Abstract
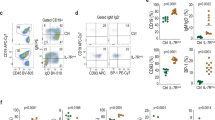
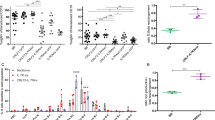
Philadelphia chromosome-like acute lymphoblastic leukemia (Ph-like ALL) is a high-risk subtype of B-ALL often associated with genetic variants that alter cytokine receptor signaling, including mutations in the interleukin-7 receptor (IL7R). To investigate whether IL7R variants are leukemia-initiating, we built mouse models expressing activated Il7r (aIL7R). B-cell intrinsic aIL7R mice developed spontaneous B-ALL, demonstrating sufficiency of Il7r activating mutations in leukemogenesis. Concomitant introduction of a knock-out allele in the associated adapter protein Lnk (encoded by Sh2b3) or a dominant-negative variant of the transcription factor Ikaros (Ikzf1) increased disease penetrance. The resulting murine leukemias displayed monoclonality and recurrent somatic Kras mutations and efficiently engrafted into immunocompetent mice. Phosphoproteomic analyses of aIL7R leukemic cells revealed constitutive Stat5 signaling and B cell receptor (BCR)-like signaling despite the absence of surface pre-BCR. Finally, in vitro treatment of aIL7R leukemic B-cells with Jak, mTOR, or Syk inhibitors blocked growth, confirming that each pathway is active in this mouse model of IL7R-driven B-ALL.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout







Similar content being viewed by others
Data availability
The mapped reads for the whole-exome sequencing have been deposited in the Sequence Read Archive database (BioProject ID: PRJNA735253). The mass spectrometry proteomics data have been deposited to the ProteomeXchange Consortium via the PRIDE [63] partner repository with the dataset identifiers PXD026438 and PXD026322.
References
Tasian SK, Loh ML, Hunger SP. Philadelphia chromosome-like acute lymphoblastic leukemia. Blood. 2017;130:2064–72.
Den Boer ML, van Slegtenhorst M, De Menezes RX, Cheok MH, Buijs-Gladdines J, Peters S, et al. A subtype of childhood acute lymphoblastic leukaemia with poor treatment outcome: a genome-wide classification study. Lancet Oncol. 2009;10:125–34.
Roberts KG, Li Y, Payne-Turner D, Harvey RC, Yang YL, Pei D, et al. Targetable kinase-activating lesions in Ph-like acute lymphoblastic leukemia. N Engl J Med. 2014;371:1005–15.
Roberts KG, Morin RD, Zhang JH, Hirst M, Zhao YJ, Su XP, et al. Genetic alterations activating kinase and cytokine receptor signaling in high-risk acute lymphoblastic leukemia. Cancer Cell. 2012;22:153–66.
Mullighan CG. The genomic landscape of acute lymphoblastic leukemia in children and young adults. Hematology-Am Soc Hematol Educ Program. 2014;2014:174–80.
Zenatti PP, Ribeiro D, Li WQ, Zuurbier L, Silva MC, Paganin M, et al. Oncogenic IL7R gain-of-function mutations in childhood T-cell acute lymphoblastic leukemia. Nat Genet. 2011;43:932–U931.
Shochat C, Tal N, Bandapalli OR, Palmi C, Ganmore I, Kronnie GT, et al. Gain-of-function mutations in interleukin-7 receptor-alpha (IL7R) in childhood acute lymphoblastic leukemias. J Exp Med. 2011;208:901–8.
Clark MR, Mandal M, Ochiai K, Singh H. Orchestrating B cell lymphopoiesis through interplay of IL-7 receptor and pre-B cell receptor signalling. Nat Rev Immunol. 2014;14:69–80.
Corfe SA, Paige CJ. The many roles of IL-7 in B cell development; Mediator of survival, proliferation and differentiation. Semin Immunol. 2012;24:198–208.
Barata JT, Durum SK, Seddon B. Flip the coin: IL-7 and IL-7R in health and disease. Nat Immunol. 2019;20:1584–93.
