survived the multiple testing threshold of false discovery rate < 5% (q < 0.05). BNGF, β-nerve growth factor; CTACK, cutaneous T-cell-attracting chemokine; GRO-α, growth-regulated oncogene α; HGF, hepatocyte growth factor; ICAM-1, intercellular adhesion molecule 1; IL, interleukin; IP-10, interferon γ-induced protein 10; MCP-1, monocyte chemoattractant protein-1; MIF, macrophage migration inhibitory factor; MIG, monokine induced by γ-interferon indicates; MIP-1b, macrophage inflammatory protein-1β; PDGF-bb, platelet-derived growth factor-bb; SCF, stem cell factor; SCGF-b, stem cell growth factor β; TRAIL, TNF-related apoptosis-inducing ligand; VEGF, vascular endothelial growth factor.
Sensitivity analyses
There was evidence for heterogeneity in the primary and the secondary MR IVW analyses as measured by Cochran Q. Alternative tests furthermore revealed varying estimates changing direction for multiple exposures (Supplementary Table S3). Cluster analyses did not reveal clusters of variants (Fig. 3).
Shown are the genetic associations for the individual variants with the exposure and outcome; lines indicate confidence intervals. When restricting to a cluster probability assignment of ≥0.8 and ≥4 variants per cluster, no clusters of variants were identified. AD, Alzheimer’s disease; CRP, C-reactive protein; HV, hippocampal volume; Lp-PLA2, lipoprotein-associated phospholipase A2. The junk cluster denotes variants with estimates that do not fit in any cluster.
Discussion
Exploring genetically predicted circulating biomarkers of immunity and inflammation in an adequately powered two-sample MR approach involving the largest GWAS datasets available, we found no association between genetically predicted circulating levels of immune cells and signaling molecules as primary exposures and AD or hippocampal volume. Similarly, none of the secondary exposures including genetically predicted levels of biomarkers of immunity and inflammation showed statistically significant associations with AD or hippocampal volume. Sensitivity analyses showed evidence for heterogeneity, but we found no clustering of variants.
Our findings are in contrast with observational studies that reported on significant associations between several circulating blood biomarkers of signaling molecules, immune cells, and AD4,6. For example, we expected to find an association between higher platelet count and higher AD risk, as we previously found that an increase of circulating platelets as measured in the blood over time increased AD risk6. However, if the risk factor is a protein biomarker, such as CRP, we can select genetic variants located in or around the coding region for that protein as instruments. This is more difficult for polygenic risk factors such as platelets, as the influence of genetic variants on such a risk factor is unlikely to be specific31. Indeed, we found substantial heterogeneity when studying immune cells and signaling molecules, but could not find meaningful clusters of genetic variants that could have a distinct effect on the risk factor, supporting the conclusion that our findings are truly null. On the other hand, our power calculations revealed that some analyses were underpowered to detect significant associations, e.g., for platelet count and AD. However, these exposures in particular did not even show suggestive associations with AD, implying that estimates are very small and probably—even if sufficiently powered—would not have survived correction for multiple testing.
The strongest suggestive association we found in our study was between CD4 cell count and AD, where higher levels of CD4 cell count increase AD risk, although only one SNP could be used as an instrument. This suggestive association is unexpected, as HIV-associated dementia is accompanied by a lower CD4 cell count32. However, it is recognized that T lymphocytes play a central role in the pathogenesis of multiple sclerosis (MS), with CD4+ T cells predominating in acute MS lesions33. Combined with the recent finding that clonally expanded CD8+ T-effector memory cells have been found in the cerebrospinal fluid of AD patients5, the role of adaptive immunity in AD, in particular T cells and the CD4 subtype, is worth investigating further in relation to AD.
