Abstract
Despite recent refinements in the diagnostic and prognostic assessment of CEBPA mutations in AML, several questions remain open, i.e. implications of different types of basic region leucin zipper (bZIP) mutations, the role of co-mutations and the allelic state. Using pooled primary data analysis on 1010 CEBPA-mutant adult AML patients, a comparison was performed taking into account the type of mutation (bZIP: either typical in-frame insertion/deletion (InDel) mutations (bZIPInDel), frameshift InDel or nonsense mutations inducing translational stop (bZIPSTOP) or single base-pair missense alterations (bZIPms), and transcription activation domain (TAD) mutations) and the allelic state (single (smCEBPA) vs. double mutant (dmCEBPA)). Only bZIPInDel patients had significantly higher rates of complete remission and longer relapse free and overall survival (OS) compared with all other CEBPA-mutant subgroups. Moreover, co-mutations in bZIPInDel patients (e.g. GATA2, FLT3, WT1 as well as ELN2022 adverse risk aberrations) had no independent impact on OS, whereas in non-bZIPInDel patients, grouping according to ELN2022 recommendations added significant prognostic information. In conclusion, these results demonstrate bZIPInDel mutations to be the major independent determinant of outcome in CEBPA-mutant AML, thereby refining current classifications according to WHO (including all dmCEBPA and smCEBPA bZIP) as well as ELN2022 and ICC recommendations (including CEBPA bZIPms).

Similar content being viewed by others
Introduction
Mutations of the gene encoding the CCAAT-enhancer binding protein alpha (CEBPA) transcription factor are common genetic alterations in acute myeloid leukemia (AML). Since their first description [1], several groups have investigated CEBPA mutations in AML (reviewed in [2]). After initial studies suggested that all patients with CEBPA mutations carry a more favorable outcome [3,4,5,6,7], subsequent analyses have consistently demonstrated that this improved prognosis is confined to biallelic or double mutations (dmCEBPA). DmCEBPA was shown to be associated with a distinct biology and to confer a more favorable clinical outcome, including higher rates of complete remission (CR), reduced relapse risk, and increased overall survival (OS), whereas single allele mutations were considered prognostically irrelevant [8,9,10,11,12,13,14]. However, several recent reports looking in more detail for the impact of individual mutations in CEBPA suggested that the specific clinical and molecular characteristics as well as the favorable prognosis were restricted to mutations within the basic leucine zipper region (bZIP) region of CEBPA, irrespective of their occurrence as double or single mutation [15,16,17]. Gene expression analysis further supports a unique biology of CEBPA bZIP mutations in AML [15, 16]. However, there is evidence that even within this mutational subgroup, biological differences may exist depending on the particular type of CEBPA bZIP mutation. Our group observed significant differences in outcome and molecular profiles when comparing patients with in-frame CEBPA bZIP mutations (either in-frame insertions/deletions or single base pair missense mutations) and patients with frameshift or nonsense mutations [16].
These findings have provided the basis for a refined biological and clinical classification of CEBPA mutations. CEBPA bZIP in-frame mutations are now being classified as favorable risk entity in the 2022 update of the European LeukemiaNet (ELN) recommendations on genetic risk classification [18] and in the International Consensus Classification [19]. However, the 2022 WHO classification still defines the CEBPA mutational class via the presence of a double mutant allelic status or a single mutant bZIP mutation regardless of the type of mutation(s)/exact DNA alteration [20]. This persistent ambiguity might be due to the fact that few reports have investigated the impact of different CEBPA mutational constellations in more detail, so the evidence supporting either of these modifications is still limited.
To gain further insights into the impact of different mutational subtypes, in particular the spectrum of CEBPA bZIP mutations, a pooled primary data analysis was performed involving detailed sequencing data as well as clinical variables and treatment outcome from more than 1000 CEBPA mutant AML patients. The main objective of this study was to investigate potential differences between different types of bZIP mutations and to examine the relevance of the allelic status (double vs. single mutant).
Our results, generated in the largest cohorts of patients with CEBPA mutant AML reported so far, strongly support the notion that the CEBPA bZIPInDel genotype introduced in this work (bZIP in-frame insertions/deletions, double and single mutant) shows a specific biology and favorable prognostic implications, whereas the other CEBPA mutational subgroups, i.e. TAD mutations as well as bZIP missense and frameshift/nonsense mutations, differ substantially with respect to most clinical as well as molecular factors studied.
This analysis establishes the basis for a more accurate refinement of current classifications and highlights the need for additional research efforts to elucidate the specific biological effects of CEBPA bZIPInDel mutations and their role in leukemogenesis.
Materials/subjects and methods
Anonymized individual patient data from 1010 adult patients aged 16–85 years (median 52 years) were collected from six European AML study groups and registries: n = 98 ALFA, n = 104 AMLCG, n = 191 HOVON, n = 200 Munich Leukemia Lab, n = 240 SAL, n = 177 MRC/NCRI. Patients included were treated between 1989 and 2019, with the majority of individuals treated between 1996 and 2016 (89.4%; Table S1). Patients were treated in prospective trials (details on study protocols are given in the supplement) involving risk stratified post induction therapy according to cytogenetic risk groups, including the option for an allogeneic hematopoietic cell transplantation (alloHCT) in CR1, or recruited to AML registries and biorepositories. Only 40 patients (3.9%) were treated after the release of the ELN 2016 guidelines, which proposed stratification by allelic status (dmCEBPA vs. smCEBPA) within the group of CEBPA-mutant patients. Otherwise, CEBPA mutations were not used for risk stratification in any of the trials. For each individual patient, a predefined minimal data set was collected, including clinical variables, i.e. age, sex, date of AML diagnosis, type of AML (de novo or secondary/therapy-related), bone marrow (BM) blast count, white blood cell (WBC) count, type of and response to induction chemotherapy, date of alloHCT in CR1, date of alloHCT beyond CR1 and events (i.e. induction failure, relapse, death) as well as genetic variables (karyotype, mutational status of NPM1, FLT3, GATA2, DNMT3A, IDH1, IDH2, WT1 and other genes, if available). Co-mutational data sufficient for genetic risk stratification according to the ELN2022 guidelines were available for 645 patients (63.8%).