Ochiai K, Maienschein-Cline M, Mandal M, Triggs JR, Bertolino E, Sciammas R, et al. A self-reinforcing regulatory network triggered by limiting IL-7 activates pre-BCR signaling and differentiation. Nat Immunol. 2012;13:300–U124.
Buchner M, Swaminathan S, Chen ZS, Muschen M. Mechanisms of pre-B-cell receptor checkpoint control and its oncogenic subversion in acute lymphoblastic leukemia. Immunol Rev. 2015;263:192–209.
Treanor LM, Zhou S, Janke L, Churchman ML, Ma ZJ, Lu TH, et al. Interleukin-7 receptor mutants initiate early T cell precursor leukemia in murine thymocyte progenitors with multipotent potential. J Exp Med. 2014;211:701–13.
Yokoyama K, Yokoyama N, Izawa K, Kotani A, Harashima A, Hozumi K, et al. In vivo leukemogenic potential of an interleukin 7 receptor alpha chain mutant in hematopoietic stem and progenitor cells. Blood. 2013;122:4259–63.
Takaki S, Sauer K, Iritani BM, Chien S, Ebihara Y, Tsuji K, et al. Control of B cell production by the adaptor protein Lnk: definition of a conserved family of signal-modulating proteins. Immunity. 2000;13:599–609.
Cheng Y, Chikwava K, Wu C, Zhang HB, Bhagat A, Pei DH, et al. LNK/SH2B3 regulates IL-7 receptor signaling in normal and malignant B-progenitors. J Clin Investig. 2016;126:1267–81.
Wang JH, Nichogiannopoulou A, Wu L, Sun L, Sharpe AH, Bigby M, et al. Selective defects in the development of the fetal and adult lymphoid system in mice with an Ikaros null mutation. Immunity. 1996;5:537–49.
Georgopoulos K, Bigby M, Wang JH, Molnar A, Wu P, Winandy S, et al. The IKAROS gene is required for the development of all lymphoid lineages. Cell. 1994;79:143–56.
Yoshida T, Ng SYM, Zuniga-Pflucker JC, Georgopoulos K. Early hematopoietic lineage restrictions directed by Ikaros. Nat Immunol. 2006;7:382–91.
Joshi I, Yoshida T, Jena N, Qi XQ, Zhang JW, Van Etten RA, et al. Loss of Ikaros DNA-binding function confers integrin-dependent survival on pre-B cells and progression to acute lymphoblastic leukemia. Nat Immunol. 2014;15:294–304.
Schwickert TA, Tagoh H, Gultekin S, Dakic A, Axelsson E, Minnich M, et al. Stage-specific control of early B cell development by the transcription factor Ikaros. Nat Immunol. 2014;15:283–93.
Katerndahl CDS, Heltemes-Harris LM, Willette MJL, Henzler CM, Frietze S, Yang RD. et al. Antagonism of B cell enhancer networks by STAT5 drives leukemia and poor patient survival. Nat Immunol. 2017;18:694
Ge Z, Gu Y, Xiao LC, Han Q, Li JY, Chen BA, et al. Co-existence of IL7R high and SH2B3 low expression distinguishes a novel high-risk acute lymphoblastic leukemia with Ikaros dysfunction. Oncotarget. 2016;7:46014–27.
Allenspach E, Shubin N, Cerosaletti K, Mikacenic C, Gorman J, MacQuivey M, et al. The autoimmune risk R262W variant of the adaptor SH2B3 improves survival in sepsis. J Immunol [In preparation/revision] 2020.
Mullighan CG, Su XP, Zhang JH, Radtke I, Phillips LAA, Miller CB, et al. Deletion of IKZF1 and Prognosis in Acute Lymphoblastic Leukemia. N Engl J Med. 2009;360:470–80.
Meffre E, Fougereau M, Argenson JN, Aubaniac JM, Schiff C. Cell surface expression of surrogate light chain (Psi L) in the absence of mu on human pro-B cell lines and normal pro-B cells. Eur J Immunol. 1996;26:2172–80. Sep
Lemmers B, Arnoulet C, Fossat C, Chambost H, Sainty D, Gabert J, et al. Fine characterization of childhood and adult acute lymphoblastic leukemia (ALL) by a proB and preB surrogate light chain-specific mAb and a proposal for a new B cell ALL classification. Leukemia. 2000;14:2103–11.