In contrast to our findings, one MR study13 found a protective effect of CRP on AD. An explanation for this difference could be the selection of instruments for CRP. In our study, we used 24 SNPs as instruments that are gene specific for CRP, thereby reducing pleiotropy. When examining CRP further by performing a cluster analysis including all genome-wide significant SNPs, we found no clusters of variants, in particular no cluster forming a biologically meaningful protective pathway of CRP on AD.
Our study has limitations. First, we could not assess competing risk by, e.g., mortality in this study, which could generate paradoxical MR estimates34. Second, we cannot exclude that the additional adjustments for body mass index, alcohol consumption, and smoking status performed in the blood cell trait GWAS12 led to collider bias (i.e., a collider between a genetic variant and confounders of the risk factor-outcome association) during instrument selection. However, the potential impact of such collider bias is likely to be less than other biases34. Third, as we used multiple proposed instruments where effect heterogeneity is likely, effect estimates need to be interpreted with caution. Fourth, for some exposures, especially those reflecting adaptive immunity, we were limited by the few known genome-wide significant genetic variants influencing these traits, potentially leading to weak instrument bias. Targeted studies incorporating further GWAS data on individual circulating adaptive immune biomarkers might reveal additional associations not captured by our approach. Furthermore, despite using the largest available datasets, some of our analyses could be limited by power to detect small but functionally relevant causal effects. This lack of power applies to both the discovery of the exposure and to the outcome. Fifth, the IGAP GWAS dataset contains mainly clinically diagnosed cases of AD (only 8% of cases and controls are pathologically confirmed), thus potentially leading to misclassification of the outcome35. Similarly, although hippocampal atrophy is a hallmark feature of AD, a recently recognized disease entity named limbic-predominant age-related TDP-43 encephalopathy has shown to be mimicking Alzheimer’s type dementia, causing hippocampal and medial temporal lobe atrophy in more than 20% of old demented people36. Sixth, the underlying study populations were of European ancestry, limiting generalizability to other ethnicities. Finally, although we have tried to deal with these factors in our study, LD, pleiotropy, canalization, and population stratification remain potential flaws in the MR approach37.
Future perspectives
SNP and biomarker studies investigating age-related diseases play a crucial role in unraveling the mechanisms underlying disease development. Apart from AD, examples of such studies include suggesting new targets for age-related macular degeneration, amyotrophic lateral sclerosis, and other neurodegenerative disorders38. These various targets need validation in suitable animal models. Creating these animal models can be challenging, as it requires undertaking standardization39. The blood–brain barrier forms an additional layer of complexity, as drug molecules are unable to reach the brain40. Nevertheless, the SNP animal model therapeutics field provides an excellent framework for studying interventions reducing risks. The pace of translation in the field of AD could be accelerated by understanding the causative events and mechanisms in the pathogenesis of AD using this framework. Integrating MR analysis when undertaking such studies could aid in the clinical translation, combined with other techniques involving genetics41,42,43. Despite the many successes in the field of genetics, in total only 53% of phenotypic variance is explained, with known AD SNPs only explaining 31% of the genetic variance44. Thus, the whole spectrum of research, including non-genetics, is needed in order to detect the functional ways to underpin the association between the immune system and the physiopathologic network that facilitates the manifestation of AD. In conclusion, this study suggests that genetically predicted circulating biomarkers of immunity and inflammation are not associated with AD risk or hippocampal volume. Future studies should assess competing risk, explore in more depth the role of adaptive immunity in AD, in particular T cells and the CD4 subtype, and confirm these findings in other ethnicities.
References
Cao, W. & Zheng, H. Peripheral immune system in aging and Alzheimer’s disease. Mol. Neurodegener. 13, 51 (2018).
Heneka, M. T. et al. Neuroinflammation in Alzheimer’s disease. Lancet Neurol. 14, 388–405 (2015).
Kunkle, B. W. et al. Genetic meta-analysis of diagnosed Alzheimer’s disease identifies new risk loci and implicates Aβ, tau, immunity and lipid processing. Nat. Genet. 51, 414–430 (2019).
