Abstract
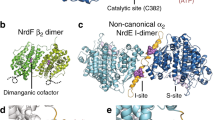
Ribonucleotide reductases (RNRs) catalyze the reduction of ribonucleotides into deoxyribonucleotides, which constitute the precursor pools used for DNA synthesis and repair. Imbalances in these pools increase mutational rates and are detrimental to the cell. Balanced precursor pools are maintained primarily through the regulation of the RNR substrate specificity. Here, the molecular mechanism of the allosteric substrate specificity regulation is revealed through the structures of a dimeric coenzyme B12–dependent RNR from Thermotoga maritima, both in complexes with four effector-substrate nucleotide pairs and in three complexes with only effector. The mechanism is based on the flexibility of loop 2, a key structural element, which forms a bridge between the specificity effector and substrate nucleotides. Substrate specificity is achieved as different effectors and their cognate substrates stabilize specific discrete loop 2 conformations. The mechanism of substrate specificity regulation is probably general for most class I and class II RNRs.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$189.00 per year
only $15.75 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
References
Jordan, A. & Reichard, P. Ribonucleotide reductases. Annu. Rev. Biochem. 67, 71–98 (1998).
Fontecave, M., Mulliez, E. & Logan, D.T. Deoxyribonucleotide synthesis in anaerobic microorganisms: the class III ribonucleotide reductase. Prog. Nucleic Acid Res. Mol. Biol. 72, 95–127 (2002).
Stubbe, J., Ator, M. & Krenitsky, T. Mechanism of ribonucleoside diphosphate reductase from Escherichia coli. Evidence for 3′-C–H bond cleavage. J. Biol. Chem. 258, 1625–1631 (1983).
Stubbe, J. Ribonucleotide reductases. Adv. Enzymol. Relat. Areas Mol. Biol. 63, 349–419 (1990).
Reichard, P. Ribonucleotide reductases: the evolution of allosteric regulation. Arch. Biochem. Biophys. 397, 149–155 (2002).
Kunz, B.A. et al. International Commission for Protection Against Environmental Mutagens and Carcinogens. Deoxyribonucleoside triphosphate levels: a critical factor in the maintenance of genetic stability. Mutat. Res. 318, 1–64 (1994).
Brown, N.C. & Reichard, P. Role of effector binding in allosteric control of ribonucleoside diphosphate reductase. J. Mol. Biol. 46, 39–55 (1969).
Eliasson, R., Pontis, E., Jordan, A. & Reichard, P. Allosteric control of three B12-dependent (class II) ribonucleotide reductases. Implications for the evolution of ribonucleotide reduction. J. Biol. Chem. 274, 7182–7189 (1999).
Andersson, J., Westman, M., Hofer, A. & Sjoberg, B.-M. Allosteric regulation of the class III anaerobic ribonucleotide reductase from bacteriophage T4. J. Biol. Chem. 275, 19443–19448 (2000).
Larsson, K.M., Andersson, J., Sjoberg, B.M., Nordlund, P. & Logan, D.T. Structural basis for allosteric substrate specificity regulation in anaerobic ribonucleotide reductases. Structure 9, 739–750 (2001).
Beck, W.S. Regulation of cobamide-dependent ribonucleotide reductase by allosteric effectors and divalent cations. J. Biol. Chem. 242, 3148–3158 (1967).
Panagou, D., Orr, M.D., Dunstone, J.R. & Blakley, R.L. A monomeric, allosteric enzyme with a single polypeptide chain. Ribonucleotide reductase from Lactobacillus leichmannii. Biochemistry 11, 2378–2388 (1972).
Chen, A.K., Bhan, A., Hopper, S., Abrams, R. & Franzen, J.S. Substrate and effector binding to ribonucleoside triphosphate reductase of Lactobacillus leichmannii. Biochemistry 13, 654–661 (1974).
Jordan, A. et al. B12-dependent ribonucleotide reductases from deeply rooted eubacteria are structurally related to the aerobic enzyme from Escherichia coli. Proc. Natl. Acad. Sci. USA 94, 13487–13492 (1997).
Eriksson, M. et al. Binding of allosteric effectors to ribonucleotide reductase protein R1: reduction of active-site cysteines promotes substrate binding. Structure 5, 1077–1092 (1997).
Uppsten, M. et al. Structure of the large subunit of class Ib ribonucleotide reductase from Salmonella typhimurium and its complexes with allosteric effectors. J. Mol. Biol. 330, 87–97 (2003).
Logan, D.T., Andersson, J., Sjöberg, B.M. & Nordlund, P. A glycyl radical site in the crystal structure of a class III ribonucleotide reductase. Science 283, 1499–1504 (1999).
Sintchak, M.D., Arjara, G., Kellogg, B.A., Stubbe, J. & Drennan, C.L. The crystal structure of class II ribonucleotide reductase reveals how an allosterically regulated monomer mimics a dimer. Nat. Struct. Biol. 9, 293–300 (2002).
