Abstract
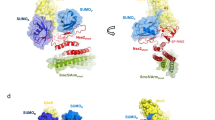
Yeast Hst2 (yHst2) is a member of the silencing information regulator 2 (Sir2) family of NAD+-dependent protein deacetylases that are implicated in transcriptional silencing, DNA repair, genome stability and longevity. The X-ray crystal structure of the full-length yHst2 protein reveals a central catalytic core domain fold that is characteristic of the other Sir2 homologs, and C- and N-terminal extensions that interact with the NAD+ and acetyl-lysine substrate-binding sites, respectively, suggesting an autoregulatory function for these domains. Moreover, the N-terminal extension mediates formation of a homotrimer within the crystal lattice. Enzymatic and sedimentation equilibrium studies using deletion constructs of yHst2 support the involvement of the N- and C-terminal yHst2 regions and trimer formation in catalysis by yHst2. Together, these studies indicate that the sequence-divergent N- and C-terminal regions of the eukaryotic Sir2 proteins may have a particularly important role in their distinct substrate-binding properties, biological activities or both.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$189.00 per year
only $15.75 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
Accession codes
References
de Ruijter, A.J., van Gennip, A.H., Caron, H.N., Kemp, S. & van Kuilenburg, A.B. Histone deacetylases (HDACs): characterization of the classical HDAC family. Biochem. J. 370, 737–749 (2003).
Timmermann, S. Histone acetylation and disease. Cell. Mol. Life Sci. 58, 728–736 (2001).
Guarente, L. Sir2 links chromatin silencing, metabolism, and aging. Genes Dev. 14, 1021–1026 (2000).
Marmorstein, R. Structure of histone deacetylases: insights into substrate recognition and catalysis. Structure 9, 1127–1133 (2001).
Frye, R.A. Phylogenetic classification of prokaryotic and eukaryotic Sir2-like proteins. Biochem. Biophys. Res. Commun. 273, 793–798 (2000).
Finnin, M.S. et al. Structures of a histone deacetylase homologue bound to the TSA and SAHA inhibitors. Nature 401, 188–193 (1999).
Landry, J. et al. The silencing protein SIR2 and its homologs are NAD-dependent protein deacetylases. Proc. Natl. Acad. Sci. USA 97, 5807–5811 (2000).
Frye, R.A. Characterization of five human cDNAs with homology to the yeast SIR2 gene: Sir2-like proteins (sirtuins) metabolize NAD and may have protein ADP-ribosyltransferase activity. Biochem. Biophys. Res. Commun. 260, 273–279 (1999).
Imai, S., Armstrong, C.M., Kaeberlein, M. & Guarente, L. Transcriptional silencing and longevity protein Sir2 is an NAD-dependent histone deacetylase. Nature 403, 795–800 (2000).
Denu, J.M. Linking chromatin function with metabolic networks: Sir2 family of NAD+-dependent deacetylases. Trends Biochem. Sci. 28, 41–48 (2003).
Smith, J.S. et al. A phylogenetically conserved NAD+-dependent protein deacetylase activity in the Sir2 protein family. Proc. Natl. Acad. Sci. USA 97, 6658–6663 (2000).
Min, J., Landry, J., Sternglanz, R. & Xu, R.M. Crystal structure of a SIR2 homolog–NAD complex. Cell 105, 269–279 (2001).
Avalos, J.L. et al. Structure of a Sir2 enzyme bound to an acetylated p53 peptide. Mol. Cell 10, 523–535 (2002).
Starai, V.J., Celic, I., Cole, R.N., Boeke, J.D. & Escalante-Semerena, J.C. Sir2-dependent activation of acetyl-CoA synthetase by deacetylation of active lysine. Science 298, 2390–2392 (2002).
Bell, S.D., Botting, C.H., Wardleworth, B.N., Jackson, S.P. & White, M.F. The interaction of Alba, a conserved archaeal chromatin protein, with Sir2 and its regulation by acetylation. Science 296, 148–151 (2002).
Vaziri, H. et al. hSIR2 (SIRT1) functions as an NAD-dependent p53 deacetylase. Cell 107, 149–159 (2001).
North, B.J., Marshall, B.L., Borra, M.T., Denu, J.M. & Verdin, E. The human Sir2 ortholog, SIRT2, is a NAD+-dependent tubulin deacetylase. Mol. Cell 11, 437–444 (2003).
Cockell, M.M., Perrod, S. & Gasser, S.M. Analysis of Sir2p domains required for rDNA and telomeric silencing in Saccharomyces cerevisiae. Genetics 154, 1069–1083 (2000).
Cuperus, G., Shafaatian, R. & Shore, D. Locus specificity determinants in the multifunctional yeast silencing protein Sir2. EMBO J. 19, 2641–2651 (2000).