Darweesh, S. K. L. et al. Inflammatory markers and the risk of dementia and Alzheimer’s disease: a meta-analysis. Alzheimers Dement. 14, 1450–1459 (2018).
Gate, D. et al. Clonally expanded CD8 T cells patrol the cerebrospinal fluid in Alzheimer’s disease. Nature 577, 399–404 (2020).
van der Willik, K. D. et al. Balance between innate versus adaptive immune system and the risk of dementia: a population-based cohort study. J. Neuroinflamm. 16, 68 (2019).
Smith, G. D. & Ebrahim, S. Data dredging, bias, or confounding. BMJ 325, 1437–1438 (2002).
Smith, G. D. & Ebrahim, S. ‘Mendelian randomization’: can genetic epidemiology contribute to understanding environmental determinants of disease? Int. J. Epidemiol. 32, 1–22 (2003).
Prins, B. P. et al. Investigating the causal relationship of C-reactive protein with 32 complex somatic and psychiatric outcomes: a large-scale Cross-Consortium Mendelian Randomization Study. PLoS Med. 13, e1001976 (2016).
Larsson, S. C. et al. Modifiable pathways in Alzheimer’s disease: Mendelian randomisation analysis. BMJ 359, j5375 (2017).
Tsui, A. & Davis, D. Systemic inflammation and causal risk for Alzheimer’s dementia: Possibilities and limitations of a Mendelian randomization approach. Aging Med. (Milton) 1, 249–253 (2018).
Astle, W. J. et al. The allelic landscape of human blood cell trait variation and links to common complex disease. Cell 167, 1415–1429 (2016). e1419.
Ligthart, S. et al. Genome analyses of >200,000 individuals identify 58 loci for chronic inflammation and highlight pathways that link inflammation and complex disorders. Am. J. Hum. Genet. 103, 691–706 (2018).
Hibar, D. P. et al. Novel genetic loci associated with hippocampal volume. Nat. Commun. 8, 13624 (2017).
Foley, C. N., Kirk, P. D. W. & Burgess, S. MR-Clust: clustering of genetic variants in Mendelian randomization with similar causal estimates. Bioinformatics 37, 531–541 (2021).
Lin, B. D. et al. 2SNP heritability and effects of genetic variants for neutrophil-to-lymphocyte and platelet-to-lymphocyte ratio. J. Hum. Genet. 62, 979–988 (2017).
Georgakis, M. K. et al. Interleukin-6 signaling effects on ischemic stroke and other cardiovascular outcomes: a Mendelian randomization study. Circ. Genom. Precis. Med. 13, e002872 (2020).
de Vries, P. S. et al. A meta-analysis of 120 246 individuals identifies 18 new loci for fibrinogen concentration. Hum. Mol. Genet. 25, 358–370 (2016).
Casas, J. P. et al. PLA2G7 genotype, lipoprotein-associated phospholipase A2 activity, and coronary heart disease risk in 10 494 cases and 15 624 controls of European Ancestry. Circulation 121, 2284–2293 (2010).
Burgess, S., Thompson, S. G. & Collaboration, C. C. G. Avoiding bias from weak instruments in Mendelian randomization studies. Int. J. Epidemiol. 40, 755–764 (2011).
Ahola-Olli, A. V. et al. Genome-wide association study identifies 27 loci influencing concentrations of circulating cytokines and growth factors. Am. J. Hum. Genet 100, 40–50 (2017).
Georgakis, M. K. et al. Genetically determined levels of circulating cytokines and risk of stroke. Circulation 139, 256–268 (2019).
Burgess, S. Sample size and power calculations in Mendelian randomization with a single instrumental variable and a binary outcome. Int. J. Epidemiol. 43, 922–929 (2014).
Burgess, S., Butterworth, A. & Thompson, S. G. Mendelian randomization analysis with multiple genetic variants using summarized data. Genet. Epidemiol. 37, 658–665 (2013).