Uhlin, U. & Eklund, H. Structure of ribonucleotide reductase protein R1. Nature 370, 533–539 (1994).
Becker, A. et al. Structure and mechanism of the glycyl radical enzyme pyruvate formate-lyase. Nat. Struct. Biol. 6, 969–975 (1999).
Eklund, H. & Fontecave, M. Glycyl radical enzymes: a conservative structural basis for radicals. Structure Fold Des. 7, R257–R262 (1999).
Himo, F. & Siegbahn, P.E. Quantum chemical studies of radical-containing enzymes. Chem. Rev. 103, 2421–2456 (2003).
Chimploy, K. & Mathews, C.K. Mouse ribonucleotide reductase control: influence of substrate binding upon interactions with allosteric effectors. J. Biol. Chem. 276, 7093–7100 (2001).
Eriksson, S. Direct photoaffinity labeling of ribonucleotide reductase from Escherichia coli. Evidence for enhanced binding of the allosteric effector dTTP by the presence of substrates. J. Biol. Chem. 258, 5674–5678 (1983).
Kabsch, W. Automatic processing of rotation diffraction data from crystals of initially unknown symmetry and cell constants. J. Appl. Cryst. 26, 795–800 (1993).
Otwinowski, Z. & Minor, W. Processing of X-ray diffraction data collected in oscillation mode. Methods Enzymol. 276, 307–326 (1997).
Collaborative Computational Project, Number 4. The CCP4 suite: programs for protein crystallography. Acta Crystallogr. D 50, 760–763 (1994).
Potterton, E., Briggs, P., Turkenburg, M. & Dodson, E. A graphical user interface to the CCP4 program suite. Acta Crystallogr. D 59, 1131–1137 (2003).
Brünger, A.T. et al. Crystallography & NMR system: a new software suite for macromolecular structure determination. Acta Crystallogr. D 54, 905–921 (1998).
Terwilliger, T.C. Maximum-likelihood density modification. Acta Crystallogr. D 56, 965–972 (2000).
Kleywegt, G.J. & Jones, A. Databases in protein crystallography. Acta Crystallogr. D 54, 1119–1131 (1998).
Murshudov, G.N., Vagin, A.A. & Dodson, E.E. Refinement of macromolecular structures by the maximum-likelihood method. Acta Crystallogr. D 54, 240–255 (1997).
van Aalten, D.M. et al. PRODRG, a program for generating molecular topologies and unique molecular descriptors from coordinates of small molecules. J. Comput. Aided Mol. Des. 10, 255–262 (1996).
Perrakis, A., Sixma, T.K., Wilson, K.S. & Lamzin, V.S. wARP: improvement and extension of crystallographic phases by weighted averaging of multiple refined dummy atomic models. Acta Crystallogr. D 53, 448–455 (1997)
Kraulis, P.J. MOLSCRIPT: a program to produce both detailed and schematic plots of protein structures. J. Appl. Crystallogr. 24, 946–950 (1991).
Esnouf, R.M. An extensively modified version of MolScript that includes greatly enhanced coloring capabilities. J. Mol. Graph. Model. 15, 132–4, 112-113 (1997).
Merritt, E.A. & Murphy, M.E.P. Raster3D version 2.0: a program for photorealistic molecular graphics. Acta Crystallogr. D 50, 869–873 (1994).
Acknowledgements
We thank staff at beamline I711 of MAX-lab (Y. Cerenius) and at the European Molecular Biology Laboratory Hamburg outstation (C. Enroth) for technical help with data collection. This work was supported by grants from the Swedish Research Council (to D.L. and P.N.) and Karolinska Institutet (to P.R.).
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Rights and permissions
About this article
Cite this article
Larsson, KM., Jordan, A., Eliasson, R. et al. Structural mechanism of allosteric substrate specificity regulation in a ribonucleotide reductase. Nat Struct Mol Biol 11, 1142–1149 (2004). https://doi.org/10.1038/nsmb838
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nsmb838
This article is cited by
-
The translational repressor 4E-BP1 regulates RRM2 levels and functions as a tumor suppressor in Ewing sarcoma tumors
Oncogene (2021)
-
Insights into the metabolism pathway and functional genes of long-chain aliphatic alkane degradation in haloarchaea
Extremophiles (2020)
-
Class Id ribonucleotide reductase utilizes a Mn2(IV,III) cofactor and undergoes large conformational changes on metal loading
JBIC Journal of Biological Inorganic Chemistry (2019)
-
Regulation of nucleotide metabolism by mutant p53 contributes to its gain-of-function activities
Nature Communications (2015)
-
The Case for an Early Biological Origin of DNA
Journal of Molecular Evolution (2014)