Abstract
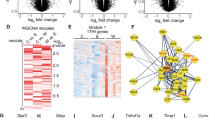
Alcohol withdrawal is a clinically important consequence and potential driver of Alcohol Use Disorder. However, susceptibility to withdrawal symptoms, ranging from craving and anxiety to seizures and delirium, varies greatly. Selectively bred Withdrawal Seizure-Prone (WSP) and Seizure-Resistant (WSR) mice are an animal model of differential susceptibility to withdrawal and phenotypes with which withdrawal severity correlates. To identify innate drivers of alcohol withdrawal severity, we performed a multi-omic study of the WSP and WSR lines and F2 mice derived from them, using genomic, genetic, and transcriptomic analyses. Genes implicated in seizures and epilepsy were over-represented among those that segregated between WSP and WSR mice and that displayed differential expression in F2 mice high and low in withdrawal. Quantitative trait locus (QTL) analysis of ethanol withdrawal convulsions identified several genome-wide significant loci and pointed to genes that modulate potassium channel function and neural excitability. Perturbations of expression of genes involved in synaptic transmission, including GABAergic and glutamatergic genes, were prominent in prefrontal cortex transcriptome. Expression QTL (eQTL) analysis fine mapped genes within the peak ethanol withdrawal QTL regions. Genetic association analysis in human subjects provided converging evidence for the involvement of those genes in severity of alcohol withdrawal and dependence. Our results reveal a polygenic network and neural signaling pathways contributing to ethanol withdrawal seizures and related phenotypes that overlap with genes modulating epilepsy and neuronal excitability.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
Data availability
Sequences of the WSP1, WSR1, WSP2, and WSR2 exomes and WSP/WSR F2 transcriptomes have been deposited in NCBI as a BioProject (accession no. PRJNA875906).
References
Hall W, Zador D. The alcohol withdrawal syndrome. Lancet. 1997;349:1897–1900.
Gilpin NW, Koob GF. Neurobiology of alcohol dependence: focus on motivational mechanisms. Alcohol Res Health. 2008;31:185–95.
Goldman D, Oroszi G, Ducci F. The genetics of addictions: uncovering the genes. Nat Rev Genet. 2005;6:521–32.
Edenberg HJ, Koller DL, Xuei X, Wetherill L, McClintick JN, Almasy L, et al. Genome-wide association study of alcohol dependence implicates a region on chromosome 11. Alcohol Clin Exp Res. 2010;34:840–52.
Walters RK, Polimanti R, Johnson EC, McClintick JN, Adams MJ, Adkins AE, et al. Transancestral GWAS of alcohol dependence reveals common genetic underpinnings with psychiatric disorders. Nat Neurosci. 2018;21:1656–69.
Zhou Z, Karlsson C, Liang T, Xiong W, Kimura M, Tapocik JD, et al. Loss of metabotropic glutamate receptor 2 escalates alcohol consumption. Proc Natl Acad Sci USA. 2013;110:16963–8.
Zhou Z, Yuan Q, Mash DC, Goldman D. Substance-specific and shared transcription and epigenetic changes in the human hippocampus chronically exposed to cocaine and alcohol. Proc Natl Acad Sci USA. 2011;108:6626–31.
Mhatre MC, Ticku MK. Chronic ethanol administration alters gamma-aminobutyric acid A receptor gene expression. Mol Pharm. 1992;42:415–22.
Herman MA, Roberto M. Cell-type-specific tonic GABA signaling in the rat central amygdala is selectively altered by acute and chronic ethanol. Addict Biol. 2016;21:72–86.
Christian DT, Alexander NJ, Diaz MR, McCool BA. Thalamic glutamatergic afferents into the rat basolateral amygdala exhibit increased presynaptic glutamate function following withdrawal from chronic intermittent ethanol. Neuropharmacology. 2013;65:134–42.
Griffin WC 3rd, Haun HL, Hazelbaker CL, Ramachandra VS, Becker HC. Increased extracellular glutamate in the nucleus accumbens promotes excessive ethanol drinking in ethanol-dependent mice. Neuropsychopharmacology. 2014;39:707–17.
Abrahao KP, Salinas AG, Lovinger DM. Alcohol and the brain: neuronal molecular targets, synapses, and circuits. Neuron. 2017;96:1223–38.
Tabarki B, AlMajhad N, AlHashem A, Shaheen R, Alkuraya FS. Homozygous KCNMA1 mutation as a cause of cerebellar atrophy, developmental delay and seizures. Hum Genet. 2016;135:1295–8.