Greco, M. F., Minelli, C., Sheehan, N. A. & Thompson, J. R. Detecting pleiotropy in Mendelian randomisation studies with summary data and a continuous outcome. Stat. Med. 34, 2926–2940 (2015).
Bowden, J., Davey, Smith, G., Haycock, P. C. & Burgess, S. Consistent estimation in Mendelian randomization with some invalid instruments using a weighted median estimator. Genet. Epidemiol. 40, 304–314 (2016).
Hartwig, F. P., Davey Smith, G. & Bowden, J. Robust inference in summary data Mendelian randomization via the zero modal pleiotropy assumption. Int. J. Epidemiol. 46, 1985–1998 (2017).
Bowden, J., Davey Smith, G. & Burgess, S. Mendelian randomization with invalid instruments: effect estimation and bias detection through Egger regression. Int. J. Epidemiol. 44, 512–525 (2015).
Burgess, S., Foley, C. N., Allara, E., Staley, J. R. & Howson, J. M. M. A robust and efficient method for Mendelian randomization with hundreds of genetic variants. Nat. Commun. 11, 376 (2020).
Verbanck, M., Chen, C. Y., Neale, B. & Do, R. Detection of widespread horizontal pleiotropy in causal relationships inferred from Mendelian randomization between complex traits and diseases. Nat. Genet. 50, 693–698 (2018).
Burgess, S., Butterworth, A. S. & Thompson, J. R. Beyond Mendelian randomization: how to interpret evidence of shared genetic predictors. J. Clin. Epidemiol. 69, 208–216 (2016).
Kaul, M. HIV-1 associated dementia: update on pathological mechanisms and therapeutic approaches. Curr. Opin. Neurol. 22, 315–320 (2009).
Chitnis, T. The role of CD4 T cells in the pathogenesis of multiple sclerosis. Int. Rev. Neurobiol. 79, 43–72 (2007).
Schooling, C. M., Lopez, P., Yang, Z., Au Yeung, S. L. & Huang, J. V. Bias from competing risk before recruitment in Mendelian Randomization studies of conditions with shared etiology. Preprint at bioRxiv https://doi.org/10.1101/716621 (2019).
Andrews, S. J., Fulton-Howard, B. & Goate, A. Interpretation of risk loci from genome-wide association studies of Alzheimer’s disease. Lancet Neurol. 19, 326–335 (2020).
Nelson, P. T. et al. Limbic-predominant age-related TDP-43 encephalopathy (LATE): consensus working group report. Brain 142, 1503–1527 (2019).
Burgess, S., Timpson, N. J., Ebrahim, S., Davey & Smith, G. Mendelian randomization: where are we now and where are we going? Int. J. Epidemiol. 44, 379–388 (2015).
Anand, A., Gupta, P. K., Sharma, N. K. & Prabhakar, S. Soluble VEGFR1 (sVEGFR1) as a novel marker of amyotrophic lateral sclerosis (ALS) in the North Indian ALS patients. Eur. J. Neurol. 19, 788–792 (2012).
Anand, A., Banik, A., Thakur, K. & Masters, C. L. The animal models of dementia and Alzheimer’s disease for pre-clinical testing and clinical translation. Curr. Alzheimer Res. 9, 1010–1029 (2012).
Goyal, K., Koul, V., Singh, Y. & Anand, A. Targeted drug delivery to central nervous system (CNS) for the treatment of neurodegenerative disorders: trends and advances. Cent. Nerv. Syst. Agents Med. Chem. 14, 43–59 (2014).
International Genomics of Alzheimer’s Disease Consortium Convergent genetic and expression data implicate immunity in Alzheimer’s disease. Alzheimers Dement. 11, 658–671 (2015).
Chen, L. et al. Genetic drivers of epigenetic and transcriptional variation in human immune cells. Cell 167, 1398–1414 (2016). e1324.