Finnin, M.S., Donigian, J.R. & Pavletich, N.P. Structure of the histone deacetylase SIRT2. Nat. Struct. Biol. 8, 621–625 (2001).
Bellamacina, C.R. The nicotinamide dinucleotide binding motif: a comparison of nucleotide binding proteins. FASEB J. 10, 1257–1269 (1996).
Polevoda, B. & Sherman, F. Nα-terminal acetylation of eukaryotic proteins. J. Biol. Chem. 275, 36479–36482 (2000).
Hubbard, S.R. Autoinhibitory mechanisms in receptor tyrosine kinases. Frontiers Biosci. 7, 330–340 (2002).
Rojas, J.R. et al. Structure of the Tetrahymena GCN5 bound to coenzyme-A and a histone H3 peptide. Nature 401, 93–98 (1999).
Yan, Y., Barlev, N.A., Haley, R.H., Berger, S.L. & Marmorstein, R. Crystal structure of yeast Esa1 suggests a unified mechanism of catalysis and substrate binding by histone acetyltransferases. Mol. Cell 6, 1195–1205 (2000).
Dutnall, R.N., Tafrov, S.T., Sternglanz, R. & Ramakrishnan, V. Structure of the histone acetyltransferase Hat1: A paradigm for the GCN5-related N-acetyltransferase superfamily. Cell 94, 427–438 (1998).
Marmorstein, R. Structure of SET domain proteins—a new twist on histone methylation. Trends Biochem. Sci. 28, 59–62 (2003).
Xiao, B. et al. Structure and catalytic mechanism of human histone methyltransferase SET7/9. Nature 421, 652–656 (2003).
Min, J., Zhang, X., Cheng, X., Grewal, S.I.S. & Xu, R.-M. Structure of the SET domain histone lysine methyltransferase Clr4. Nat. Struct. Biol. 9, 828–832 (2002).
Trievel, R.C., Beach, B.M., Dirk, L.M.A., Houtz, R.L. & Hurley, J.H. Structure and catalytic mechanism of a SET domain protein methyltransferase. Cell 111, 91–103 (2002).
Jacobs, S.A. et al. The active site of the SET domain is constructed on a knot. Nat. Struct. Biol. 9, 833–838 (2002).
Brunger, A.T. et al. Crystallography & NMR system: a new software suite for macromolecular structure determination. Acta Crystallogr. D 54, 905–921 (1998).
Terwilliger, T.C. & Berendzen, J. Automated MAD and MIR structure solution. Acta Crystallogr. D 55, 849–861 (1999).
Schneider, T.R. & Sheldrick, G.M. Substructure solution with SHELXD. Acta Crystallogr. D 58, 1772–1779 (2002).
de La Fortelle, E. & Bricogne, G. SHARP: maximum-likelihood heavy-atom parameter refinement for multiple isomorphous replacement and multiwavelength anomalous diffraction methods. Methods Enzymol. 276, 472–494 (1997).
Jones, T.A. A graphics model building and refinement system for macromolecules. J. Appl. Crystallogr. 11, 268–272 (1978).
Johnson, M.L., Correia, J.J., Yphantis, D.A. & Halvorson, H.R. Analysis of data from the analytical ultracentrifuge by nonlinear least-squares techniques. Biophys. J. 36, 575–588 (1981).
Bitterman, K.J., Anderson, R.M., Cohen, H.Y., Latorre-Esteves, M. & Sinclair, D.A. Inhibition of silencing and accelerated aging by nicotinamide, a putative negative regulator of yeast sir2 and human SIRT1. J. Biol. Chem. 277, 45099–45107 (2002).
Acknowledgements
We thank L. Guarente, L. Pillus, D. Speicher and S. Harper for useful discussions, and A. Joachimiak, R. Zhang, N. Duke and the SBC-CAT staff for access to and assistance with the 19BM beamline at APS. This work was supported by the US National Institutes of Health grants to R.M. and by a grant from the Commonwealth Universal Research Enhancement Program, Pennsylvania Department of Health, awarded to the Wistar Institute.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Rights and permissions
About this article
Cite this article
Zhao, K., Chai, X., Clements, A. et al. Structure and autoregulation of the yeast Hst2 homolog of Sir2. Nat Struct Mol Biol 10, 864–871 (2003). https://doi.org/10.1038/nsb978
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nsb978
This article is cited by
-
Large-scale reverse docking profiles and their applications
BMC Bioinformatics (2012)
-
The NAD+-dependent protein deacetylase activity of SIRT1 is regulated by its oligomeric status
Scientific Reports (2012)
-
Chromatin affinity-precipitation using a small metabolic molecule: its application to analysis of O-acetyl-ADP-ribose
Cellular and Molecular Life Sciences (2012)
-
Primers on chromatin
Nature Structural & Molecular Biology (2007)
-
Nuclear export modulates the cytoplasmic Sir2 homologue Hst2
EMBO reports (2006)