Abstract
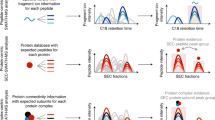
Protein complexes form, dissociate and re-form in order to perform specific cellular functions. In this two-pronged protocol, noncovalent protein complexes are initially isolated by affinity purification for subsequent identification of the components by liquid chromatography high-resolution mass spectrometry (LC-MS) on a hybrid LTQ Orbitrap Velos. In the second prong of the approach, the affinity-purification strategy includes a chemical cross-linking step to 'freeze' a series of concurrently formed, heterogeneous protein subcomplex species that are visualized by gel electrophoresis. This branch of the methodology amalgamates standard and well-practiced laboratory methods to reveal compositional changes that occur in protein complex architecture. By using mouse N-terminally tagged streptavidin-binding peptide–hemagglutinin–TANK-binding kinase 1 (SH-TBK1), we chemically cross-linked the affinity-purified complex of SH-TBK1 with the homobifunctional lysine-specific reagent bis(sulfosuccinimidyl) suberate (BS3), and we separated the resultant protein complexes by denaturation and by silver-stained one- and two-dimensional SDS-PAGE. We observed a range of cross-linked TBK1 complexes of variable pI and Mr and confirmed them by immunoblotting. LC-MS analysis of in situ–digested cross-linked proteins shows differences in the composition of the TBK1 subcomplexes. The protocol is inherently simple and can be readily extended to the investigation of a range of protein complexes. From cell lysis to data generation by LC-MS, the protocol takes approximately 2.5 to 5.5 d to perform.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout


Similar content being viewed by others
References
Rigaut, G. et al. A generic protein purification method for protein complex characterization and proteome exploration. Nat. Biotechnol. 17, 1030–1032 (1999).
Gingras, A.C., Gstaiger, M., Raught, B. & Aebersold, R. Analysis of protein complexes using mass spectrometry. Nat. Rev. Mol. Cell. Biol. 8, 645–654 (2007).
Kocher, T. & Superti-Furga, G. Mass spectrometry-based functional proteomics: from molecular machines to protein networks. Nat. Methods 4, 807–815 (2007).
Collins, M.O. & Choudhary, J.S. Mapping multiprotein complexes by affinity purification and mass spectrometry. Curr. Opin. Biotechnol. 19, 324–330 (2008).
Li, Y. The tandem affinity purification technology: an overview. Biotechnol. Lett. 33, 1487–1499 (2011).
Gavin, A.C. et al. Functional organization of the yeast proteome by systematic analysis of protein complexes. Nature 415, 141–147 (2002).
Ho, Y. et al. Systematic identification of protein complexes in Saccharomyces cerevisiae by mass spectrometry. Nature 415, 180–183 (2002).
Butland, G. et al. Interaction network containing conserved and essential protein complexes in Escherichia coli. Nature 433, 531–537 (2005).
Gavin, A.C. et al. Proteome survey reveals modularity of the yeast cell machinery. Nature 440, 631–636 (2006).
Krogan, N.J. et al. Global landscape of protein complexes in the yeast Saccharomyces cerevisiae. Nature 440, 637–643 (2006).
Glatter, T., Wepf, A., Aebersold, R. & Gstaiger, M. An integrated workflow for charting the human interaction proteome: insights into the PP2A system. Mol. Syst. Biol. 5, 237 (2009).
Behrends, C., Sowa, M.E., Gygi, S.P. & Harper, J.W. Network organization of the human autophagy system. Nature 466, 68–76 (2010).
Pichlmair, A. et al. Viral immune modulators perturb the human molecular network by common and unique strategies. Nature 487, 486–490 (2012).
Ewing, R.M. et al. Large-scale mapping of human protein-protein interactions by mass spectrometry. Mol. Syst. Biol. 3, 89 (2007).
Haura, E.B. et al. Using iTRAQ combined with tandem affinity purification to enhance low-abundance proteins associated with somatically mutated EGFR core complexes in lung cancer. J. Proteome Res. 10, 182–190 (2011).
Haura, E.B. et al. Optimisation of downscaled tandem affinity purifications to identify core protein complexes. J. Integr. OMICS 2, 55–68 (2012).