The information collected included complete sequencing results (performed either by Sanger sequencing or next generation sequencing; NGS) of the entire CEBPA gene (Genbank Accession No. NM_004364.2). All retrieved CEBPA sequences were evaluated for the precise localization of the mutation, i.e. bZIP vs. transcription activation domains [17] 1 and 2, allelic status (smCEBPA vs. dmCEBPA) as well as the type of mutation, i.e. insertions/deletions either in-frame or frameshift, missense mutations as well as nonsense mutations.
This study was performed in accordance with the Declaration of Helsinki, all clinical studies and data registries were approved by the local institutional review boards, and written informed consent was obtained from all patients through the participating centers.
For statistical analysis, comparisons of categorical variables between groups were done with the Chi-squared test. Continuous variables were compared with the Kruskal-Wallis-Test between groups. OS was calculated from date of study entry until death, or last follow-up visit. Relapse free survival (RFS) was calculated from date of first remission until date of relapse, date of death, or date of last follow-up visit. Survival endpoints were analyzed with the Kaplan-Meier method. Cox regression models were fitted to estimate hazard ratios. AlloHCT as adjusting variable in multiple models was modeled as time-dependent covariate. Univariate and multivariate logistic regression models were used to estimate odds ratios for achievement of CR1. Individual patient data of the different study groups were pooled. All Cox regression models were stratified for study group. In logistic regression models study group was modeled as a factor. To estimate variability between study groups, all analyses were conducted per study group and interaction of CEBPA categories with study group was assessed. Additionally, the estimates from the study groups were pooled via inverse variance method and heterogeneity statistics were estimated. AlloHCT was modeled as time-dependent covariate. A landmark of 3 months from study entry was applied to reduce bias due to early deaths disqualifying patients for alloHCT.
Results
CEBPA mutational status
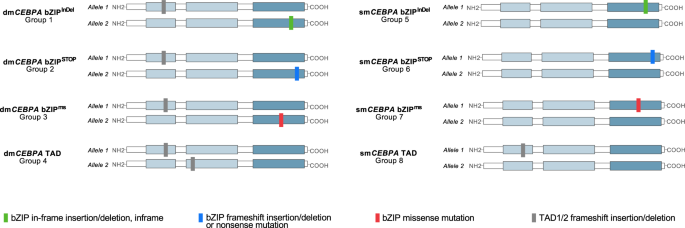
Of the 1010 patients, 661 patients (65.4%) showed mutations affecting the bZIP-domain of CEBPA encompassing amino acids (AA) 272–358 [1]. As illustrated in Fig. 1, mutations in the CEBPA bZIP region were typically in-frame insertions or deletions (n = 491 pts; 74.3%; referred to as bZIPInDel), i.e. (multiples of) 3 bp affecting the DNA-binding-, fork- or bZIP-region. Frameshift insertions/deletions or nonsense mutations in bZIP, causing premature termination of transcription (referred to as bZIPSTOP), as well as missense mutations (bZIPms), causing single AA changes, were less common and were observed in 81 (12.3%) and 89 (13.5%) of patients, respectively. Interestingly, the different types of mutations showed a non-random distribution, with bZIPms mainly clustering in certain critical AA positions in the DNA-binding basic region, i.e. AA297 and AA300. In contrast, most bZIPInDel mutations affected the hinge/fork region of the bZIP-domain, i.e., the 14 AAs preceding the first leucine residue of the leucine zipper at position 317 [21], with the most frequent InDel mutations at AA312 and AA313 (Fig. 1). In contrast to the bZIP region, the TAD domains almost uniformly harbored frameshift InDel or nonsense mutations, creating a premature termination codon.
To assess the impact of different mutational constellations, 8 subgroups were generated, taking into account both type and site of mutation (bZIPInDel vs. bZIPms vs. bZIPSTOP vs. TAD) and allelic status (dmCEBPA vs. smCEBPA). The characteristic mutational constellations of the different mutational subgroups (dmCEBPA bZIPInDel (Gr1), dmCEBPA bZIPSTOP (Gr2), dmCEBPA bZIPms (Gr3), dmCEBPA TAD (Gr4), smCEBPA bZIPInDel (Gr5), smCEBPA bZIPSTOP (Gr6), smCEBPA bZIPms (Gr7) and smCEBPA TAD (Gr8)) are illustrated in Fig. 2.