Lemmers B, Gauthier L, Guelpa-Fonlupt V, Fougereau M, Schiff C. The human (Psi L+mu(-)) proB complex: Cell surface expression and biochemical structure of a putative transducing receptor. Blood. 1999;93:4336–46.
Goldberg L, Gough SM, Lee F, Dang C, Walker RL, Zhu YLJ, et al. Somatic mutations in murine models of leukemia and lymphoma: Disease specificity and clinical relevance. Genes Chromosomes Cancer. 2017;56:472–83.
Gough SM, Goldberg L, Pineda M, Walker RL, Zhu YJ, Bilke S, et al. Progenitor B-1 B-cell acute lymphoblastic leukemia is associated with collaborative mutations in 3 critical pathways. Blood Adv. 2017;1:1749–59.
Lu LW, Chaudhury P, Osmond DG. Regulation of cell survival during B lymphopoiesis: Apoptosis and Bcl-2/Bax content of precursor B cells in bone marrow of mice with altered expression of IL-7 and recombinase-activating gene-2. J Immunol. 1999;162:1931–40.
Malin S, McManus S, Cobaleda C, Novatchkova M, Delogu A, Bouillet P, et al. Role of STAT5 in controlling cell survival and immunoglobulin gene recombination during pro-B cell development. Nat Immunol. 2010;11:171–U197.
Grillot DAM, Merino R, Pena JC, Fanslow WC, Finkelman FD, Thompson CB, et al. bcl-x exhibits regulated expression during B cell development and activation and modulates lymphocyte survival in transgenic mice. J Exp Med. 1996;183:381–91.
Maude SL, Tasian SK, Vincent T, Hall JW, Sheen C, Roberts KG, et al. Targeting JAK1/2 and mTOR in murine xenograft models of Ph-like acute lymphoblastic leukemia. Blood. 2012;120:3510–8.
Tasian SK, Doral MY, Borowitz MJ, Wood BL, Chen IM, Harvey RC, et al. Aberrant STAT5 and PI3K/mTOR pathway signaling occurs in human CRLF2-rearranged B-precursor acute lymphoblastic leukemia. Blood. 2012;120:833–42.
Tasian SK, Teachey DT, Li Y, Shen F, Harvey RC, Chen IM, et al. Potent efficacy of combined PI3K/mTOR and JAK or ABL inhibition in murine xenograft models of Ph-like acute lymphoblastic leukemia. Blood. 2017;129:177–87.
Hurtz C, Wertheim GB, Loftus JP, Blumenthal D, Lehman A, Li Y, et al. Oncogene-independent BCR-like signaling adaptation confers drug resistance in Ph-like ALL. J Clin Investig. 2020;130:3637–53.
Cramer SD, Hixon JA, Andrews C, Porter RJ, Rodrigues GOL, Wu XL, et al. Mutant IL-7R alpha and mutant NRas are sufficient to induce murine T cell acute lymphoblastic leukemia. Leukemia. 2018;32:1795–9.
Geron I, Savino AMS, Tal N, Brown J, Turati V, James C, et al. An instructive role for IL7RA in the development of human B-cell precursor leukemia. BioRxiv; 2020. https://doi.org/10.1101/2020.01.27.919951.
Damia G, Komschlies KL, Faltynek CR, Ruscetti FW, Wiltrout RH. Administration of recombinant human interleukin-7 alters the frequency and number of myeloid progenitor cells in the bone-marrow and spleen of mice. Blood. 1992;79:1121–9.
Komschlies KL, Gregorio TA, Gruys ME, Back TC, Faltynek CR, Wiltrout RH. Administration of recombinant human IL-7 to mice alters the composition of b-lineage cells and t-cell subsets, enhances t-cell function, and induces regression of established metastases. J Immunol. 1994;152:5776–84.