Blagoev, B. et al. A proteomics strategy to elucidate functional protein-protein interactions applied to EGF signaling. Nat. Biotechnol. 21, 315–318 (2003).
Bouwmeester, T. et al. A physical and functional map of the human TNF-α/NF-κB signal transduction pathway. Nat. Cell. Biol. 6, 97–105 (2004).
Brajenovic, M., Joberty, G., Kuster, B., Bouwmeester, T. & Drewes, G. Comprehensive proteomic analysis of human Par protein complexes reveals an interconnected protein network. J. Biol. Chem. 279, 12804–12811 (2004).
Gloeckner, C.J., Boldt, K., Schumacher, A. & Ueffing, M. Tandem affinity purification of protein complexes from mammalian cells by the Strep/FLAG (SF)-TAP tag. Methods Mol. Biol. 564, 359–372 (2009).
Goncalves, A. et al. Functional dissection of the TBK1 molecular network. PLoS ONE 6, e23971 (2011).
Bertwistle, D., Sugimoto, M. & Sherr, C.J. Physical and functional interactions of the Arf tumor suppressor protein with nucleophosmin/B23. Mol. Cell. Biol. 24, 985–996 (2004).
Vanacova, S. et al. A new yeast poly(A) polymerase complex involved in RNA quality control. PLoS Biol. 3, 986–997 (2005).
Bürckstümmer, T. et al. An efficient tandem affinity purification procedure for interaction proteomics in mammalian cells. Nat. Methods 3, 1013–1019 (2006).
Riedel, C.G. et al. Protein phosphatase 2A protects centromeric sister chromatid cohesion during meiosis I. Nature 441, 53–61 (2006).
Soulat, D. et al. The DEAD-box helicase DDX3X is a critical component of the TANK-binding kinase 1-dependent innate immune response. EMBO J. 27, 2135–2146 (2008).
Brehme, M. et al. Charting the molecular network of the drug target Bcr-Abl. Proc. Natl. Acad. Sci. USA 106, 7414–7419 (2009).
Baumann, C.L. et al. CD14 is a coreceptor of Toll-like receptors 7 and 9. J. Exp. Med. 207, 2689–2701 (2010).
Wojcik, J. et al. A potent and highly specific FN3 monobody inhibitor of the Abl SH2 domain. Nat. Struct. Mol. Biol. 17, 519–527 (2010).
Li, Y., Franklin, S., Zhang, M.J. & Vondriska, T.M. Highly efficient purification of protein complexes from mammalian cells using a novel streptavidin-binding peptide and hexahistidine tandem tag system: application to Bruton's tyrosine kinase. Protein Sci. 20, 140–149 (2011).
Pichlmair, A. et al. IFIT1 is an antiviral protein that recognizes 5′-triphosphate RNA. Nat. Immunol. 12, 624–630 (2011).
Kachaner, D. et al. Plk1-dependent phosphorylation of optineurin provides a negative feedback mechanism for mitotic progression. Mol. Cell. 45, 553–566 (2012).
Gingras, A.C., Aebersold, R. & Raught, B. Advances in protein complex analysis using mass spectrometry. J. Physiol. 563, 11–21 (2005).
Figeys, D. Mapping the human protein interactome. Cell. Res. 18, 716–724 (2008).
Gstaiger, M. & Aebersold, R. Applying mass spectrometry-based proteomics to genetics, genomics and network biology. Nat. Rev. Genet. 10, 617–627 (2009).
Sabido, E., Selevsek, N. & Aebersold, R. Mass spectrometry-based proteomics for systems biology. Curr. Opin. Biotechnol. 23, 591–597 (2012).
Fonslow, B.R. et al. Native capillary isoelectric focusing for the separation of protein complex isoforms and subcomplexes. Anal. Chem. 82, 6643–6651 (2010).
Andersen, J.S. et al. Proteomic characterization of the human centrosome by protein correlation profiling. Nature 426, 570–574 (2003).
Bennett, K.L., Matthiesen, T. & Roepstorff, P. Probing protein surface topology by chemical surface labeling, crosslinking, and mass spectrometry. Methods Mol. Biol. 146, 113–131 (2000).