In this cohort, patients with dmCEBPA predominantly harbored a combination of a bZIP and a TAD mutation (n = 475, 87%). In rare cases (n = 11), a combination of two bZIP mutations was present, in all of which one of the two mutations was a bZIPInDel mutation. These cases were assigned to Gr1. 60 dmCEBPA patients (11%) showed alterations affecting only the TAD regions, most frequently being a combination of TAD1 and TAD2 mutations (Gr4). In 10 patients with dmCEBPA, more than two CEBPA mutations were detected, including the following combinations: 2 bZIPInDel+TAD (assigned to Gr1); bZIPInDel+2 TAD (assigned to Gr1, n = 2), bZIPInDel + bZIPms+TAD (assigned to Gr1, n = 2), bZIPSTOP+2 TAD (assigned to Gr2); bZIPms+2 TAD (assigned to Gr3), 2 bZIPms+2 TAD (assigned to Gr3), 3 TAD (assigned to Gr4),
CEBPA mutational subgroups and clinical characteristics
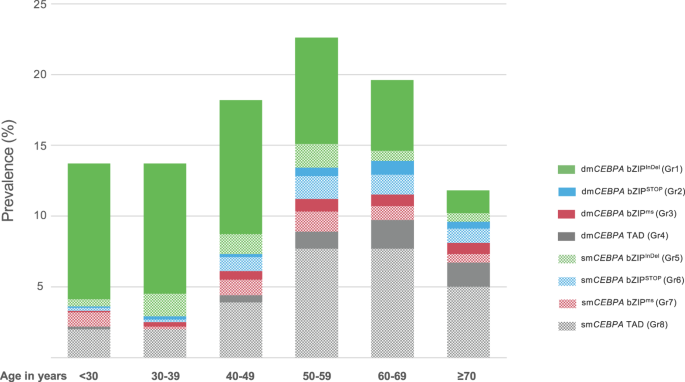
As outlined in Table 1, the association of several clinical parameters differed substantially between the defined mutational subgroups. Patients with bZIPInDel mutations, i.e., Gr1 and Gr5, were significantly younger (median age Gr1 42.2 years [IQR 31–54.9]; Gr5 47 years [IQR 39–58]) than those without bZIPInDel mutations (groups 2–4 and groups 6–8) (median age 52–64 years). They also had a higher prevalence of de novo AML (98% and 96%) compared to patients without bZIPInDel mutations (groups 2–4 and groups 6–8), with the latter more frequently evolving as secondary disease after prior MDS or as tAML (rate of de novo AML 81–94%) (Table 1). Categorizing age in 10-year intervals (Fig. 3, Table S2), a continuous decrease in the occurrence of bZIPInDel mutations (especially dmCEBPA bZIPInDel) was seen with increasing age, whereas bZIPSTOP and bZIPms mutations and alterations affecting only the TAD regions were particularly common in older individuals and less prevalent in patients up to the age of 40 years. Other clinical parameters did not differ significantly between subgroups.
Association of CEBPA mutations with other molecular and cytogenetic abnormalities
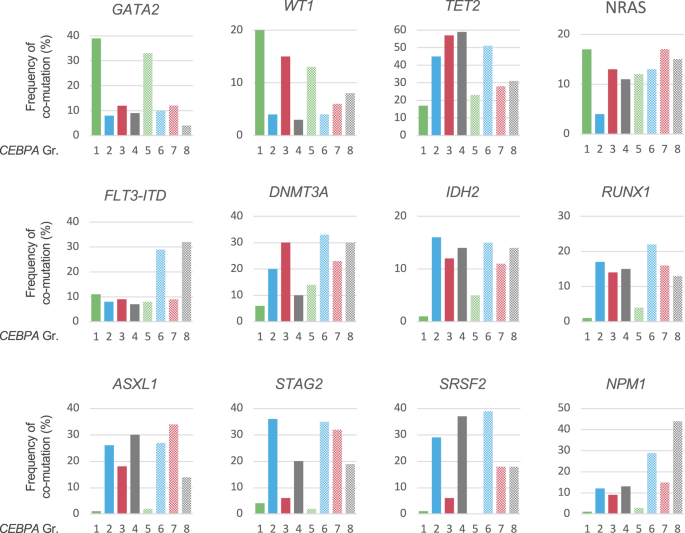
Additional mutations were identified in 861/1010 CEBPA-mutant patients (85.2%). Figure 4 illustrates the distribution of co-mutations in the different CEBPA mutational subgroups. Significant differences were observed for several genes, the most striking being GATA2, which was found mutated in 39 and 33% of patients in Gr1 and Gr5, but only 4–12% in patients carrying non-bZIPInDel mutations (groups 2–4 and 6–8), and NPM1, with only 1 and 3% of patients affected in Gr1 and Gr5, but 10–44% in groups 2–4 and 6–8 (p < 0.001). In general, the spectrum of co-mutations of patients with bZIPInDel mutations differed markedly from the other CEBPA subgroups, the latter more frequently carrying mutations in genes associated with AML after prior MDS, such as spliceosome mutations (i.e. SRSF2, SF3B1, U2AF1 or ZRSR2) as well as alterations associated with DNA-methylation (i.e. DNMT3A, TET2, IDH1 and IDH2) (Fig. 4, Table S3). Besides GATA2, patients with bZIPInDel mutations were more likely to harbor mutations in WT1 (Gr1 bZIPInDel 20%, Gr5 13%, all other groups 3–15%).
Most CEBPA-mutant patients showed a normal karyotype (Table 1). In patients with bZIPInDel mutations, the predominant cytogenetic aberrations were del 9q (Gr1 n = 24/92; Gr5 n = 5/20) and +21 (Gr1 n = 19/92; Gr5 n = 4/20). Among the other CEBPA mutational subgroups, chromosomal abnormalities were more diverse and included −7, −5, +8 and several others, thereby reflecting the spectrum of changes seen in AML in general.