Hu, Y. S., Xin, J., Hu, Y., Zhang, L. & Wang, J. Analyzing the genes related to Alzheimer’s disease via a network and pathway-based approach. Alzheimers Res. Ther. 9, 29 (2017).
Ridge, P. G. et al. Assessment of the genetic variance of late-onset Alzheimer’s disease. Neurobiol. Aging 41, e213–200 (2016). e220.
Lin, B. D. et al. Heritability and GWAS studies for monocyte-lymphocyte ratio. Twin Res. Hum. Genet. 20, 97–107 (2017).
Ferreira, M. A. et al. Quantitative trait loci for CD4:CD8 lymphocyte ratio are associated with risk of type 1 diabetes and HIV-1 immune control. Am. J. Hum. Genet. 86, 88–92 (2010).
Ward-Caviness, C. K. et al. Mendelian randomization evaluation of causal effects of fibrinogen on incident coronary heart disease. PLoS ONE 14, e0216222 (2019).
Grallert, H. et al. Eight genetic loci associated with variation in lipoprotein-associated phospholipase A2 mass and activity and coronary heart disease: meta-analysis of genome-wide association studies from five community-based studies. Eur. Heart J. 33, 238–251 (2012).
Matteini, A. M. et al. Novel gene variants predict serum levels of the cytokines IL-18 and IL-1ra in older adults. Cytokine 65, 10–16 (2014).
Interleukin 1 Genetics Consortium Cardiometabolic effects of genetic upregulation of the interleukin 1 receptor antagonist: a Mendelian randomisation analysis. Lancet Diabetes Endocrinol. 3, 243–253 (2015).
Barbalic, M. et al. Large-scale genomic studies reveal central role of ABO in sP-selectin and sICAM-1 levels. Hum. Mol. Genet. 19, 1863–1872 (2010).
Pare, G. et al. Genome-wide association analysis of soluble ICAM-1 concentration reveals novel associations at the NFKBIK, PNPLA3, RELA, and SH2B3 loci. PLoS Genet. 7, e1001374 (2011).
Kunkle, B. W. et al. Author Correction: Genetic meta-analysis of diagnosed Alzheimer’s disease identifies new risk loci and implicates Abeta, tau, immunity and lipid processing. Nat. Genet. 51, 1423–1424 (2019).
Acknowledgements
This work was supported by the European Union’s Horizon 2020 research and innovation program (grant numbers 667375 [“CoSTREAM”] and 666881 [“SVDs@target”]) and Alzheimer Nederland. The funding source had no role in study design, collection, analysis, interpretation of data, writing of the report, or decision to submit the article for publication.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
41398_2021_1400_MOESM1_ESM.docx
Circulating biomarkers of immunity and inflammation and risk of Alzheimer’s disease and hippocampal volume: A mendelian randomization study
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Fani, L., Georgakis, M.K., Ikram, M.A. et al. Circulating biomarkers of immunity and inflammation, risk of Alzheimer’s disease, and hippocampal volume: a Mendelian randomization study. Transl Psychiatry 11, 291 (2021). https://doi.org/10.1038/s41398-021-01400-z
Received:
Revised:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41398-021-01400-z
This article is cited by
-
The role of peripheral inflammatory insults in Alzheimer’s disease: a review and research roadmap
Molecular Neurodegeneration (2023)
-
Mendelian randomization highlights significant difference and genetic heterogeneity in clinically diagnosed Alzheimer’s disease GWAS and self-report proxy phenotype GWAX
Alzheimer's Research & Therapy (2022)
-
The role of the adaptive immune system and T cell dysfunction in neurodegenerative diseases
Journal of Neuroinflammation (2022)
-
Immune system-wide Mendelian randomization and triangulation analyses support autoimmunity as a modifiable component in dementia-causing diseases
Nature Aging (2022)
-
Systemic inflammatory markers in relation to cognitive function and measures of brain atrophy: a Mendelian randomization study
GeroScience (2022)