Krapivinsky G, Medina I, Krapivinsky L, Gapon S, Clapham DE. SynGAP-MUPP1-CaMKII synaptic complexes regulate p38 MAP kinase activity and NMDA receptor-dependent synaptic AMPA receptor potentiation. Neuron. 2004;43:563–74.
Fehr C, Shirley RL, Belknap JK, Crabbe JC, Buck KJ. Congenic mapping of alcohol and pentobarbital withdrawal liability loci to a <1 centimorgan interval of murine chromosome 4: identification of Mpdz as a candidate gene. J Neurosci. 2002;22:3730–8.
Bubier JA, Jay JJ, Baker CL, Bergeson SE, Ohno H, Metten P, et al. Identification of a QTL in Mus musculus for alcohol preference, withdrawal, and Ap3m2 expression using integrative functional genomics and precision genetics. Genetics. 2014;197:1377–93.
Crabbe JC, Kosobud A, Young ER, Tam BR, McSwigan JD. Bidirectional selection for susceptibility to ethanol withdrawal seizures in Mus musculus. Behav Genet. 1985;15:521–36.
Metten P, Schlumbohm JP, Huang LC, Greenberg GD, Hack WR, Spence SE, et al. An alcohol withdrawal test battery measuring multiple behavioral symptoms in mice. Alcohol. 2018;68:19–35.
McClearn GE, Wilson JR, Meredith JE. The use of isogenic and heterogenic mouse stocks in behavioral research. In: G Lindzey DT (ed). Contributions to behavior-genetic analysis: the mouse as a prototype. Appleton-Century-Crofts: New York, 1970, pp 3–22.
Crabbe JC, Spence SE, Huang LC, Cameron AJ, Schlumbohm JP, Barkley-Levenson AM, et al. Ethanol drinking in Withdrawal Seizure-Prone and -Resistant selected mouse lines. Alcohol. 2013;47:381–9.
Hartmann MC, Holbrook SE, Haney MM, Crabbe JC, Rosenwasser AM. Affective behavior in withdrawal seizure-prone and withdrawal seizure-resistant mice during long-term alcohol abstinence. Alcohol Clin Exp Res. 2019;43:1478–85.
Philibin SD, Cameron AJ, Schlumbohm JP, Metten P, Crabbe JC. Ethanol withdrawal-induced motor impairment in mice. Psychopharmacology. 2012;220:367–78.
Bergeson SE, Kyle Warren R, Crabbe JC, Metten P, Gene Erwin V, Belknap JK. Chromosomal loci influencing chronic alcohol withdrawal severity. Mamm Genome. 2003;14:454–63.
Mark GP, Finn DA. The relationship between hippocampal acetylcholine release and cholinergic convulsant sensitivity in withdrawal seizure-prone and withdrawal seizure-resistant selected mouse lines. Alcohol Clin Exp Res. 2002;26:1141–52.
Plotkin SR, Banks WA, Cohn CS, Kastin AJ. Withdrawal from alcohol in withdrawal seizure-prone and -resistant mice: evidence for enkephalin resistance. Pharm Biochem Behav. 2001;68:379–87.
Snelling C, Tanchuck-Nipper MA, Ford MM, Jensen JP, Cozzoli DK, Ramaker MJ, et al. Quantification of ten neuroactive steroids in plasma in Withdrawal Seizure-Prone and -Resistant mice during chronic ethanol withdrawal. Psychopharmacology. 2014;231:3401–14.
Keir WJ, Morrow AL. Differential expression of GABAA receptor subunit mRNAs in ethanol-naive withdrawal seizure resistant (WSR) vs. withdrawal seizure prone (WSP) mouse brain. Brain Res Mol Brain Res. 1994;25:200–8.
Mason JN, Eshleman AJ, Belknap JK, Crabbe JC, Loftis JM, Macey TA, et al. NMDA receptor subunit mRNA and protein expression in ethanol-withdrawal seizure-prone and -resistant mice. Alcohol Clin Exp Res. 2001;25:651–60.
Langfelder P, Horvath S. WGCNA: an R package for weighted correlation network analysis. BMC Bioinform. 2008;9:559.
Szklarczyk D, Gable AL, Lyon D, Junge A, Wyder S, Huerta-Cepas J, et al. STRING v11: protein-protein association networks with increased coverage, supporting functional discovery in genome-wide experimental datasets. Nucleic Acids Res. 2019;47:D607–d613.