CEBPA mutational subgroups and response to treatment
Treatment response data were available for 992 patients (98.2%). Outcome analysis showed comparability of CR rates as well as RFS and OS between study groups (Table S1). Regarding initial response to therapy within the eight different subgroups, patients with bZIPInDel mutations achieved the highest rates of CR1, with 94.3% in Gr1 (OR 6.38 [3.83–10.63], p < 0.001) and 92.1% in Gr5 (OR 4.51 [1.71–11.86], p = 0.002) compared to CR-rates of 73.1–79.6% observed in the other subgroups (Table 1, Table S4).
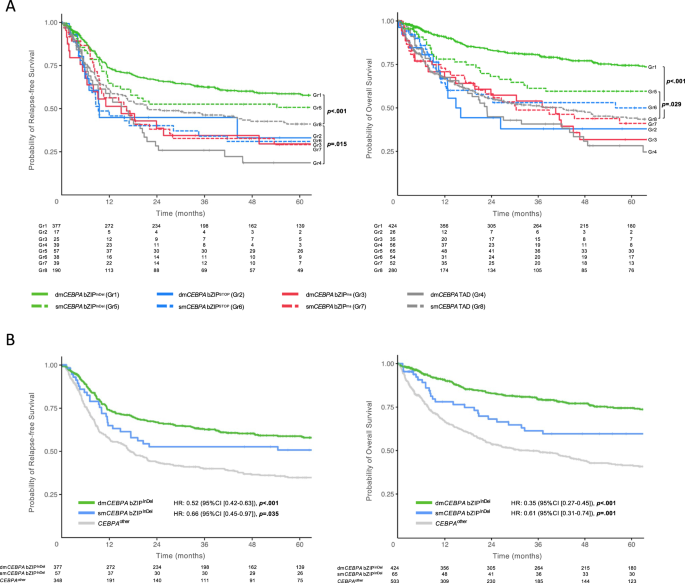
When analyzed according to the eight different mutational subgroups, patients carrying bZIPInDel mutations had a more favorable outcome (RFS and OS) than patients without these mutations (Fig. 5A). In detail, patients in Gr1 showed the longest RFS (median 152 months (HR 0.60 [0.47–0.77]); p < 0.001) and OS (median 215 months, HR 0.37 [0.29–0.46], p < 0.001). Outcome of Gr5 patients was less favorable than Gr1 (median RFS 64 months (HR 0.77 [0.51–1.15]), p = 0.21; median OS 126 months HR = 0.65 [0.44 to 0.96], p = 0.029), but still better than for the other subgroups (median RFS ranged between 9.4 and 21.8 months, median OS 15.7 and 70.9 months) (Tables S5 and S6).
In a subsequent analysis combining groups 2–4 and 6–8 as CEBPAother (reference group), patients with dmCEBPA bZIPInDel as well as smCEBPA bZIPInDel both demonstrated significantly better CR1, RFS and OS compared with CEBPAother patients, which was confirmed in multivariate analysis taking into account the individual study groups and the patient age (CR1: dmCEBPA bZIPInDel OR 5.82 [3.62–9.36], p < 0.001), smCEBPA bZIPInDel OR 4.13 [1.59–10.68], p = 0.003; RFS: dmCEBPA bZIPInDel HR 0.52 [0.42–0.63], p < 0.001, smCEBPA bZIPInDel HR 0.66 [0.45–0.97], p = 0.035; OS: dmCEBPA bZIPInDel HR 0.35 [0.28–0.43], p < 0.001, smCEBPA bZIPInDel HR 0.61 [0.42–0.89], p = 0.011 (Fig. 5B, Tables S7–S9).
AlloHCT performed in CR1 showed no benefit in bZIPInDel patients (HR 1.19 [0.81 to 1.75], p = 0.178) (Fig. S1, Table S10).
Impact of co-mutations on outcome in CEBPA bZIPInDel mutant patients
Several studies have reported on the effect of co-mutations [22,23,24,25,26,27,28] in CEBPA-mutant AML. In patients with dmCEBPA bZIPInDel, a potential concurrent effect on outcome was assessed for the most common alterations previously associated with outcome, i.e. GATA2, TET2, WT1, CSF3R and FLT3-ITD. The RFS in the few CSF3R-mutant patients (n = 15) was significantly reduced in univariate analysis (dmCEBPA bZIPInDel /CSF3Rwt: RFS HR 0.24 [0.14–0.44], p < 0.001) (Fig. S2), which remained significant in multivariate analysis (Table S11), however, this effect did not translate into a difference in overall survival (OS HR 1.42 [0.52–3.89]; p = 0.491). The presence of a co-mutation in TET2 was also associated with worse RFS (dmCEBPA bZIPInDel /TET2wt: RFS HR 0.61 [0.4–0.95], p = 0.028) and OS, (dmCEBPA bZIPInDel /TET2wt: OS HR 0.56 [0.34–0.93], p = 0.025) (Fig. S3), though this effect lost its significance in multivariate analysis (Table S12). FLT3-ITD-positive patients had a shorter RFS in univariate (dmCEBPA bZIPInDel without FLT3-ITD: RFS HR 0.60 [0.38–0.96], p = 0.031) (Fig. S4), but not in multivariate analysis (Table S13). In contrast, the presence of GATA2 and WT1 co-mutations did not significantly affect RFS or OS in this group (Figs. S5, S6).