Bennett, K.L. et al. Chemical cross-linking with thiol-cleavable reagents combined with differential mass spectrometric peptide mapping—a novel approach to assess intermolecular protein contacts. Protein Sci. 9, 1503–1518 (2000).
Sorensen, P. et al. Identification of protein-protein interfaces implicated in CD80-CD28 costimulatory signaling. J. Immunol. 172, 6803–6809 (2004).
Wiener, M.C., Sachs, J.R., Deyanova, E.G. & Yates, N.A. Differential mass spectrometry: a label-free LC-MS method for finding significant differences in complex peptide and protein mixtures. Anal. Chem. 76, 6085–6096 (2004).
Zybailov, B. et al. Statistical analysis of membrane proteome expression changes in Saccharomyces cerevisiae. J. Proteome Res. 5, 2339–2347 (2006).
Ross, P.L. et al. Multiplexed protein quantitation in Saccharomyces cerevisiae using amine-reactive isobaric tagging reagents. Mol. Cell. Proteomics 3, 1154–1169 (2004).
Thompson, A. et al. Tandem mass tags: a novel quantification strategy for comparative analysis of complex protein mixtures by MS/MS. Anal. Chem. 75, 1895–1904 (2003).
Ong, S.E. et al. Stable isotope labeling by amino acids in cell culture, SILAC, as a simple and accurate approach to expression proteomics. Mol. Cell. Proteomics 1, 376–386 (2002).
Gerber, S.A., Rush, J., Stemman, O., Kirschner, M.W. & Gygi, S.P. Absolute quantification of proteins and phosphoproteins from cell lysates by tandem MS. Proc. Natl. Acad. Sci. USA 100, 6940–6945 (2003).
Gordon, J.A. Use of vanadate as protein-phosphotyrosine phosphatase inhibitor. Methods Enzymol. 201, 477–482 (1991).
Abmayr, S.M., Yao, T., Parmely, T. & Workman, J.L. Preparation of nuclear and cytoplasmic extracts from mammalian cells. Curr. Protoc. Mol. Biol. 12, 12.1 (2006).
Cardi, D. et al. Heterologous expression and affinity purification of eukaryotic membrane proteins in view of functional and structural studies: the example of the sarcoplasmic reticulum Ca2+-ATPase. Methods Mol. Biol. 601, 247–267 (2010).
Rappsilber, J., Ishihama, Y. & Mann, M. Stop and go extraction tips for matrix-assisted laser desorption/ionization, nanoelectrospray, and LC/MS sample pretreatment in proteomics. Anal. Chem. 75, 663–670 (2003).
Shevchenko, A., Wilm, M., Vorm, O. & Mann, M. Mass spectrometric sequencing of proteins silver-stained polyacrylamide gels. Anal. Chem. 68, 850–858 (1996).
Kratochwill, K. et al. Alanyl-glutamine dipeptide restores the cytoprotective stress proteome of mesothelial cells exposed to peritoneal dialysis fluids. Nephrol. Dial. Transplant 27, 937–946 (2012).
Kratochwill, K. et al. Interleukin-1 receptor-mediated inflammation impairs the heat shock response of human mesothelial cells. Am. J. Pathol. 178, 1544–1555 (2011).
Kratochwill, K. et al. Stress responses and conditioning effects in mesothelial cells exposed to peritoneal dialysis fluid. J. Proteome Res. 8, 1731–1747 (2009).
Ryzhakov, G. & Randow, F. SINTBAD, a novel component of innate antiviral immunity, shares a TBK1-binding domain with NAP1 and TANK. EMBO J. 26, 3180–3190 (2007).
Chau, T.L. et al. Are the IKKs and IKK-related kinases TBK1 and IKK-epsilon similarly activated? Trends Biochem. Sci. 33, 171–180 (2008).
Colinge, J., Masselot, A., Giron, M., Dessingy, T. & Magnin, J. OLAV: towards high-throughput tandem mass spectrometry data identification. Proteomics 3, 1454–1463 (2003).