Morrissey PJ, Conlon P, Braddy S, Williams DE, Namen AE, Mochizuki DY. Administration of IL-7 to mice with cyclophosphamide-induced lymphopenia accelerates lymphocyte repopulation. J Immunol. 1991;146:1547–52.
Digel W, Schmid M, Heil G, Conrad P, Gillis S, Porzsolt F. Human interleukin-7 induces proliferation of neoplastic-cells from chronic lymphocytic-leukemia and acute leukemias. Blood. 1991;78:753–9.
Martin-Lorenzo A, Hauer J, Vicente-Duenas C, Auer F, Gonzalez-Herrero I, Garcia-Ramirez I, et al. Infection exposure is a causal factor in B-cell precursor acute lymphoblastic leukemia as a result of Pax5-inherited susceptibility. Cancer Discov. 2015;5:1328–43.
Nakayama J, Yamamoto M, Hayashi K, Satoh H, Bundo K, Kubo M, et al. BLNK suppresses pre-B-cell leukemogenesis through inhibition of JAK3. Blood. 2009;113:1483–92.
Appelmann I, Rillahan CD, de Stanchina E, Carbonetti G, Chen C, Lowe SW, et al. Janus kinase inhibition by ruxolitinib extends dasatinib- and dexamethasone-induced remissions in a mouse model of Ph plus ALL. Blood. 2015;125:1444–51.
Abdelrasoul H, Vadakumchery A, Werner M, Lenk L, Khadour A, Young M, et al. Synergism between IL7R and CXCR4 drives BCR-ABL induced transformation in Philadelphia chromosome-positive acute lymphoblastic leukemia. Nature. Communications. 2020;11:12.
Mocsai A, Ruland J, Tybulewicz VLJ. The SYK tyrosine kinase: a crucial player in diverse biological functions. Nat Rev Immunol. 2010;10:387–402.
Loftus JP, Yahiaoui A, Brown PA, Niswander LM, Bagashev A, Wang M, et al. Combinatorial efficacy of entospletinib and chemotherapy in patient-derived xenograft models of infant acute lymphoblastic leukemia. Haematologica. 2021;106:1067–78.
Hobeika E, Thiemann S, Storch B, Jumaa H, Nielsen PJ, Pelanda R, et al. Testing gene function early in the B cell lineage in mb1-cre mice. Proc Natl Acad Sci USA. 2006;103:13789–94.
Rickert RC, Roes J, Rajewsky K. B lymphocyte-specific, Cre-mediated mutagenesis in mice. Nucleic Acids Res. 1997;25:1317–8.
Becker-Herman S, Meyer-Bahlburg A, Schwartz MA, Jackson SW, Hudkins KL, Liu CH, et al. WASp-deficient B cells play a critical, cell-intrinsic role in triggering autoimmunity. J Exp Med. 2011;208:2033–42.
Wray-Dutra MN, Chawla R, Thomas KR, Seymour BJ, Arkatkar T, Sommer KM, et al. Activated CARD11 accelerates germinal center kinetics, promoting mTORC1 and terminal differentiation. J Exp Med. 2018;215:2445–61.
Bandaranayake AD, Correnti C, Ryu BY, Brault M, Strong RK, Rawlings DJ. Daedalus: a robust, turnkey platform for rapid production of decigram quantities of active recombinant proteins in human cell lines using novel lentiviral vectors. Nucleic Acids Res. 2011;39:e143.
Schwartz MA, Kolhatkar NS, Thouvenel C, Khim S, Rawlings DJ. CD4(+) T cells and CD40 participate in selection and homeostasis of peripheral b cells. J Immunol. 2014;193:3492–502.
McKenna A, Hanna M, Banks E, Sivachenko A, Cibulskis K, Kernytsky A, et al. The Genome Analysis Toolkit: A MapReduce framework for analyzing next-generation DNA sequencing data. Genome Res. 2010;20:1297–303.
Dunn T, Berry G, Emig-Agius D, Jiang Y, Lei S, Iyer A, et al. Pisces: an accurate and versatile variant caller for somatic and germline next-generation sequencing data. Bioinformatics. 2019;35:1579–81.