Impact of ELN2022 adverse molecular and cytogenetic alterations on outcome in CEBPA bZIPInDel mutant AML
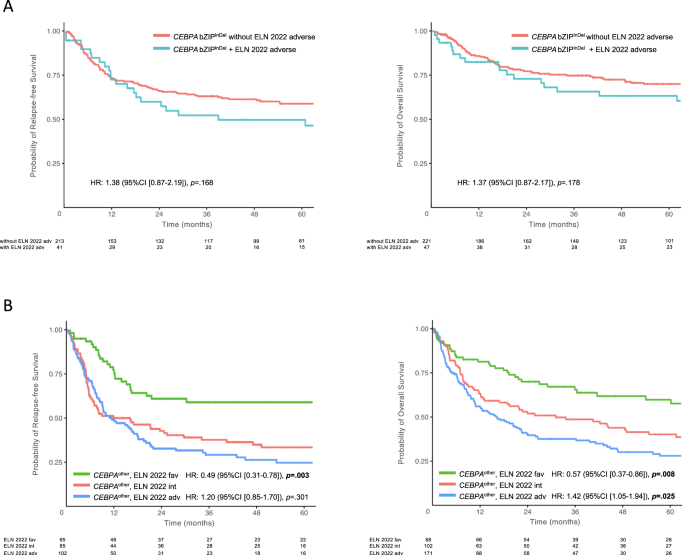
To investigate whether cytogenetic and molecular risk factors according to the novel ELN2022 recommendations had an effect on outcome in bZIPInDel patients, further outcome analysis was performed within this subgroup. In total, 48 patients harbored a combination of CEBPA bZIPInDel mutations and adverse cytogenetic or molecular abnormalities, including complex karyotypes (n = 9), monosomy 5 or 7 (n = 6) or mutations in ASXL1 (n = 5), EZH2 (n = 8), RUNX1 (n = 6), SF3B1 (n = 1), SRSF2 (n = 1), STAG2 (n = 8), TP53 (n = 2) and U2AF1 (n = 2). As depicted in Fig. 6A, the presence of ELN2022 adverse genetic factors did not significantly affect the favorable outcome in CEBPA bZIPInDel-mutant patients (RFS: HR 1.37 [0.87–2.17], p = 0.178 and OS: HR 1.46 [0.9–2.37], p = 0.129) (Tables S14, S15).
Impact of ELN2022 mutational subgroups in CEBPA mutant patients without bZIPInDel (subgroups 2–4 and 6–8)
Recent analysis suggested that certain co-mutations, in particular mutant NPM1, might have an effect in patients with smCEBPA [29]. To gain further insights on the impact of co-mutations in patients without bZIPInDel mutations, i.e. groups 2–4 and 6–8 (CEBPAother), a combined analysis based on the ELN2022 guidelines was performed for these patients. A total of 345 patients (63.8%) had sufficient cytogenetic and molecular data to allow reclassification according to the ELN2022 risk groups. Most of these patients (n = 183/345) (53%) were assigned to the adverse risk group, predominantly due to ASXL1 (n = 85), RUNX1 (n = 63), SRSF2 (n = 56) and STAG2 (n = 55) mutations, while only 27 patients (14.8%) had poor risk cytogenetics.
In the patients re-assigned to the favorable risk group (n = 88), all but one patient (showing a t(8;21)) had NPM1-mutations. As depicted in Fig. 6B, the outcome of these groups showed statistically significant differences, with the median RFS not reached and a median OS of 154 months in the CEBPAother/ELN2022 favorable risk group compared to 16 months and 31 months in the CEBPAother/ELN2022 intermediate risk group and 12 months and 16 months in the CEBPAother/ELN2022 adverse risk group (p < 0.001; for multivariate analysis see Tables S16 and S17).
Discussion
This study examined the prognostic impact of different CEBPA mutational subgroups in detail in a large cohort of patients. The prevalence of mutations in CEBPA ranges between 5 and 10 [8], therefore the 1010 CEBPA-mutant patients investigated in this study correspond to a total of 10.000–20.000 adult patients with AML. Based on these large numbers, this analysis for the first time allowed us to address several questions which had remained unclear or controversial in previous investigations, namely the impact of missense mutations in the bZIP region, the impact of co-mutations and cytogenetics in patients with bZIPInDel mutations as well as the prognosis of CEBPA mutant patients without CEBPA bZIPInDel mutations.