Parikh, J.R. et al. multiplierz: an extensible API based desktop environment for proteomics data analysis. BMC Bioinformatics 10, 364 (2009).
Elias, J.E., Haas, W., Faherty, B.K. & Gygi, S.P. Comparative evaluation of mass spectrometry platforms used in large-scale proteomics investigations. Nat. Methods 2, 667–675 (2005).
Acknowledgements
We thank T. Bürckstümmer from Haplogen GmbH, Vienna, for providing the mouse SH-TBK1 entry vector, A.C. Müller for valuable advice on iTRAQ quantification and desalting techniques; and all the members of the Superti-Furga and Bennett laboratories for helpful discussions. Work in our laboratory is supported by the Austrian Academy of Sciences, the Austrian Federal Ministry for Science and Research (Gen-Au projects, APP-III and BIN-III), the Austrian Science Fund FWF and the Central Bank of the Republic of Austria. E.L.R. and R.S. are supported by the GenAu APP-III program (no. 820965), and M.L.H. by the Central Bank of the Republic of Austria (no. 14252).
Author information
Authors and Affiliations
Contributions
E.L.R. developed the protocol, conducted the experiments, interpreted the data and drafted the manuscript. R.S. developed sections of the protocol, participated in optimizing the tandem affinity purification methodology, wrote sections of the manuscript and provided valuable feedback in the review process. K.K. provided intellectual and technical expertise in optimization of the 2D SDS-PAGE, performed the 2D SDS-PAGE experiments, interpreted the gel images and wrote the 2D SDS-PAGE sections of the manuscript. M.L.H. provided technical support with 1D SDS-PAGE and sample preparation for LC-MS, and assisted in writing sections of the manuscript and proofreading of the revised versions. M.G. and G.S.-F. provided intellectual expertise on tandem affinity purifications and valuable feedback during the revision process. K.L.B. and E.L.R. conceived the notion of the protocol. K.L.B. was responsible for project supervision, data interpretation, manuscript writing and providing grant support.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Table 1
SH-tagged TBK1 protein complexes. Complexes are separated by: (A) one- and (B) two-dimensional sodium dodecyl sulphate polyacrylamide gel electrophoresis (1D- and 2D-SDS-PAGE) on 3-8% tris-acetate gradient gels. The different TBK1 sub-complexes and interacting proteins are: (a) SH-TBK1_M+TBK1_H, HS90A, HS90B, AZI2, TANK; (b) SH-TBK1_M+TBK1_H, TANK; (c) SH-TBK1_M+TBK1_H, HS90A, HS90B; (d) HS90A, HS90B; (e) HS90A, HS90B, TBKBP1; (f) SH-TBK1_M, TBK1_H, HS90A, HS90B; (g) SH-TBK1_M, TBK1_H, TBKBP1; (h) SH-TBK1_M, TBK1_H. Bold numbers in (A) are the molecular mass ranges of the 1D-SDS-PAGE. Unique peptide counts, spectral counts (or number of MSMS spectra matched to peptides), and sequence coverage are given for each sub-complex. (PDF 333 kb)
Rights and permissions
About this article
Cite this article
Rudashevskaya, E., Sacco, R., Kratochwill, K. et al. A method to resolve the composition of heterogeneous affinity-purified protein complexes assembled around a common protein by chemical cross-linking, gel electrophoresis and mass spectrometry. Nat Protoc 8, 75–97 (2013). https://doi.org/10.1038/nprot.2012.133
Published:
Issue Date:
DOI: https://doi.org/10.1038/nprot.2012.133
This article is cited by
-
TASL is the SLC15A4-associated adaptor for IRF5 activation by TLR7–9
Nature (2020)
-
A combinatorial screen of the CLOUD uncovers a synergy targeting the androgen receptor
Nature Chemical Biology (2017)
-
JAGN1 deficiency causes aberrant myeloid cell homeostasis and congenital neutropenia
Nature Genetics (2014)
-
Deregulated expression of TANK in glioblastomas triggers pro-tumorigenic ERK1/2 and AKT signaling pathways
Oncogenesis (2013)
-
The CRAPome: a contaminant repository for affinity purification–mass spectrometry data
Nature Methods (2013)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.