Ridgway LD, Kim EY, Dryer SE. MAGI-1 interacts with Slo1 channel proteins and suppresses Slo1 expression on the cell surface. Am J Physiol Cell Physiol. 2009;297:C55–65.
Tanemoto M, Abe T, Uchida S, Kawahara K. Mislocalization of K+ channels causes the renal salt wasting in EAST/SeSAME syndrome. FEBS Lett. 2014;588:899–905.
Shirley RL, Walter NA, Reilly MT, Fehr C, Buck KJ. Mpdz is a quantitative trait gene for drug withdrawal seizures. Nat Neurosci. 2004;7:699–700.
Hu JH, Park JM, Park S, Xiao B, Dehoff MH, Kim S, et al. Homeostatic scaling requires group I mGluR activation mediated by Homer1a. Neuron. 2010;68:1128–42.
Kammermeier PJ. Endogenous homer proteins regulate metabotropic glutamate receptor signaling in neurons. J Neurosci. 2008;28:8560–7.
Medina-Kauwe LK, Tobin AJ, De Meirleir L, Jaeken J, Jakobs C, Nyhan WL, et al. 4-Aminobutyrate aminotransferase (GABA-transaminase) deficiency. J Inherit Metab Dis. 1999;22:414–27.
Youle RJ, van der Bliek AM. Mitochondrial fission, fusion, and stress. Science. 2012;337:1062–5.
Tonduti D, Dorboz I, Imbard A, Slama A, Boutron A, Pichard S, et al. New spastic paraplegia phenotype associated to mutation of NFU1. Orphanet J Rare Dis. 2015;10:13.
Zhang R, Engler A, Taylor V. Notch: an interactive player in neurogenesis and disease. Cell Tissue Res. 2018;371:73–89.
Noebels JL. Sodium channel gene expression and epilepsy. Novartis Found Symp. 2002;241:109–20. discussion 120-103, 226-132
Reynolds C, King MD, Gorman KM. The phenotypic spectrum of SCN2A-related epilepsy. Eur J Paediatr Neurol. 2020;24:117–22.
Erol A, Karpyak VM. Sex and gender-related differences in alcohol use and its consequences: contemporary knowledge and future research considerations. Drug Alcohol Depend. 2015;156:1–13.
Sharrett-Field L, Butler TR, Reynolds AR, Berry JN, Prendergast MA. Sex differences in neuroadaptation to alcohol and withdrawal neurotoxicity. Pflug Arch. 2013;465:643–54.
Sullivan JT, Sykora K, Schneiderman J, Naranjo CA, Sellers EM. Assessment of alcohol withdrawal: the revised clinical institute withdrawal assessment for alcohol scale (CIWA-Ar). Br J Addict. 1989;84:1353–7.
Zhou Z, Roy A, Lipsky R, Kuchipudi K, Zhu G, Taubman J, et al. Haplotype-based linkage of tryptophan hydroxylase 2 to suicide attempt, major depression, and cerebrospinal fluid 5-hydroxyindoleacetic acid in 4 populations. Arch Gen Psychiatry. 2005;62:1109–18.
Khalili AA, Loudet O. eqtl: tools for analyzing eQTL experiments: a complementary to Karl Broman’s ‘qtl’ package for genome-wide analysis. 2012.
Broman KW, Wu H, Sen S, Churchill GA. R/qtl: QTL mapping in experimental crosses. Bioinformatics. 2003;19:889–90.
Clayton D. snpStats: SnpMatrix and XSnpMatrix classes and methods. R package version 1.16.0. 2014.
Acknowledgements
The authors wish to thank many former technicians, interns, post-doctoral fellows, and graduate students of the Crabbe lab whose efforts in breeding, testing, and scoring withdrawal behaviors were invaluable. This work was supported by the Intramural Research Program of the NIAAA; NIAAA Grant AA10760 (to JCC); and US Department of Veteran’s Affairs Grant BX000313 (to JCC).
Author information
Authors and Affiliations
Contributions
ZZ, PM, JCC, and DG designed the research project. ZZ, PM, HS, CAH, and CM carried out or contributed to the experiments. ZZ, QY, and P-HS analyzed the data. ZZ, PM, JCC, and DG wrote the paper.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
About this article
Cite this article
Zhou, Z., Metten, P., Yuan, Q. et al. Genetic and genomic signatures in ethanol withdrawal seizure-prone and seizure-resistant mice implicate genes involved in epilepsy and neuronal excitability. Mol Psychiatry 27, 4611–4623 (2022). https://doi.org/10.1038/s41380-022-01799-x
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41380-022-01799-x