An important conclusion from our study is that it clearly supports the previous findings on the unique behavior of bZIP mutations compared to other types of mutations in CEBPA. The results confirm that irrespective of the allelic state, CEBPA bZIPInDel mutations define a distinct subgroup characterized by younger age and a specific co-mutational profile, including a high rate of GATA2 and WT1 mutations and mutual exclusiveness of other subtype-defining lesions like mutations in NPM1. In addition, CEBPA bZIPInDel mutations were associated with a very favorable outcome. Patients with bZIPInDel mutations, especially those with double mutant CEBPA, demonstrated a 5-year overall survival rate above 75%, indicating that these patients should be considered as one of the AML subgroups with the best response to conventional treatment. In line with this, alloHCT performed in CR1 did not improve outcome in this group. However, relapsed bZIPInDel patients appear to benefit from alloHCT as part of salvage treatment which is most strikingly demonstrated in the small subgroup of patients with CEBPA bZIPInDel mutations and CSF3R co-mutations, which showed a highly significantly decreased RFS, but no difference in OS due to successful salvage treatment. These results confirm similar observations in pediatric patients [15]. Co-mutations in several other genes have been associated with prognosis in CEBPA-mutant AML. In particular, GATA2 co-mutations were reported to confer a better prognosis in dmCEBPA patients by some groups [23, 30], although this was not confirmed by others [22, 24]. The data presented here provide a possible explanation for these discrepant reports. GATA2 mutations were predominantly found in patients with bZIPInDel mutations, with a significantly lower prevalence in patients with other dmCEBPA mutations (Gr.2–4). Given that GATA2 mutations had no impact on outcome when analysis was restricted to bZIPInDel patients (Fig. S5), this suggests that GATA2-mutation status might be a surrogate parameter for bZIPInDel-mutations, and therefore associated with better outcome in some studies. The same might be true for several other mutations, i.e., FLT3 and TET2, which have a significantly higher prevalence in non-bZIPInDel patients and had previously been shown to be associated with inferior outcome in CEBPA-mutant patients in some studies [23, 28, 31,32,33]. Very recently Tet2-mutations have been demonstrated to enhance aggressiveness of Cebpa-mutant disease in animal models [34]. Although significant differences were observed for mutant TET2 and FLT3 in univariate analysis, multivariable analysis did not confirm an independent effect of these alterations in bZIPInDel patients. In addition, a combined analysis of adverse molecular and cytogenetic prognostic factors according to ELN 2022 recommendations failed to indicate a significant prognostic impact in the bZIPInDel group. Taken together, these data suggest that CEBPA bZIPInDel-mutant patients represent a unique subgroup of patients with AML.
In contrast, patients with bZIPSTOP, bZIPms or TAD mutations, irrespective of allelic status, showed a different biology and worse outcome. In our previous analysis, bZIPms mutations were grouped and analyzed together with bZIPInDel as “bZIP in-frame”, which corresponds to the definitions of the current ELN and ICC guidelines [18, 19]. However, in the current analysis of a larger cohort of patients, bZIPms mutations were clearly associated with an inferior outcome when evaluated separately, and were clinically and molecularly distinct from bZIPInDel mutations, while sharing more commonality with the other CEBPA subgroups.
WHO continues to include biallelic CEBPA mutations as a defined subgroup [20]. However, our results indicate that patients with dmCEBPA without bZIPInDel mutations, i.e. patients showing either two TAD mutations, TAD and bZIPSTOP or TAD and bZIPms, differ substantially in biology and outcome, suggesting that only bZIPInDel mutations and not bZIPms mutations or any other dmCEBPA mutations should be included in this specific AML subgroup (Fig. 7). This extends previously published data by El-Sharkawi et al. which already provided evidence for a differential effect of different double mutant constellations [35].
Interestingly, our data indicate that the different CEBPA bZIP mutational subtypes, i.e. in-frame InDel mutations, InDel mutations inducing frameshift and missense mutations are distributed in a non-random way in the bZIP region, raising the possibility that mutation location impacts on the functional consequences. As illustrated in Fig. 1, CEBPAInDel mutations significantly clustered in the first part of the leucine zipper (between AA310-317) as well as the fork region (AA303-309), whereas missense mutations were significantly more common in the basic region (especially in several highly conserved amino acids, e.g. R300 and R297), which are directly involved in DNA-binding (reviewed in [36]). In contrast, the fork or hinge-region of bZIP-proteins is considered to be especially important for the spacing of the two alpha-helices of the leucine zipper, which in turn could influence either binding specificity and/or affinity of the DNA-binding [21] as well as the interaction with other proteins. CEBP proteins bind DNA as homo and heterodimers, and the CEBPA-interactome appears to be complex and still not completely understood [37].
Even though the presence of adverse risk aberrations according to the ELN 2022 recommendations was rare in CEBPA bZIPInDel mutant patients, knowledge of their prognostic implications is crucial for the choice of post-remission treatment, as it might abrogate the presumed prognostic advantage and low risk of relapse in these AML patients. In this analysis, patients with CEBPA bZIPInDel demonstrated superior survival irrespective of concurrent high-risk features. However, it is important to note that the impact of adverse genetic factors according to ELN 2022 in CEBPA mutant AML may vary depending on the specific chromosomal or molecular abnormalities.
Aside from CEBPA bZIPInDel, the other mutational subgroups do not appear to have an independent prognostic value. Analysis based on concomitant cytogenetic and molecular alterations according to current ELN 2022 recommendations within the CEBPAother patient group showed that they conformed to the expected risk stratification group, with no evidence that the CEBPA mutation had substantially changed the outcome. For example, a more favorable outcome in these groups was usually attributable to a concomitant NPM1 mutation.
Although our study represents the largest cohort of CEBPA-mutant AML, the analysis also has some limitations, in particular the retrospective nature of the analysis covering a period of almost three decades in which patients were treated. Consequently, none of the patients included were treated with novel targeted agents, e.g. tyrosine kinase inhibitors or Venetoclax/HMA-based therapies, which might affect outcome, at least in subgroups, and the impact of such agents on the different subgroups will be an important issue for future analyses.
In conclusion, CEBPA bZIPInDel-mutant AML represent a subset of AML with profoundly distinct disease biology and clinical outcomes. Further research efforts aimed at elucidating the underlying molecular mechanisms and identifying additional genetic and epigenetic alterations that interact with CEBPA mutations are necessary to harness the full potential of CEBPA bZIPInDel mutations in improving the management and prognosis of these AML patients.
Data availability
The datasets generated during and/or analyzed during the current study are available from the corresponding author on reasonable request.
References
Pabst T, Mueller BU, Zhang P, Radomska HS, Narravula S, Schnittger S, et al. Dominant-negative mutations of CEBPA, encoding CCAAT/enhancer binding protein-alpha (C/EBPalpha), in acute myeloid leukemia. Nat Genet. 2001;27:263–70.