Wang K, Li MY, Hakonarson H. ANNOVAR: functional annotation of genetic variants from high-throughput sequencing data. Nucleic Acids Res. 2010;38:7.
Hale M, Lee B, Honaker Y, Leung WH, Grier AE, Jacobs HM, et al. Homology-directed recombination for enhanced engineering of chimeric antigen receptor T Cells. Mol Ther Methods Clin Dev. 2017;4:192–203.
Eng JK, Jahan TA, Hoopmann MR. Comet: An open-source MS/MS sequence database search tool. Proteomics. 2013;13:22–24.
Deutsch EW, Mendoza L, Shteynberg D, Farrah T, Lam H, Tasman N, et al. A guided tour of the trans-proteomic pipeline. Proteomics. 2010;10:1150–9.
MacLean B, Tomazela DM, Shulman N, Chambers M, Finney GL, Frewen B, et al. Skyline: an open source document editor for creating and analyzing targeted proteomics experiments. Bioinformatics. 2010;26:966–8.
Perez-Riverol Y, Csordas A, Bai JW, Bernal-Llinares M, Hewapathirana S, Kundu DJ. et al. The PRIDE database and related tools and resources in 2019: improving support for quantification data. Nucleic Acids Res. 2019;47:D442–D450.
Acknowledgements
The authors would like to thank Karen Sommer and Andrea Repele for laboratory management, Jennifer Haddock for administrative support, Beth Lawlor for helpful comments and critical review of the manuscript, the University of Washington Histology and Imaging Core, and the Northwest Genomics Center at the University of Washington. This work was supported by the National Institutes of Health under award numbers: TL1-TR000422 (to KRT), DP3-DK111802 (to DJR), NCI-R01CA201135-A1 (to RGJ), 1K08CA184418 (to SKT), 1U01CA232486 (to SKT), 1U01CA243072 (SKT), 1K08DK114568-01 (to EJA), Department of Defense Translational Team Science Award CA180683P1 (to SKT), the V Foundation for Cancer Research (to SKT), the Seattle Children’s Research Institute (SCRI) Center for Immunity and Immunotherapies (CIIT) Program for Cell and Gene Therapy (PCGT), the Children’s Guild Association Endowed Chair in Pediatric Immunology (to DJR), and the Hansen Investigator in Pediatric Innovation Endowment (to DJR). The content is solely the responsibility of the authors and does not necessarily represent the official views of the National Institutes of Health.
Author information
Authors and Affiliations
Contributions
KRT, EJA, and NDC: designed and performed experiments, analyzed data, and wrote and/or edited the manuscript. KRT and EJA designed the aIL7R murine model. MNW, JPL, SK, and AZ: designed, performed, and interpreted the experiments. AET and HDL analyzed data. KG provided the Ikzf1 flox mice and edited the manuscript. SKT provided the PDX models, interpreted data, and edited the manuscript. RGJ and DJR conceived of and supervised the study, interpreted data, and edited the manuscript.
Corresponding authors
Ethics declarations
Competing interests
SKT receives research funding from Incyte Corporation, Gilead Sciences, and MacroGenics, Inc. for unrelated studies and is a member of the scientific advisory board of Aleta Biotherapeutics for unrelated studies.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
About this article
Cite this article
Thomas, K.R., Allenspach, E.J., Camp, N.D. et al. Activated interleukin-7 receptor signaling drives B-cell acute lymphoblastic leukemia in mice. Leukemia 36, 42–57 (2022). https://doi.org/10.1038/s41375-021-01326-x
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41375-021-01326-x
This article is cited by
-
Chemical genetic control of cytokine signaling in CAR-T cells using lenalidomide-controlled membrane-bound degradable IL-7
Leukemia (2024)
-
In Utero Development and Immunosurveillance of B Cell Acute Lymphoblastic Leukemia
Current Treatment Options in Oncology (2022)
-
An instructive role for Interleukin-7 receptor α in the development of human B-cell precursor leukemia
Nature Communications (2022)
-
Interleukin-7 receptor α mutational activation can initiate precursor B-cell acute lymphoblastic leukemia
Nature Communications (2021)