Wilhelmson AS, Porse BT. CCAAT enhancer binding protein alpha (CEBPA) biallelic acute myeloid leukaemia: cooperating lesions, molecular mechanisms and clinical relevance. Br J Haematol. 2020;190:495–507.
Preudhomme C, Sagot C, Boissel N, Cayuela JM, Tigaud I, de Botton S, et al. Favorable prognostic significance of CEBPA mutations in patients with de novo acute myeloid leukemia: a study from the Acute Leukemia French Association (ALFA). Blood. 2002;100:2717–23.
Barjesteh van Waalwijk van Doorn-Khosrovani S, Erpelinck C, Meijer J, van Oosterhoud S, van Putten WL, et al. Biallelic mutations in the CEBPA gene and low CEBPA expression levels as prognostic markers in intermediate-risk AML. Hematol J. 2003;4:31–40.
Fröhling S, Schlenk RF, Stolze I, Bihlmayr J, Benner A, Kreitmeier S, et al. CEBPA mutations in younger adults with acute myeloid leukemia and normal cytogenetics: prognostic relevance and analysis of cooperating mutations. J Clin Oncol. 2004;22:624–33.
Bienz M, Ludwig M, Leibundgut EO, Mueller BU, Ratschiller D, Solenthaler M, et al. Risk assessment in patients with acute myeloid leukemia and a normal karyotype. Clin Cancer Res. 2005;11:1416–24.
Schlenk RF, Döhner K, Krauter J, Fröhling S, Corbacioglu A, Bullinger L, et al. Mutations and treatment outcome in cytogenetically normal acute myeloid leukemia. N Engl J Med. 2008;358:1909–18.
Pabst T, Mueller BU. Complexity of CEBPA dysregulation in human acute myeloid leukemia. Clin Cancer Res. 2009;15:5303–7.
Wouters BJ, Löwenberg B, Erpelinck-Verschueren CA, van Putten WL, Valk PJ, Delwel R. Double CEBPA mutations, but not single CEBPA mutations, define a subgroup of acute myeloid leukemia with a distinctive gene expression profile that is uniquely associated with a favorable outcome. Blood. 2009;113:3088–91.
Dufour A, Schneider F, Metzeler KH, Hoster E, Schneider S, Zellmeier E, et al. Acute myeloid leukemia with biallelic CEBPA gene mutations and normal karyotype represents a distinct genetic entity associated with a favorable clinical outcome. J Clin Oncol. 2010;28:570–7.
Green CL, Koo KK, Hills RK, Burnett AK, Linch DC, Gale RE. Prognostic significance of CEBPA mutations in a large cohort of younger adult patients with acute myeloid leukemia: impact of double CEBPA mutations and the interaction with FLT3 and NPM1 mutations. J Clin Oncol. 2010;28:2739–47.
Rockova V, Abbas S, Wouters BJ, Erpelinck CA, Beverloo HB, Delwel R, et al. Risk stratification of intermediate-risk acute myeloid leukemia: integrative analysis of a multitude of gene mutation and gene expression markers. Blood. 2011;118:1069–76.
Taskesen E, Bullinger L, Corbacioglu A, Sanders MA, Erpelinck CA, Wouters BJ, et al. Prognostic impact, concurrent genetic mutations, and gene expression features of AML with CEBPA mutations in a cohort of 1182 cytogenetically normal AML patients: further evidence for CEBPA double mutant AML as a distinctive disease entity. Blood. 2011;117:2469–75.
Fasan A, Haferlach C, Alpermann T, Jeromin S, Grossmann V, Eder C, et al. The role of different genetic subtypes of CEBPA mutated AML. Leukemia. 2014;28:794–803.
Tarlock K, Lamble A, Wang J, Gerbing RB, Ries RE, Loken MR, et al. CEBPA bZip mutations are associated with favorable prognosis in de novo AML: a report from the children’s oncology group. Blood. 2021;138:1137–47.
Taube F, Georgi JA, Kramer M, Stasik S, Middeke JM, Röllig C, et al. CEBPA mutations in 4708 patients with acute myeloid leukemia: differential impact of bZIP and TAD mutations on outcome. Blood. 2022;139:87–103.
Wakita S, Sakaguchi M, Oh I, Kako S, Toya T, Najima Y, et al. Prognostic impact of CEBPA bZIP domain mutation in acute myeloid leukemia. Blood Adv. 2022;6:238–47.
Döhner H, Wei AH, Appelbaum FR, Craddock C, DiNardo CD, Dombret H, et al. Diagnosis and management of AML in adults: 2022 recommendations from an international expert panel on behalf of the ELN. Blood. 2022;140:1345–77.
Arber DA, Orazi A, Hasserjian RP, Borowitz MJ, Calvo KR, Kvasnicka HM, et al. International consensus classification of myeloid neoplasms and acute leukemias: integrating morphologic, clinical, and genomic data. Blood. 2022;140:1200–28.
Khoury JD, Solary E, Abla O, Akkari Y, Alaggio R, Apperley JF, et al. The 5th edition of the World Health Organization Classification of haematolymphoid tumours: myeloid and histiocytic/dendritic neoplasms. Leukemia. 2022;36:1703–19.
Johnson PF. Identification of C/EBP basic region residues involved in DNA sequence recognition and half-site spacing preference. Mol Cell Biol. 1993;13:6919–30.
Greif PA, Dufour A, Konstandin NP, Ksienzyk B, Zellmeier E, Tizazu B, et al. GATA2 zinc finger 1 mutations associated with biallelic CEBPA mutations define a unique genetic entity of acute myeloid leukemia. Blood. 2012;120:395–403.
Grossmann V, Haferlach C, Nadarajah N, Fasan A, Weissmann S, Roller A, et al. CEBPA double-mutated acute myeloid leukaemia harbours concomitant molecular mutations in 76·8% of cases with TET2 and GATA2 alterations impacting prognosis. Br J Haematol. 2013;161:649–58.
Marceau-Renaut A, Guihard S, Castaigne S, Dombret H, Preudhomme C, Cheok M. Classification of CEBPA mutated acute myeloid leukemia by GATA2 mutations. Am J Hematol. 2015;90:E93–4.
Su L, Gao S, Tan Y, Lin H, Liu X, Liu S, et al. CSF3R mutations were associated with an unfavorable prognosis in patients with acute myeloid leukemia with CEBPA double mutations. Ann Hematol. 2019;98:1641–6.
Zhang Y, Wang F, Chen X, Zhang Y, Wang M, Liu H, et al. Companion gene mutations and their clinical significance in AML with double mutant CEBPA. Cancer Gene Ther. 2020;27:599–606.
He J, Liu J, Shen H, Wang Z, Cao L, Wu P, et al. Companion gene mutations and their clinical significance in AML with double or single mutant CEBPA. Int J Hematol. 2022;116:71–80.
Wang T, Hua H, Wang Z, Wang B, Cao L, Qin W, et al. Frequency and clinical impact of WT1 mutations in the context of CEBPA-mutated acute myeloid leukemia. Hematology. 2022;27:994–1002.
Dufour A, Schneider F, Hoster E, Benthaus T, Ksienzyk B, Schneider S, et al. Monoallelic CEBPA mutations in normal karyotype acute myeloid leukemia: independent favorable prognostic factor within NPM1 mutated patients. Ann Hematol. 2012;91:1051–63.
Fasan A, Eder C, Haferlach C, Grossmann V, Kohlmann A, Dicker F, et al. GATA2 mutations are frequent in intermediate-risk karyotype AML with biallelic CEBPA mutations and are associated with favorable prognosis. Leukemia. 2013;27:482–5.
Konstandin NP, Pastore F, Herold T, Dufour A, Rothenberg-Thurley M, Hinrichsen T, et al. Genetic heterogeneity of cytogenetically normal AML with mutations of CEBPA. Blood Adv. 2018;2:2724–31.
Tien FM, Hou HA, Tang JL, Kuo YY, Chen CY, Tsai CH, et al. Concomitant WT1 mutations predict poor prognosis in acute myeloid leukemia patients with double mutant CEBPA. Haematologica. 2018;103:e510–e3.
Wang H, Chu TT, Han SY, Qi JQ, Tang YQ, Qiu HY, et al. FLT3-ITD and CEBPA mutations predict prognosis in acute myelogenous leukemia irrespective of hematopoietic stem cell transplantation. Biol Blood Marrow Transpl. 2019;25:941–8.
Heyes E, Wilhelmson AS, Wenzel A, Manhart G, Eder T, Schuster MB, et al. TET2 lesions enhance the aggressiveness of CEBPA-mutant acute myeloid leukemia by rebalancing GATA2 expression. Nat Commun. 2023;14:6185.
El-Sharkawi D, Sproul D, Allen CG, Feber A, Wright M, Hills RK, et al. Variable outcome and methylation status according to CEBPA mutant type in double-mutated acute myeloid leukemia patients and the possible implications for treatment. Haematologica. 2018;103:91–100.
Miller M. The importance of being flexible: the case of basic region leucine zipper transcriptional regulators. Curr Protein Pept Sci. 2009;10:244–69.
Ramberger E, Sapozhnikova V, Kowenz-Leutz E, Zimmermann K, Nicot N, Nazarov PV, et al. PRISMA and BioID disclose a motifs-based interactome of the intrinsically disordered transcription factor C/EBPα. iScience. 2021;24:102686.
Acknowledgements
The authors thank all centers and participating physicians of the SAL, MLL, HOVON, MRC, AMLCG and ALFA study groups who entered their patients into the studies or registries.
Funding
Open Access funding enabled and organized by Projekt DEAL.
Author information
Authors and Affiliations
Contributions
Contribution: JAG, and CT designed the study; performed the research; assembled, analyzed, interpreted the data; and wrote the manuscript; SS and MM contributed molecular analyses and performed data analysis; M.K. performed the statistical analyses; CR, TH, PV, DL, TH, ND, FT, JMM, UP, HS, CB, CMT, CH, SK, WEB, BJW, UK, JB, WH, KS, ELB, RKH, AB, GE, KM, MRT, AD, HD, CP, LF, MB and RG contributed clinical and molecular data; and all authors approved the final version of the manuscript.
Corresponding author
Ethics declarations
Competing interests
C.T. is CEO and co-owner of AgenDix GmbH; all other authors have nothing to disclose.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Georgi, JA., Stasik, S., Kramer, M. et al. Prognostic impact of CEBPA mutational subgroups in adult AML. Leukemia 38, 281–290 (2024). https://doi.org/10.1038/s41375-024-02140-x
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41375-024-02140-x
This article is cited by
-
CEBPA bZIP in-frame mutations in acute myeloid leukemia: prognostic and therapeutic implications
Blood Cancer Journal (2024)